* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Current Status and Future Prospects for Public
Pathogenomics wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Genome evolution wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Pharmacogenomics wikipedia , lookup
Genetic code wikipedia , lookup
Genetic drift wikipedia , lookup
Medical genetics wikipedia , lookup
Designer baby wikipedia , lookup
Heritability of IQ wikipedia , lookup
Behavioural genetics wikipedia , lookup
Population genetics wikipedia , lookup
Quantitative trait locus wikipedia , lookup
Genetically modified organism containment and escape wikipedia , lookup
Genetically modified food wikipedia , lookup
Genetic testing wikipedia , lookup
Human genetic variation wikipedia , lookup
Genetically modified crops wikipedia , lookup
Microevolution wikipedia , lookup
History of genetic engineering wikipedia , lookup
Genome (book) wikipedia , lookup
Genetic engineering wikipedia , lookup
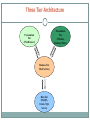
Working in Partnership to Attain Priority Crop Genetic Resource, Genomics, and Genetic Improvement Research Goals PETER BRETTING U S DA - A G R I C U LT U R A L R E S E A R C H S E R V I C E O F F I C E O F N AT I O N A L P R O G R A M S PETER .BRET [email protected] .GOV H T T P : / / W W W. A R S . U S D A . G O V/ R E S E A R C H / P R O GRAMS/PROGRAMS.HTM?NP_CODE=301 Outline for Presentation American Seed Research Summit USDA/ARS National Program in Plant Genetic Resources, Genomics, and Genetic Improvement Plant Genetic Resources and Information Management Plant Genomics, Genetic Analyses and Genome Databases Maize and soybeans: genotyping and nested association mapping Genetic Improvement of Crops Maize: GEM Project Conclusion American Seed Research Summit Research, Education, and Policy Goals and Strategies Strengthen public and private partnerships to accomplish national seed research priorities Coordinate and engage industry stakeholders to support stable funding for seed and breeding education, research, and development Attract and develop a pool of diverse, high-quality plant researchers Ensure that the regulatory system governing the development and implementation of new technology is efficient, effective, and science-based. Develop an education and advocacy program to communicate the value of seed and crop research to the public. USDA/ARS National Program in Plant Genetic Resources, Genomics, and Genetic Improvement (Crop Genes R’ Us!) Largest NP, with 125+ projects, and about $128 million gross annual budget. Conducted by about 300 scientists at more than 50 ARS locations. Extensive public and private sector partnerships. Goal: deliver crop genetic, genomic and bioinformatic tools, information, genetic resources, and improved crop varieties to enhance U. S. agricultural productivity and security. NP 301 Action Plan Research Components Plant and Microbial Genetic Resource Management Crop Informatics, Genomics, and Genetic Analyses Genetic Improvement of Crops The USDA/ARS National Plant Germplasm System (NPGS) One of the largest national genebank systems. More than 510,000 samples of more than 13,400 plant species. Large collections of the major staple crops important to U. S. and world agriculture. Large holdings of crops without major collections at international agricultural research centers, e.g., cotton, soybean, various horticultural and “specialty” crops. Germplasm Resources Information Network (GRIN): an international standard. USDA National Plant Germplasm System (NPGS) Plant Genetic Resource Management in Genebanks Acquisition Characterization Maintenance Evaluation Regeneration Enhancement Documentation and Data Management Distribution Avoidance of Cross Pollination and Seed Mixing Extensive quality assessment when the seed enters the NPGS Procedures and protocols for regeneration Physical isolation Cages Hand pollination Care in storage and distribution to prevent mixing Maize in the NPGS Isolation from sources of out crossing Regenerate ~400 accessions/year Sometimes plantings yield as few as 5,000 seeds for some accessions GRIN-Global GRIN = Germplasm Resources Information Network. http://www.ars-grin.gov/ The genebank information management system for the NPGS, and for Canada’s genebank system (GRIN-Canada). The Global Crop Diversity Trust asked ARS and Bioversity International (an International Agricultural Research Center) to enhance and expand GRIN to address global germplasm information management needs. The Trust awarded ARS a 3-year, $1.4 million grant to develop GRIN-Global; ARS is devoting $900K in-kind support to the project. ARS effort is located in Beltsville, MD and Ames, IA. GRIN-Global Based on GRIN, but can be implemented in both a system-wide and “stand-alone” local management mode Supports multiple users via a “user-friendly” interface Maintains linkages with other databases and interoperates with existing systems Advanced querying, custom and third-party applications Three Tier Architecture Presentation Tier (Windows Desktop Client) Presentation Tier (Web Browser) Business Tier (Web Service) Data Tier (MySQL, Oracle, SQL Server) GRIN-Global On-line ordering/request capability Database-flexible, free of recurrent licensing costs, with interface and database schema source code open and available without restriction to further development Designed to serve as the global standard plant genebank information management system Future Prospects Trends in demand for NPGS germplasm and information vs. NPGS budget NPGS Web Page Access Germplasm Distributions 2,500,000 200000 2,000,000 180000 160000 1,500,000 1,000,000 NPGS Budget 45,000,000 40,000,000 35,000,000 140000 30,000,000 120000 25,000,000 100000 20,000,000 80000 15,000,000 60000 2008 2007 2006 2005 0 2004 0 2003 5,000,000 2002 20000 2001 2008 2007 2006 2005 2004 2003 2002 2001 2000 1999 0 10,000,000 2000 40000 1999 500,000 Result?: mismatch between expanding demand for PGR and static NPGS capacity to manage it. NP 301 Action Plan Research Components Plant and Microbial Genetic Resource Management Crop Informatics, Genomics, and Genetic Analyses Genetic Improvement of Crops Genomic Information and Research Tools Genetic markers: polymorphic and heritable—simple sequence repeats (SSRs) and single nucleotide polymorphisms (SNPs) Expressed sequence tags (ESTs) Genetic maps Quantitative trait loci (QTLs) Physical maps Complete and partial genome sequences Application of Genomic Information to Genetic Analyses: The Nested Association Mapping (NAM) Genetic Map [E. Buckler, J. Holland, M. McMullen (USDA/ARS), and many university and private-sector collaborators] NAM is the most powerful tool for dissection of the genetic bases of quantitative traits for any species – period. Linkage Mapping Recent recombination High power Low resolution Analysis of 2 alleles Moderate marker density Genome scan Association Mapping Historic recombination Low power High resolution Analysis of many alleles High marker density Candidate gene testing Nested Association Mapping Recent and ancient recombination High Power High resolution Analysis of many alleles Moderate genetic marker density High projected marker density SSD NAM × Tzi8 Tx303 P39 Oh7B Oh43 NC358 NC350 MS71 Mo18W M37W M162W Ky21 Ki3 Ki11 Il14H Hp301 CML69 CML52 CML333 CML322 CML277 CML247 CML228 CML103 B97 Nested Association Analysis 25 DL B73 F1s 1 2 200 Yu et al. (2008) Genetics 178: 539 NAM Genotyping and Genetic Map •Genotyping with more than 1500 single nucleotide polymorphism (SNP) genetic markers •Map consists of 1106 loci (38 composite loci) and ~1400 cM genetic distance, therefore an average marker density of 1.3 cM/marker. •It is a composite or consensus map and distances and potentially order can not be assumed to translate directly to individual family maps. Maize Phenomics: Massively Parallel Phenotyping of The Nested Association Mapping Population THE MAIZE DIVERSITY PROJECT Experimental Evaluation of NAM NAM lines evaluated in 2006 – 2007 at each of the following sites: Aurora, NY Champaign, IL Columbia, MO Clayton, NC Homestead, FL 11 Total Environments Ponce, PR + Tassel branch angle Tassel main spike length Tassel branch no. Partitioning Genetic Variance B73 X NC358 B73 X P39 B73 X CML103 RIL1 B73 X IL14H B73 X Tx303 B73 X B97 X M162W B73 X Tzi8 Line-to-line variationB73 within a family – dueB73 to B73 B73 B73 differences between alleles from X X X X B73 and X Ki11 NC350 Oh7Bof that Ki3 alleles from DiverseKy21 Line founder B73 B73 family.B73 B737 B73 X Mo18W B73 X CML52 B73 X M37W X Oh43 B73 X CML322 RIL2 … X MS71 B73 X CML228 xHp301 B73 X CML277 B73 X CML247 …RIL Among-Family Genetic Variance: σ2F RIL199 RIL200 RIL1 RIL2 Family-to-family variation – due to Within-Family Genetic Variances differences among different founders. σ2 σ2G(F)1 B73 X CML69 B73 x CML333 G(F)2 199 RIL200 The Maize Genetics and Genomics Database (Maize GDB) (C. Lawrence et al.) USDA/ARS, AMES, IA www.maizegdb.org/ Phenotypes Integrating structural and genetic maps with maize genomic sequence Genome annotation Research community support Soybean Nested Association Mapping (NAM) Project (P. Cregan, D. Hyten, ARS-Beltsville) Collaborators: USDA/ARS Iowa St. Univ. Univ. of Nebraska Univ. of Illinois Univ. of Tennessee Univ. of Maryland Agricultural Research Service Funding Support Soybean NAM Design focused on yield Populations will be restricted to Maturity Group III Hub parent is IA3023 Lines crossed with IA3023 included both elite and exotic germplasm 121 total lines selected by research community – Crosses made the summer of 2008 Parents were genotyped with the Universal Soy Linkage Panel 1.0 What’s next with the Soybean NAM? During population development, the number of families will be reduced to 40 based on field observations during the inbreeding process. Each population will consist of 250 F5 RILs – Total size: 10,000 lines Phenotyping will be done with a ‘connected’ incomplete block experimental design (Tested in 30 environments) Best Linear Unbiased Predictions (BLUPs) of grain yield, i.e., breeding values for every RIL The Universal Soy Linkage Panel 1.0 (USLP 1.0) With United Soybean Board and collaborator funding 180 sets of the USLP 1.0 (17,280 genotypes) were acquired for gene/QTL Discovery ARS Collaborators Ames, IA Beltsville, MD Columbia, MO Raleigh, NC Stoneville, MS Urbana, IL Wooster, OH State University Collaborators N. Carolina St. Univ. N. Dakota St. Univ. Ohio State Univ. So. Illinois Univ. S. Dakota St. Univ. Univ. of Arkansas Univ. of Georgia Univ. of Illinois Univ. of Minnesota Univ. of Missouri Univ. of Nebraska Virginia Tech Universal Soy Linkage Panel (USLP 1.0) 1,536 SNPs selected from 3,110 SNPs mapped on the Soybean Consensus Map SNPs have diverse allele frequencies Average polymorphism in bi-parental crosses Elite cultivars= 458 PI landraces = 544 Elite crossed with PI landrace = 590 Spread throughout the genome Assayed using the Illumina GoldenGate assay 192 DNA samples run in three days Large orders reduce cost from $11,000 per 96 DNA samples to $5,500 per 96 DNA samples The Universal Soy Linkage Panel 1.0 (USLP 1.0) for Gene/QTL Discovery - Traits Under Study in Collaborative Projects Enhanced Seed Composition Resistance to Biotic Stress Resistance to Abiotic Stress Reduced linolenic acid oil Elevated oleic acid oil Lower saturates Higher protein Soybean Cyst Nematode Soybean Rust Phytophthora Root Rot Foliar Feeding Insects Soybean Aphid Drought Iron Deficiency Chlorosis Seed Yield – Assessment and enhancement of genetic diversity Source of Genetic Improvement for Soybean USDA germplasm collection is the source for new genetic variation for soybean improvement for every important trait USDA Soybean Collection (21,000 accessions) 1,116 wild soybeans 17 Introduced landraces (86% of the Germplasm base) 18,090 accessions collected mostly from Asia (Landraces) 513 Public cultivars released after Hyten et al. 2006, PNAS 103: 16666-16671 1946 Soy HapMap • The United Soybean Board recently agreed to fund a $2.9M • project to characterize the entire USDA Soybean Germplasm Collection of 21,000+ wild and cultivated soybeans with 50,000 SNP DNA markers. The Illumina’s Beadstation will allow genotyping of this collection with 50,000 SNPs within three years • New gene discovery through association analysis • Enables breeders to select germplasm with greatest potential for • agronomic improvement Decipher the signatures of selection (allele frequency changes) associated with soybean yield improvement over 75 years of soybean breeding to help understand yield Soybean Genomics and Improvement Lab USDA, ARS, BARC-West Beltsville, Maryland 60,800 SNP Infinium Chip Research by R. Shoemaker (ARS-Ames), C. Vance (ARS-St. Paul) and collaborators at N. Dakota State Univ. and Iowa State Univ. Iron is a limiting growth factor on 30% of cropland Iron is also a major nutritional deficiency in much of the world Using global gene expression profiling to identify genes involved in iron metabolism Currently evaluating how to control the genetic expression to enhance iron balance in commercial plant varieties. Developing molecular markers to speed the public release of commercial varieties with improved iron efficiency. Mapping & Cloning Genes Responsible for Soybean Protein Research by R. Shoemaker (ARS-Ames), C. Vance (ARS-St. Paul), collaborators at the Univ. of Illinois and Univ. of Nebraska The locus of a major trait that controls seed protein content was mapped Chromosome 20 High-throughput transcript sequencing and mapping identified candidate genes Currently identifying the specific and evaluating modifications to change the protein content to 15% Soluble Carbohydrates Soybean Seed Composition 18% Oil several (sucrose, stachyose, raffinose, others) genes 14% Moisture, ash, other 38% Protein 15% Insoluble Carbohydrates SNP Haplotypes and Linkage Disequilibrium (R. Shoemaker, ARS-Ames) Haplotype and Allele States for varieties NP 301 Action Plan Research Components Plant and Microbial Genetic Resource Management Crop Informatics, Genomics, and Genetic Analyses Genetic Improvement of Crops Germplasm Enhancement of Maize Germplasm Enhancement of Maize (GEM) Project A collaborative effort of public and private sector researchers to broaden and enhance the maize germplasm base. More than 60 collaborators. Two permanent breeding sites: Ames, IA for development of 25% tropical and temperate exotic Raleigh, NC for development of 50% tropical GEM is administered by the USDA-ARS Plant Introduction Research Unit (PIRU) located in Ames, IA; and the Plant Science Research Unit (PSRU) in Raleigh, NC Technical Steering Group (TSG) provides guidelines for research, germplasm, and methods. Germplasm Enhancement of Maize GEM Project Cooperators Category Number Private US companies 29 Public US universities 13 USDA-ARS research units 8 Non-government organization 1 Private international companies 11 Public international institutes 4 Total 66 Germplasm Enhancement of Maize GEM Objectives Manage and coordinate a multi-site cooperative program for germplasm evaluation, development, and information sharing Evaluate diverse maize germplasm for adaptation, yield, stress resistance, and key value-added traits (VATs) Develop and release enhanced germplasm with key traits Develop innovative means of managing and transferring information to the maize community Germplasm Enhancement of Maize GEM Germplasm Releases Year # Lines Released 2001 2002 2003 2003 2003 2003 2003 2004 2004 2004 2004 2005 2005 2005 2006 2006 2007 2007 2007 2008 2009 2009 Total 1* 2* 29** 1* 1 42 9* 14 2 1 9 9 1 19 13 3 10 10 1* 13 7 5 202 Institution Germplasm Attributes USDA-ARS Ames Univ. of Delaware NC State Univ. Ohio State Univ. Univ. of Delaware USDA-ARS Ames NC State Univ. USDA-ARS Ames Texas A&M Univ. Univ. of Wisconsin NC State Univ. USDA-ARS Ames Univ. of Delaware NC State Univ. USDA-ARS Ames NC State Univ. USDA-ARS Ames NC State Univ. Truman State Univ. USDA-ARS Ames USDA-ARS Ames USDA-ARS Raleigh Resistance to 1st brood ECB (non-DIMBOA) Yield, resistance to anthracnose and GLS Yield, earlier flowering, GLS, Fusarium Resistance Yield, Fusarium resistance VAT Temperate adaptation, GLS, VAT Yield, VAT, GLS Temperate adaptation, yield, VAT Stress tolerance, yield, CEW, grain mold resistance Superior nutritional quality/yield Yield, earlier flowering, VAT Temperate adaptation, yield, VAT High protein Yield, earlier flowering, VAT Yield, VAT Yield, earlier flowering Protein, oil, high starch for ethanol 50% exotics; disease resistance Amylose maize VII line (GEMS-0067) Temperate adaptation, yield, VAT, waxy lines Protein, oil 50% exotic; protein, oil * Crop Science registered and ** 20 of these 29 lines were Crop Science registered. Germplasm Enhancement of Maize Number of Releases by Traits Germplasm Traits Number Protein above 13% 33 Amino Acid index (Met. Lys. Trp.) 17 Oil above 4.5% 13 Starch compositional properties 24 High Amylose (70%) 1 Waxy 2 Fumonisin (reduced level) 5 Aflatoxin (reduced level) 6 Leaf disease resistance (GLS, NLB, & SLB) 104 Other (yield, Y/M, etc.) 12 Germplasm Enhancement of Maize New GEM Initiatives Allelic diversity New un-sampled races being tapped Goal to assess ~300 races for adaptation in US New elite exotic sources being acquired More than 40 germplasm sources acquired from Thailand, Peru, Nigeria, Argentina, Chile, and France Provide resistance to exotic diseases Shade house to reduce photoperiod response Make tropical introgressions in Ames, IA 23 tropical sources (11 races) successful (so far) Double haploid research Explore feasibility with exotic germplasm Germplasm Enhancement of Maize Fusarium Ear Rot Susceptible Line Resistant Line GEMS-0002 Public Release Bill Dolezal, Pioneer Hi-Bred Int’l, Woodland, CA (2005) Integrated national programmatic approach, with extensive academic and private sector collaborations/partnerships Exploit untapped genetic diversity in genebanks, and breeding populations Plant genomics, gene discovery Genetic markers & Bioinformatics Genome Databases Genetic Enhancement and Breeding American Seed Research Summit Research, Education, and Policy Goals and Strategies Strengthen public and private partnerships to accomplish national seed research priorities Coordinate and engage industry stakeholders to support stable funding for seed and breeding education, research, and development Attract and develop a pool of diverse, high-quality plant researchers Ensure that the regulatory system governing the development and implementation of new technology is efficient, effective, and science-based. Develop an education and advocacy program to communicate the value of seed and crop research to the public. Thanks! Thanks to the NCCPB and ASTA for the invitation to speak Thanks to USDA/ARS scientists for sharing information and data Thanks to our partners in the seed industry and academia for their invaluable collaboration