* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download 91.510_ch9_2
Genetic code wikipedia , lookup
Gene expression wikipedia , lookup
Biochemistry wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Expression vector wikipedia , lookup
Magnesium transporter wikipedia , lookup
Ancestral sequence reconstruction wikipedia , lookup
Metalloprotein wikipedia , lookup
Point mutation wikipedia , lookup
Interactome wikipedia , lookup
Western blot wikipedia , lookup
Protein purification wikipedia , lookup
Structural alignment wikipedia , lookup
Protein–protein interaction wikipedia , lookup
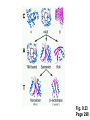
Protein structure (Part 2 of 2) Copyright notice Many of the images in this powerpoint presentation are from Bioinformatics and Functional Genomics by Jonathan Pevsner (ISBN 0-471-21004-8). Copyright © 2003 by John Wiley & Sons, Inc. These images and materials may not be used without permission from the publisher. We welcome instructors to use these powerpoints for educational purposes, but please acknowledge the source. The book has a homepage at http://www.bioinfbook.org Including hyperlinks to the book chapters. Many databases explore protein structures SCOP CATH Dali Domain Dictionary FSSP Page 293 Structural Classification of Proteins (SCOP) SCOP describes protein structures using a hierarchical classification scheme: Classes Folds Superfamilies (likely evolutionary relationship) Families Domains Individual PDB entries http://scop.mrc.lmb.cam.ac.uk/scop/ Page 293 Page 297 SCOP statistics (October, 2002) Class All a All b a/b a+b … Total # folds 151 110 113 208 686 # superfamilies 252 205 185 295 1073 # families 393 337 438 454 1827 Page 298 Class, Architecture, Topology, and Homologous Superfamily (CATH) database CATH clusters proteins at four levels: C Class (a, b, a&b folds) A Architecture (shape of domain, e.g. jelly roll) T Topology (fold families; not necessarily homologous) H Homologous superfamily http://www.biochem.ucl.ac.uk/basm/cath_new Page 293 Fig. 9.23 Page 298 Fig. 9.24 Page 299 Fig. 9.24 Page 299 Fig. 9.25 Page 300 Fig. 9.25 Page 300 Page 301 Fig. 9.27 Page 302 Fig. 9.28 Page 303 Dali Domain Dictionary Dali contains a numerical taxonomy of all known structures in PDB. Dali integrates additional data for entries within a domain class, such as secondary structure predictions and solvent accessibility. Page 302 Fig. 9.29 Page 303 Fig. 9.30 Page 304 Fig. 9.30 Page 304 Fig. 9.30 Page 304 Fold classification based on structure-structure alignment of proteins (FSSP) FSSP is based on a comprehensive comparison of PDB proteins (greater than 30 amino acids in length). Representative sets exclude sequence homologs sharing > 25% amino acid identity. The output includes a “fold tree.” http://www.ebi.ac.uk/dali/fssp Page 293 Fig. 9.31 Page 305 FSSP: fold tree Fig. 9.32 Page 306 Fig. 9.33 Page 307 Fig. 9.34 Page 307 Approaches to predicting protein structures There are about >20,000 structures in PDB, and about 1 million protein sequences in SwissProt/ TrEMBL. For most proteins, structural models derive from computational biology approaches, rather than experimental methods. The most reliable method of modeling and evaluating new structures is by comparison to previously known structures. This is comparative modeling. An alternative is ab initio modeling. Page 303-305 Approaches to predicting protein structures obtain sequence (target) fold assignment comparative modeling ab initio modeling build, assess model Page 308 Comparative modeling of protein structures [1] Perform fold assignment (e.g. BLAST, CATH, SCOP); identify structurally conserved regions [2] Align the target (unknown protein) with the template. This is performed for >30% amino acid identity over a sufficient length [3] Build a model [4] Evaluate the model Page 305 Errors in comparative modeling Errors may occur for many reasons [1] Errors in side-chain packing [2] Distortions within correctly aligned regions [3] Errors in regions of target that do not match template [4] errors in sequence alignment [5] use of incorrect templates Page 306 Comparative modeling In general, accuracy of structure prediction depends on the percent amino acid identity shared between target and template. For >50% identity, RMSD is often only 1 Å. Page 306 Baker and Sali (2000) Page 308 Comparative modeling Many web servers offer comparative modeling services. Examples are SWISS-MODEL (ExPASy) Predict Protein server (Columbia) WHAT IF (CMBI, Netherlands) Page 309 Ab initio protein structure prediction Ab initio prediction can be performed when a protein has no detectable homologs. Protein folding is modeled based on global free-energy minimum estimates. The “Rosetta Stone” methods was applied to sequence families lacking known structures. For 80 of 131 proteins, one of the top five ranked models successfully predicted the structure within 6.0 Å RMSD (Bonneau et al., 2002). Page 309-310 Protein structure and human disease In some cases, a single amino acid substitution can induce a dramatic change in protein structure. For example, the DF508 mutation of CFTR alters the a helical content of the protein, and disrupts intracellular trafficking. Other changes are subtle. The E6V mutation in the gene encoding hemoglobin beta causes sicklecell anemia. The substitution introduces a hydrophobic patch on the protein surface, leading to clumping of hemoglobin molecules. Page 311 Protein structure and human disease Disease Cystic fibrosis Sickle-cell anemia “mad cow” disease Alzheimer disease Protein CFTR hemoglobin beta prion protein amyloid precursor protein Table 9.5 Page 312