* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Further Details of Mechanism
Butyric acid wikipedia , lookup
Lactate dehydrogenase wikipedia , lookup
Fatty acid metabolism wikipedia , lookup
Photosynthesis wikipedia , lookup
Fatty acid synthesis wikipedia , lookup
Biosynthesis wikipedia , lookup
Adenosine triphosphate wikipedia , lookup
Metalloprotein wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Nicotinamide adenine dinucleotide wikipedia , lookup
Light-dependent reactions wikipedia , lookup
Evolution of metal ions in biological systems wikipedia , lookup
Microbial metabolism wikipedia , lookup
Electron transport chain wikipedia , lookup
Photosynthetic reaction centre wikipedia , lookup
Biochemistry wikipedia , lookup
NADH:ubiquinone oxidoreductase (H+-translocating) wikipedia , lookup
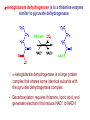
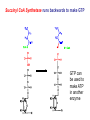
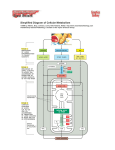
The Citric Acid Cycle Can Be a Multistep Catalyst • Oxaloacetate is regenerated • The cycle is a mechanism for oxidizing acetyl CoA to CO2 by NAD+ and Q • The cycle itself is not a pathway for a net degradation of any cycle intermediates • Cycle intermediates can be shared with other pathways, which may lead to a resupply or net decrease in cycle intermediates 1. Citrate Synthase • Citrate formed from acetyl CoA and oxaloacetate • Only cycle reaction with C-C bond formation Proposed mechanism of citrate synthase Citrate synthase generates a new C-C bond using acetyl CoA in a Claisen condensation 1. The enolate nucleophile is generated by “push-pull” acid/base catalysis 2. The enolate attacks the -keto position of oxaloacetate in a reversible aldol reaction 3. Hydrolysis of the thioester, also catalyzed by Asp375 and His274, is exothermic and essentially irreversible. This drives the reaction to completion. 2. Aconitase • Elimination of H2O from citrate to form C=C bond of cis-aconitate • Stereospecific addition of H2O to cis-aconitate to form 2R,3S-Isocitrate Reaction of Aconitase 3. Isocitrate Dehydrogenase • Oxidative decarboxylation of isocitrate to -ketoglutarate (-kg) (a metabolically irreversible reaction) • One of four oxidation-reduction reactions of the cycle • Hydride ion from the C-2 of isocitrate is transferred to NAD+ to form NADH • Oxalosuccinate is decarboxylated to -kg Spontaneous CO2 production can occur with a -keto acid -keto acids are unstable in protic solvents • They will pick up a proton on the ketone oxygen and this triggers decarboxylation. • Generation of CO2 makes this reaction irreversible and highly exothermic. • How do you capture the energy from this process? • In biochemistry, the energy of “combustion” is partially harvested by coupling most decarboxylation reactions to an oxidation reaction. Isocitrate dehydrogenase reaction Aconitase uses an FeS cluster as a Lewis acid catalyst Isocitrate dehydrogenase captures the energy of decarboxylation •The oxidation of an alcohol to a ketone in oxalosuccinate is reversible and near equilibrium. •Lewis acid catalyzed decarboxylation of the -keto acid drives this reaction to 100% completion. •Energy of decarboxylation is “captured” in the reduced NADH through the coupling of these two reactions on the same enzyme. The Citric Acid Cycle Oxidizes AcetylCoA • Table 12.2 4. The a-Ketoglutarate Dehydrogenase Complex Structure of a-Ketoglutarate dehydrogenase complex • Similar to pyruvate dehydrogenase complex • Same coenzymes, identical mechanisms E1 - -ketoglutarate dehydrogenase (with TPP) E2 - succinyltransferase (with flexible lipoamide prosthetic group) E3 - dihydrolipoamide dehydrogenase (with FAD) -ketoglutarate dehydrogenase is is a thiamine enzyme similar to pyruvate dehydrogenase -ketoglutarate dehydrogenase is a large protein complex that shares some identical subunits with the pyruvate dehydrogenase complex • Decarboxylation requires thiamine, lipoic acid, and generates electrons that reduce NAD+ to NADH Succinyl CoA Synthetase runs backwards to make GTP GTP can be used to make ATP in another enzyme Thioester is used to make a phosphoanhydride bond Phosphohistidine is high energy attacked to make a phosphodiester bond 6. The Succinate Dehydrogenase (SDH) Complex • Located on the inner mitochondrial membrane (other components are dissolved in the matrix) • Dehydrogenation is stereospecific; only the trans isomer is formed • Substrate analog malonate is a competitive inhibitor of the SDH complex Reaction of the succinate dehydrogenase complex (a) Ubiquinone, (b) Plastoquinone • Hydrophobic tail of each is composed of 6 to 10 five-carbon isoprenoid units • The isoprenoid chain allows these quinones to dissolve in lipid membranes • Three oxidation states of ubiquinone • Ubiquinone is reduced in two one-electron steps via a semiquinone free radical intermediate. Reactive center is shown in red. Succinate dehydrogenase contains an FAD cofactor Eh′ = -0.040 V (NAD+ is -0.320 V) 7. Fumarase • Stereospecific trans addition of water to the double bond of fumarate to form L-malate Fumarase catalyzes hydration of an alkene Class II fumarase (human and yeast) uses acid/base chemistry to catalyze trans addition of water via a carbanion intermediate Histidine Lysine Class I fumarase (bacteria) uses an iron-sulfur cluster to catalyze the same chemistry. It has not been studied experimentally, but it is assumed to work by a mechanism similar to aconitase. 8. Malate Dehydrogenase Reduced Coenzymes Fuel the Production of ATP • Each acetyl CoA entering the cycle nets: (1) 3 NADH (2) 1 QH2 (3) 1 GTP (or 1 ATP) • Oxidation of each NADH yields 2.5 ATP • Oxidation of each QH2 yields 1.5 ATP • Complete oxidation of 1 acetyl CoA = 10 ATP Glucose degradation via glycolysis, citric acid cycle, and oxidative phosphorylation Three point attachment of prochiral substrates to enzymes • Chemically identical groups a1 and a2 of a prochiral molecule can be distinguished by the enzyme • Fates of carbon atoms in the cycle • Carbon atoms from acetyl CoA (red) are not lost in the first turn of the cycle Reducing Power • Electrons of reduced coenzymes flow toward O2 • This produces a proton flow and a transmembrane potential • Oxidative phosphorylation is the process by which the potential is coupled to the reaction: ADP + Pi ATP Reduced Coenzymes Conserve Energy from Biological Oxidations • Amino acids, monosaccharides and lipids are oxidized in the catabolic pathways • Oxidizing agent - accepts electrons, is reduced • Reducing agent - loses electrons, is oxidized • Oxidation of one molecule must be coupled with the reduction of another molecule Ared + Box Aox + Bred Diagram of an electrochemical cell • Electrons flow through external circuit from Zn electrode to the Cu electrode Reduction Potentials Cathode (Reduction) Half-Reaction Li+(aq) + e- -> Li(s) K+(aq) + e- -> K(s) Ca2+(aq) + 2e- -> Ca(s) Na+(aq) + e- -> Na(s) Zn2+(aq) + 2e- -> Zn(s) Cu2+(aq) + 2e- -> Cu(s) O3(g) + 2H+(aq) + 2e- -> O2(g) + H2O(l) F2(g) + 2e- -> 2F-(aq) Standard Potential E° (volts) -3.04 -2.92 -2.76 -2.71 -0.76 0.34 2.07 2.87 Standard reduction potentials and free energy • Relationship between standard free-energy change and the standard reduction potential: DGo’ = -nFDEo’ n = # electrons transferred F = Faraday constant (96.48 kJ V-1) DEo’ = Eo’electron acceptor - Eo’electron donor Example Suppose we had the following voltaic cell at 25o C: Cu(s)/Cu+2 (1.0 M) // Ag+(1.0 M)/ Ag (s) What would be the cell potential under these conditions? Example Suppose we had the following voltaic cell at 25o C: Cu(s)/Cu+2 (1.0 M) // Ag+(1.0 M)/ Ag (s) What would be the cell potential under these conditions? Ag+ + e- ---> Ag0 Cu+2 + 2e- ----> Cu0 E0red = + 0.80 v E0red = + 0.337 v Example: Biological Systems Both NAD+ and FAD are oxidizing agents The question is which would oxidize which? OR Which one of the above is the spontaneous reaction? in which DG is negative To be able to answer the question We must look into the “electron donation” capabilities of NADH and FADH2 i.e. reduction potentials of NADH and FADH2 Remember, DEo’ = Eo’electron acceptor - Eo’electron donor For a spontaneous reaction DEo ’ must be positive Therefore, rearrange Add the two reactions electron acceptor electron donor Regulation of the Citric Acid Cycle • Pathway controlled by: (1) Allosteric modulators (2) Covalent modification of cycle enzymes (3) Supply of acetyl CoA (4) Regulation of pyruvate dehydrogenase complex controls acetyl CoA supply Regulation of the PDH complex • Increased levels of acetyl CoA and NADH inhibit E2, E3 in mammals and E. coli Regulation of mammalian PDH complex by covalent modification • Phosphorylation/dephosphorylation of E1 Further regulation of the PDH complex Pyruvate dehydrogenase kinase (PDK) • PDK is activated by NADH and acetyl CoA (leads to inactivation of the PDH complex) • PDK is inhibited by pyruvate and ADP (leads to activation of the PDH complex) Pyruvate dehydrogenase phosphatase (PDP) • PDP activity is stimulated by Ca2+ (leads to an activation of the PDH complex) Control points in the citric acid cycle Rate is adjusted to meet the cell’s need for ATP. Three allosteric enzyme control points: PDH - inhibited by NADH, acetyl CoA, and ATP. Isocitrate dehydrogenase stimulated by ADP; inhibited by ATP and NADH a-ketoglutarate dehydrogenase— inhibited by NADH, succinyl CoA, high energy charge. The Glyoxylate Cycle • Pathway for the formation of glucose from noncarbohydrate precursors in plants, bacteria and yeast (not animals) • Glyoxylate cycle leads from 2-carbon compounds to glucose • In animals, acetyl CoA is not a carbon source for the net formation of glucose (2 carbons of acetyl CoA enter cycle, 2 are released as 2 CO2) Glyoxylate cycle - formation of glucose • Formation of glucose from acetyl CoA (or any substrate that is a precursor to acetyl CoA) • Ethanol or acetate can be metabolized to acetyl CoA and then to glucose via the glyoxylate cycle • Stored seed oils in plants are converted to carbohydrates during germination Isocitrate lyase: first bypass enzyme of glyoxylate Malate synthase: second bypass enzyme of glyoxylate Glyoxylate cycle in germinating castor beans • Conversion of acetyl CoA to glucose requires the transfer of metabolites among three metabolic compartments (1) The glyoxysome (2) The cytosol (3) The mitochondrion