* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Proteins File
Implicit solvation wikipedia , lookup
Rosetta@home wikipedia , lookup
Protein design wikipedia , lookup
Bimolecular fluorescence complementation wikipedia , lookup
Structural alignment wikipedia , lookup
Cooperative binding wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Circular dichroism wikipedia , lookup
Protein folding wikipedia , lookup
Protein moonlighting wikipedia , lookup
Alpha helix wikipedia , lookup
Homology modeling wikipedia , lookup
Protein domain wikipedia , lookup
Protein purification wikipedia , lookup
List of types of proteins wikipedia , lookup
Protein mass spectrometry wikipedia , lookup
Western blot wikipedia , lookup
Protein structure prediction wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
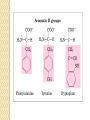
CHM 708: MEDICINAL CHEMISTRY Proteins a-Amino-acids: Building blocks of proteins Proteins are made up of 20 major aminoacids with different side chains Peptide bonds Primary Structure A protein consists of a long chain (50 or more) or polymer of a-amino-acids, linked together by peptide (amide) bonds. Short chains are called peptides. The free NH3+ end is called the N-terminus, and the free CO2- end the C terminus. The primary structure of a protein is the sequence of amino-acids in the chain. The sequence of -N–C–C- segments in the chain is called the backbone. Secondary Structure This refers to the local spatial arrangement of the backbone atoms of segments of the protein chain. Three main types: ◦ a-Helix ◦ b-Sheet ◦ Irregular coils. All are stabilised by H-bonding between backbone NH and C=O groups. a-Helix b-Strand (anti-parallel) b-Strand (parallel) b-Sheet Tertiary structure This is the overall arrangement of all the atoms in the protein, i.e., its overall shape. Every protein has a natural tertiary structure – most stable shape. It is active only in that shape. Tertiary structure is determined by primary structure. Tertiary structure is stabilised by H-bonds, ionic bonds, polar interactions, hydrophobic interactions, dispersion (Van der Waals) forces, and certain types of covalent bonds. Tertiary structures Fatty-acid binding protein Myoglobin Multi-subunit Proteins Some complex proteins consist of two or more amino-acid chains. Each chain is called a subunit and has its own tertiary structure. The arrangement of subunits in the overall protein is called the quaternary structure. Quaternary structure is stabilised by the same forces as tertiary structure. So is the binding of small molecules to specific sites on the surface of the protein. Quaternary structure Yellow: a subunits Red: b subunits Haemoglobin Functions of proteins Structural proteins (skin, hair, nails, etc.) Muscle proteins (able to contract, use energy to do work). Transport proteins. Cell membrane proteins. ◦ Membrane channels and pores ◦ Molecular recognition and signalling. Enzymes: catalysts of biological reactions. ◦ Membrane-bound ◦ Intra-cellular ◦ Extra-cellular. Protein-ligand binding Most proteins function by, or as a result of, reversibly binding a small molecule (ligand). Each ligand has a specific binding site (active site), into which it fits. The binding of a particular ligand to a protein can be modulated by other ligands binding to the same protein. Drug molecules binding to the protein at the active site or other sites can interfere with its normal functioning.