* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download MedicalAspectsVariations
SNP genotyping wikipedia , lookup
Transgenerational epigenetic inheritance wikipedia , lookup
Polymorphism (biology) wikipedia , lookup
Gene therapy wikipedia , lookup
Non-coding DNA wikipedia , lookup
Hardy–Weinberg principle wikipedia , lookup
Whole genome sequencing wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Dominance (genetics) wikipedia , lookup
Human genome wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Genome evolution wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Heritability of IQ wikipedia , lookup
History of genetic engineering wikipedia , lookup
Genetic testing wikipedia , lookup
Genetic drift wikipedia , lookup
Genetic engineering wikipedia , lookup
Designer baby wikipedia , lookup
Pharmacogenomics wikipedia , lookup
Population genetics wikipedia , lookup
Behavioural genetics wikipedia , lookup
Genome (book) wikipedia , lookup
Microevolution wikipedia , lookup
Medical genetics wikipedia , lookup
Human genetic variation wikipedia , lookup
Quantitative trait locus wikipedia , lookup
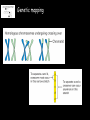
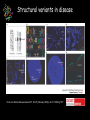
The medical relevance of genome variability Gabor T. Marth, D.Sc. Department of Biology, Boston College [email protected] Medical Genomics Course – Debrecen, Hungary, May 2006 Lecture overview 1. Phenotypic effects caused by known genetic variants 2. Genetic mapping to find genetic variants that cause diseases – linkage analysis and association studies 3. Genome-wide association mapping resources – the HapMap 4. Structural and epigenetic variations in disease 1. Phenotypic effects caused by known genetic variants Many SNPs do have phenotypic effects some notable genetic diseases: cystic fibrosis cycle-cell anemia Badano and Katsanis, NRG 2002 Genetic variants in Pharmacogenetics Evans and Rellig, Science 1999 Genetic variants in Pharmacogenetics Evans and Rellig, Science 1999 Using genotype information in the drug development pipeline Roses. NRG 2004 Are all genetic variants functional? ~ 10 million known SNPs SNPs, on the scale of the genome, can be described well with the “neutral theory” of sequence variations the vast majority of SNPs likely to have no functional effects 0.4 0.3 0.2 0.1 0 16kb 16 kb 12 kb 12 kb 0.00 5.00 10.00 8kb 8 kb 15.00 20.00 25.00 4 kb 30.00 4 kb 35.00 40.00 How do we find the few functional variants in the background of millions of non-functional SNPs? 2. Genetic mapping to find genetic variants that cause diseases – linkage analysis and association studies Genetic mapping Allelic association (linkage) • allelic association is the nonrandom assortment between alleles i.e. it measures how well knowledge of the allele state at one site permits prediction at another marker site functional site • significant allelic association between a marker and a functional site permits localization (mapping) even without having the functional site in our collection • allelic association, and the use of genetic markers is the basis for mapping functional alleles Mendelian diseases have simple inheritance genotype inheritance genotype + phenotype inheritance Linkage analysis compares the transmission of marker genotype and phenotype in families Complex disease – complex inheritance Badano and Katsanis, NRG 2002 Allele frequency and relative risk Brinkman et al. Nature Reviews Genetics advance online publication; published online 14 March 2006 | doi:10.1038/nrg1828 Association study strategies • region(s) interrogated: single gene, list of candidate genes (“candidate gene study”), or entire genome (“genome scan”) • direct or indirect: causative variant • single-SNP marker or multiSNP haplotype marker • single-stage or multi-stage marker that is co-inherited with causative variant causative variant Association study strategies for economy, one cannot genotype every SNP in thousands of clinical samples: marker selection is the process where a subset of all available SNPs is chosen 1. hypothesis driven (i.e. based on gene function) 2. LD-driven – based entirely on the reduction of redundancy presented by the linkage disequilibrium (LD) between SNPs; tags represent other SNPs they are correlated with causative variant Case-control association testing • genotyping cases and controls at various polymorphisms clinical cases • searching for markers with “significant” marker allele frequency differences between cases and controls; these marker signify regions of possible causative alleles AF(controls) clinical controls AF(cases) Marker selection depends on genome LD Daly et al. NG 2001 3. Genome-wide association mapping resources – the HapMap The HapMap resource • goal: to map out human allele and association structure of at the kilobase scale • deliverables: a set of physical and informational reagents LD structure in four human populations International HapMap Consortium, Nature 2005 Genome-wide scans for human diseases SNPs in Complement Factor H (CFH) gene are associated with Age-related Macular Degeneration (AMD) Klein et al, Science 2005 4. Structural and epigenetic variants in disease Structural variants in disease Feuk et al. Nature Reviews Genetics 7, 85–97 (February 2006) | doi:10.1038/nrg1767 Structural variations and phenotype Feuk et al. Nature Reviews Genetics 7, 85–97 (February 2006) | doi:10.1038/nrg1767 Epigenetics and cancer Baylin at al. NRC 2006.