* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download protein - Portal UniMAP
Gel electrophoresis wikipedia , lookup
Ribosomally synthesized and post-translationally modified peptides wikipedia , lookup
Size-exclusion chromatography wikipedia , lookup
Paracrine signalling wikipedia , lookup
Signal transduction wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Gene expression wikipedia , lookup
Genetic code wikipedia , lookup
Expression vector wikipedia , lookup
Ancestral sequence reconstruction wikipedia , lookup
Magnesium transporter wikipedia , lookup
Biosynthesis wikipedia , lookup
Point mutation wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Bimolecular fluorescence complementation wikipedia , lookup
Homology modeling wikipedia , lookup
Interactome wikipedia , lookup
Metalloprotein wikipedia , lookup
Biochemistry wikipedia , lookup
Protein purification wikipedia , lookup
Western blot wikipedia , lookup
Two-hybrid screening wikipedia , lookup
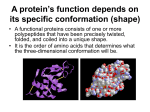
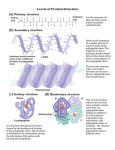
PTT 103 BIOCHEMISTRY PROTEIN Pn Khadijah Hanim Abdul Rahman Protein structure - Protein (polypeptides) are organic compound made of amino acids arranged in a linear form and folded into specific conformation Protein - essential part of organisms - most diverse functions Function of proteins: Catalysis (enzymes), structure (collagen, elastin), movement (actin, tubulin), Defense (keratin, fibrinogen, thrombin), regulation (insulin), transport (function as carriers across membrane), storage (casein), stress response Protein structure When cell synthesizes a polypeptides, the chain folded spontaneously This folding is reinforced by variety of bonds between the chain In a complex structure of protein, several levels of the structural organization of proteins : a) primary structure c) tertiary structure b) secondary structure d) quaternary structure a) Primary structure Primary structure of protein is its unique sequence of amino acids forming its polypeptide chain Every polypeptides has a specific amino acid sequence the primary structure of a protein is starting from the amino-terminal (N) end to the carboxyl-terminal (C) end. The interactions between amino acid residues determine the protein’s 3-D structure and its functional role. Primary structure of enzyme lysozyme b) Secondary structure Most proteins have segments of their polypeptide chain repeatedly coiled or folded in patterns. These coiled & folded referred as secondary structure. 2 types of secondary structure : - α-helix stabilized by hydrogen bond - β-pleated sheet between carbonyl & amino groups in the polypeptide’s backbone α-helix Rigid, rod like structure that forms when a polypeptides chain twists into a right-handed helical conformation Hydrogen bond form between amino group (NH) of each amino acid and the carbonyl group of the amino acid four residue away (H bond form between 4 amino acid) There are 3.6 a.a residues per-turn helix. R group extend outward from the helix Because of several structural constraints, certain a.a do not foster α-helical formation Glycine R group is too small that the p.p chain may be too flexible. Proline: contains a rigid ring that prevents NCα bond from rotating. Proline also has no NH group available to form H bonds that are crucial in α-helix Bulky R groups a.a are also incompatible with α-helix structure. β-pleated sheet Form when two or more polypeptide chain segments line up side by side Each individual segment = β-strand Each β-strand is fully extended β-pleated sheet stabilized by hydrogen bonds form between the polypeptide backbone N-H and carbonyl groups of adjacent chains Two β-pleated sheet : - parallel - polypeptide chain arranged in same direction - antiparallel - polypeptide chain arranged in opposite direction - more stable because fully colinear H bonds form. Usually mixed parellal-antiparallel β-pleated sheet observed in proteins combination of α-helix and β-pleated sheet secondary structure = supersecondary structure Supersecondary structure : a) βαβ unit two parallel β-pleated sheets connected by α-helix fragment b) β-meander two antiparallel β-sheets are connected by polar amino acids and glycines to effect an abrupt change in direction of the polypeptide chain (reverse or β-turns) b meander c) αα-units two α-helices separated by loop or nonhelical segment d) β-barrel- form when various β-sheet configurations fold back on themselves e) Greek key- antiparallel βsheet doubles back on itself in a pattern that resemble a common greek pottery design c) Tertiary structure The term tertiary structure refers to the unique 3-D conformations that globular protein assume as they fold into their native structures (biologically active). The α-helices and β-pleated sheets are folded into compact globule. Protein folding occurs as consequence of interactions between the side chains in their primary structure Tertiary structure has several important features: Many p.p fold in such fashion that a.a residues that are distant from each other in the primary structure come into close proximity Because of efficient packing as the p.p chain folds, globular protein are compact. Most water molecules are excluded from protein’s interior making interactions between both polar and non-polar groups possible. Large globular protein contain several compact units called domains. Domains: structural independent segments that have specific functions. Interactions that stabilize tertiary structure 1) Hydrophobic interactions As polypeptide folds, amino acids with hydrophobic (nonpolar) side chain are brought close to each other, out of contact with water. 2) Electrostatic interactions Interaction occurs between ionic groups of opposite charge (referred as salt bridge) 3) Hydrogen bonds Large number of hydrogen bond form within a protein’s interior and on its surface Examples of amino acid side chains that may hydrogen bond to each other: Two alcohols: ser, thr, and tyr. Alcohol and an acid: asp and tyr Two acids: asp and glu Alcohol and amine: ser and lys Alcohol and amide: ser and asn 4) Covalent bond Created by chemical reactions that alter a polypeptide's structure during or after its synthesis eg. Disulphide bond (strong linkage) Protect protein structure from adverse changes in pH or salt concentrations d) Quaternary structure Proteins consist of two or more polypeptide chains aggregated into one functional macromolecules Many proteins, esp those with high molecular weight are composed of several polypeptide chains. In proteins that consist of more than 1 polypeptide chain, each polypeptide is called subunit Polypeptide subunits assemble and held together by noncovalent interaction eg H bonding, hydrophobic effect, electrostatic interaction Loss of protein structure Considering the small differences in the free energy of folded and unfolded protein- protein structure is sensitive to environment factors. Many physical & chemical agents can disrupt protein’s native conformation The process of structure disruption = denaturation Denaturing conditions : Strong acid or base – changes in pH result in protonation of some protein side group, which alter/disrupt hydrogen bonding & salt bridge. As a protein approaches its isoelectric point, it becomes insoluble and precipitates from solution. Organic solvents – water-soluble organic solvents eg. Ethanol interfere with hydrophobic interaction because they interact with nonpolar R groups and form H bond with water and polar protein group. Detergents – these amphiphatic molecules disrupt hydrophobic interaction causing proteins to unfold into extended polypeptide chains (amphiphatic = contain nonpolar and polar components) Reducing agents – eg. Urea, βmercaptoethanol, will convert disulfide bridge (S-S) to sulfhydryl group (SH) urea disrupt H bond & hydrophobic interaction Heavy metal ions – mercury (Hg+) and lead (Pb2+) disrupt salt bridge by forming ionic bond with negatively charge group. Temperature change – as temp increase, the rate of molecular vibration increase. So weak H bond will be disrupt and protein will unfold. Mechanical stress – stirring & grinding actions disrupt the delicate balance of forces that maintain protein strcuture. eg. Foam formed when egg white is beaten vigorously contains denatured protein Fibrous proteins Fibrous protein exist as a long stranded molecules Contain high proportions of secondary structures ; α-helices and β-pleated sheets Most are structural protein eg. α-keratin, collagen, silk fibroin α-keratin Found in hair, wool, skin, horns, fingernails is an α-helical polypeptides Each polypeptide has three domain : - an amino terminal ‘head’ - a central rodlike α-helical domain - a carboxyl terminal ‘tail’ Two keratin polypeptides associate to form = coiled coil dimer Two antiparallel rows of these dimer form a supercoiled structure called a protofilament (H bonds & disulfide bond aid the formation of protofilament) Hundreds of filaments, each containing 4 protofilaments form macrofibril Each hair cells (fiber) contain several macrofibrils Physical properties of the α-keratins are reflected in their amino acids compositions. They have a regular α-helix structure because they lack of proline and have alanine and leucine. Because R groups are on the outside of the α-helices, the high hydrophobic a.a content makes it insoluble in water. Its cys residues and the formation of interhelix disulfide bridges make it resistant to stretching. Collagen The most abundant protein in vertebrates Synthesized by - connective tissue cells mostly found in fibrous tissues such as : tendon, ligament and skin, in cornea, cartilage, bone, blood vessels, the gut Extremely strong Collagen is composed of three lefthanded polypeptide helices that are twisted around each other to form a triple helix (stabilized by hydrogen bonding) The amino acid composition of collagen is distinctive - high content of glycine, proline and lysine - very little amount of cysteine (unlike αkeratin) Silk fibroin Silk protein - form spider webs, cocoon and nests - consist of the fibrous protein fibroin Considered to be β-keratin - polypeptide chains arranged in antiparallel βpleated sheet comformation Its primary structure mainly consists of the amino acid sequence (Gly-Ser-Gly-Ala-Gly-Ala)n Because the β-pleated sheets are loosely bonded to each other, they slide over each other easily. This arrangements gives silk fibers their flexibility. Globular protein Globular protein have a spherical shape, compact and water-soluble In their function, usually require them to bind precisely to other molecules Each protein has a unique and complex surface that contains cavities and clefts whose structure is complementary to specific ligands. After ligand binding, a conformational change occurs in the protein that is linked to a biochemical event. Most enzymes are globular Myoglobin & hemoglobin are typical example of globular protein Both are hemoprotein and each is involved in oxygen metabolism Unlike fibrous proteins which only play a structural function, globular proteins can act as: 1) Enzymes, by catalyzing organic reactions taking place in the organism in mild conditions and with a great specificity. 2) Messengers, by transmitting messages to regulate biological processes. This function is done by hormones, i.e. insulin etc. 3) Transporters of other molecules through membranes 4) Stocks of amino acids. 5) Regulatory roles are also performed by globular proteins rather than fibrous proteins. a) Myoglobin Myoglobin - oxygen-transport protein found in the muscle tissue of vertebrates in general and in almost all mammals. Diving mammals – high conc of myoglobin in the muscle tissue (brown in colour) has the ability to store oxygen by binding it to an iron atom (heme) It is found abundantly in the tissues of diving mammals, e.g., the whale, the seal, and the dolphin. High concentrations of myoglobin in these animals, allows them to store sufficient oxygen to remain underwater for long periods. Protein component = globin (single polypeptides chain (153 a.a) with 8 section of α-helix) Each myoglobin molecule contains one heme prosthetic group Each heme consist of porphyrin ring with Fe2+ in the center Free heme [Fe2+] has a high affinity for O2 and is irreversibly oxidized to form hematin [Fe3+] b) Hemoglobin Hemoglobin is a roughly spherical molecule found in red blood cells Function = transport oxygen from lung to every tissues in the body Composed - two α-chain - two β-chain The protein contain four subunits, designated α and β. Each subunit contain a heme group that bind with oxygen Protein technology: Purification of protein - - - After protein has been isolated, several methods can be used to enhance purification. Salting out Technique in which high concentrations of salts such as [(NH4)2SO4] are used to precipitate proteins. This technique removes impurities Dialysis is routinely used to remove lowmolecular weight impurities such as salts, solvents and detergents. Chromatography Chromatography techniques can be used to separate protein mixtures on the basis of molecular properties, such as size, shape and weight or binding affinities. In all chromatographic methods, protein mixture is dissolved in liquid (mobile phase). As protein molecules passed thru stationary phase (solid matrix), they separate because of different distributions between 2 phases. Gel-filtration chromatography A column packed with gelatenous polymer separates molecules according to size and shape. Molecules larger than gel pores are excluded and move faster thru the column. Molecules that are smaller than gel pores diffuse in and out of pores- their movement thru the column is retarded. The smaller the molecular weight, the slower they move. Differences in these rates separate protein mixture into bands, which are then collected separately. Ion-exchange chromatography Separates protein based on their charge. Anion-exchange resins- consist of +vely charged materials. Binds reversibly with protein’s negatively charged groups. Cation-exchange resins: bind positively charged groups. After proteins that do not bind to resin are removed, the protein of interest is recovered by appropriate change in solvent pH/salt conc. (a change in pH alters the charge). Affinity Chromatography Uses unique properties of proteins. Uses a special non-covalent binding affinity between protein and special molecule- ligand. Ligand is covalently bound to insoluble matrix which is placed in the column. After nonbinding protein molecules have passed thru the column, the protein of interest is removed by altering the conditions that affect binding (pH/salt conc.). Electrophoresis Proteins are electrically charged- they move in an electric field. In electrophoresis- molecules separate from each other because of differences in their net charge. Positive net charge migrate toward the –vely charged electrode and negative net charge migrate to +vely charged electrode. No charge=not moving. Electrophoresis using polyacrylamide/agarose gel. The gel separate proteins on the basis of their molecular weight and shape. During purification, specific bands may be excised from the gel after visualization. Each protein containing fragment is then eluted with buffer. Gel electrophoresis also used to assess the purity of protein samples. Staining the gel with dye Coomassie blue is commanly used to assess the success of a purfication step.