* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Detection of chromosome 2 and chromosome 7 within X-ray
Medical genetics wikipedia , lookup
Genealogical DNA test wikipedia , lookup
Saethre–Chotzen syndrome wikipedia , lookup
Human genome wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Point mutation wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
Genomic imprinting wikipedia , lookup
Designer baby wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
Genomic library wikipedia , lookup
Microevolution wikipedia , lookup
Gene expression programming wikipedia , lookup
Epigenetics of human development wikipedia , lookup
DNA supercoil wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Segmental Duplication on the Human Y Chromosome wikipedia , lookup
Hybrid (biology) wikipedia , lookup
Comparative genomic hybridization wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Genome (book) wikipedia , lookup
Skewed X-inactivation wikipedia , lookup
Y chromosome wikipedia , lookup
X-inactivation wikipedia , lookup
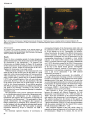
Mutagenesis vol.12 no.2 pp.55-59, 1997 Detection of chromosome 2 and chromosome 7 within X-rayor colchicine-induced micronuclei by fluorescence in situ hybridization K.Wuttke, CStreffer and W.-U.MUIler1 Institut fllr Medizinische Strahlenbiologie, Universitatsklinikum Essen, D45122 Essen, Germany 'To whom correspondence should be addressed The occurrence of chromosome 2 and chromosome 7 within micronuclei of binucleated lymphocytes induced by X-rays or colchicine was scored using the whole chromosome painting technique. The observed frequency of involvement in micronucleus formation was compared with the yield that would be expected theoretically, when the random participation of each chromosome is assumed. No difference was observed between the expected and observed inclusion of chromosome 2 or chromosome 7 into micronuclei after X-ray exposure (2.5 Gy). This was also the case for chromosome 2, but not for chromosome 7 after colchicine treatment (0.04/0.06 ug/ml); chromosome 7 was detected approximately 1.5 times more frequently in micronucleus formation than would be expected from the assumption of a random distribution. Introduction Genetic changes are central to the initiation and progression of neoplasias. A marker of chromosomal damage on the cytogenetic level is the formation of micronuclei: one possible fate of acentric fragments induced by clastogens as well as of lagging chromosomes induced by spindle poisons is micronucleation, which occurs after mitosis during nuclear membrane formation. The cytokinesis block method using cytochalasin B allows detection of cells which have undergone division, as binucleated cells, and micronuclei occurring in such cells can be used effectively to detect clastogenic or aneugenic effects (Mtlller and Streffer, 1994). Independent of the micronucleus-inducing agent, it is assumed that all chromosomes participate in aberrations in a random way depending on their DNA content. In general, this appears to be true. However, there are some indications that cytogenetic damage is not always distributed randomly, but that some chromosomes are involved more frequently than others (e.g. Brogger, 1977; Hayata and Dutrillaux, 1985; Tawn, 1988; Natarajan et ai, 1992; Hando et ai, 1994; Richard et ai, 1994; and additional literature cited in these papers). To test this possibility, we combined the cytokinesis block micTonucleus assay with the whole chromosome painting technique for chromosomes 2 and 7 of human lymphocytes after exposure to X-rays or to the spindle poison, colchicine. This allowed the simultaneous determination of the chromosome 2 and chromosome 7 status of micronuclei induced by clastogenic (X-rays) or aneuploidy-inducing (colchicine) agents. Materials and methods Cell culture and treatment Lymphocytes were isolated from the blood of a female donor by density centrifugation over Ficoll-Hypaque Qymphocyte separation medium; © UK Environmental Mutagen Society/Oxford University Press 1997 Fresenius, Bad Homburg, Germany). They were cultured at a density of 0.5X106 cells/ml in Roswell Park Memorial Institute (RPMI) 1640 medium (Gibco, Eggenstein, Germany) supplemented with 259b fetal calf serum and antibiotic/antimycotic solution (Gibco). The lymphocytes were stimulated by phytohaemagglutinin (PHA; Gibco). The cells were cultivated in multi-well tissue culture plates (Falcon) in an incubator (Heraeus, Osterode, Germany) at 37°C and 5% CO?. Depending on the experimental procedure, the lymphocytes were either irradiated with 2.5 Gy X-rays (240 kV, 0.5 mm copper filter, dose rate 1 Gy/ min) shortly before PHA stimulation or treated with colchicine (Sigma, Deisenhofen, Germany) 44 h after PHA stimulation. Due to inter-individual variation in the sensitivity of the cells to the spindle poison, the lymphocytes were exposed to different concentrations of colchicine (0.02, 0.04, 0.06 or 0.08 ug/ml). Fluorescence in situ hybridization was performed with those slides revealing the highest frequency of micronuclei (0.04/0.06 |ig/ml). Induction of binucleated cells Cytochalasin B (Sigma) was added to the culture at a final concentration of 5 |ig/ml 44 h after the initiation of the culture, and the cells were harvested 24 h later. Slide preparation Cells were transferred to precleaned slides using a cytocentrifuge (48 g, 5 min) and air-dried for 2 h. They were then fixed in cold methanol for 15 min and air-dried overnight. The slides were stored at -20°C till processed for hybridization. Whole chromosome painting by fluorescence in situ hybridization (FISH) FISH was performed using probes specific for whole chromosomes. The slides were hybridized simultaneously with Spectrum Green-conjugated DNA probes specific for chromosome 2 (Vysis-Company, Stuttgart-Fasanenhof, Germany) and Spectrum Orange-conjugated DNA probes specific for chromosome 7 (Vysis-Company). The fluorescent dye 4,6-diamidino-2-pbenylindole (DAPI; 150 ng/ml) was used to counterstain all chromosomes. In general, hybridization, washes and detection of bound probes were carried out according to the probe manufacturer's instructions, except for a few modifications. As the probes were developed for the detection of chromosomes in metaphase, pretreatment of the slides prior to denaturation was found to be necessary for the staining of chromosomes in interphase. Briefly, slides were pretreated by RNase (100 Ug/ml; Boehringer) for 1 h at 37°C, washed three times in 2X sodium chloride/sodium citrate (SSC) at room temperature, pretreated with pepsin (50 ilg/ml; Sigma) for 7 min at 37°C, washed twice in phosphate-buffered saline (PBS) and once in PBS + 50 mM MgCl2 at room temperature, fixed in 1% formaldehyde for 10 min at room temperature and washed in PBS at room temperature. Afterwards, the slides were dehydrated in ethanol (70, 90, 100, 100%; 3 min each) and air-dried. The DNA in the denaturation mix was denatured at 73°C for 5 min, and applied to slides that were denatured at 71°C for 5 min in 70% formamide/2x SSC (pH 7-8) and dehydrated in a 70, 85, 100% ethanol series. The slides were covered by a 22X22 mm coverslip and sealed with rubber cement Hybridization was performed at 37°C overnight in a moist chamber. The slides were air-dried after extensive washing of the slides in 50% fonnamide/2X SSC (pH 7-8) at 45°C for 30 min, followed by one wash in 2X SSC for 10 min and one wash for 5 min in 2X SSC + 0.1% MM0. Scoring procedure Microscopic analysis was performed on a Leica Diaplan fluorescence microscope that was connected to a laser scanning unit at a magnification of X1000. Both main nuclei and micronuclei had to be visible with DAPI to exclude artefacts. By changing the filter sets, the Spectrum Orange and Spectrum Green signals were clearly visible within the main nuclei. From time to time, chromosomes 2 and 7 were also stained in metaphase spreads to act as controls for the specificity of the method. At the start of die study, some signals in main nuclei and micronuclei were scanned to become familiar with the differentiation of artefacts. A total of 1000 X-ray-induced micronuclei and 1000 colchicine-induced micronuclei were analysed. These two groups of 1000 micronuclei were pooled from two blood samples obtained at different times from die same person; both samples showed the same result 55 K.Wuttke et al A D*t»«tl*n of mm* «T i n h u w n ttotlon of • ( r i M t n t of ohrMOtMt 2 within a «I or- anuo I «u* • ff • blnuolaatsd ly»phooyU Fig. 1. (A) Detection of chromosome 2 (green fluorescence) and chromosome 7 (red fluorescence) in a metaphase spread of a human lymphocyte. (B) Detection of a fragment of chromosome 2 within a micronucleus of a binuclcated lymphocyte (green fluorescence); theredfluorescenceindicates the location of chromosome 7. Statistical analysis As a measure of the statistical uncertainty of the observed frequency of involvement of the individual chromosomes in micronucleus formation, 95% confidence intervals were calculated (Sachs, 1984). Derivation of the expected values is explained in the Results section. Results Figure 1A shows a metaphase spread of a human lymphocyte painted simultaneously with chromosome-specific DNA probes for chromosome 2 and chromosome 7. As expected, four chromosomes are brightly labelled. In Figure IB, the painting of these chromosomes in the interphase nuclei of a binucleated lymphocyte is shown. Within the micronucleus of this cell, a fragment of chromosome 2 was made visible. Different fixation techniques for the binucleated cells were tested to find out the one that gave the best basis for a good signal after whole chromosome painting. Ah" fixation protocols which included the use of acetic acid led to a chromosome paint of minor quality, whereas the use of absolute methanol (-20°C) after cytocentrifugation of the cells onto slides gave good results. However, in this case, the attachment of the cells to the surface of the slides was not good enough, so that cells could be lost during denaturation. A post-fixation procedure after RNase and pepsin treatment (see Material and methods) was found to be necessary. According to this scheme, the hybridization led to good chromosome painting for interphase chromatin as well. The micronucleus assay and the subsequent fluorescence in situ hybridization protocol using whole chromosome painting allowed the determination of the composition of micronuclei with regard to the involvement of (fragments of) chromosome 2 and chromosome 7. The participation of these specific chromosomes was investigated after exposure of human lymphocytes to clastogenic X-ray irradiation (2.5 Gy) or after treatment with the aneuploidy-inducing spindle poison colchicine (0.04/0.06 (ig/ml). The total number of analysed micronuclei induced by X-rays or colchicine was 1000 in each case. The theoretical expected frequency of involvement in 56 micronucleus formation of the chromosomes under study can be calculated as follows: the chance of a target being damaged by X-rays depends on its size. Consequently, for radiationinduced micronuclei, the length of the different chromosomes must be considered. As the relative length of the individual chromosomes in metaphase may not be representative for the corresponding chromosomes in interphase, a more reliable indicator is the relative DNA content of the chromosomes which is correlated with their length. According to Mendelsohn et al. (1973), the relative content of the DNA of chromosomes 2 and chromosomes 7 is 8.0 and 5.3 respectively as a percentage of the total of the diploid female genome. Consequently, the theoretical frequency expected of involvement of these chromosomes in micronucleus composition is 80 for chromosome 2 and 53 for chromosome 7 per 1000 analysed micronuclei (see Figure 2). For cholchicine-induced micronuclei, the probability of the single chromosomes becoming involved in micronucleus formation is independent of their length because of the spindledamaging action of the agent. We also have to consider that we cannot distinguish between the two copies of each autosome. Consequently, the expected frequency of chromosome integration in this case is based on the assumption of a random involvement, 1000/23 = 44 (see Figure 2; for obvious reasons, no confidence interval can be given here). After exposure to X-rays, good agreement was found between the expected and observed frequencies for both chromosome 2 and chromosome 7 (P > 0.05) (see Figure 2). Table I shows the classification of the analysed micronuclei with regard to their chromosome 2 and chromosome 7 status. The quotient resulting from the division of the quantity of chromosome 2-positive micronuclei by the one of chromosome 7-positive micronuclei (70/49) is 1.43 and is in good accordance with the quotient calculated from the relative DNA contents of both chromosomes (8.00/5.30 = 1.51). After colchicine treatment of the cells, similar values for expected and observed frequencies of involvement were found for chromosome 2 (Figure 2). In contrast, this was not the Detection of chromosomes 2 and 7 in micronuclei using FISH Table I. Classification of the analysed micronuclei induced by X-rays or colchicine with regard to their chromosome 2 and chromosome 7 status Signal-negative micronuclei after Total no. analysed micronuclei after colcichine X-rays Chromosome 2 Chromosome 7 Ratio of chromosome 2/7 1000 1000 X-rays 1000 1000 930 951 colchicine 965 933 Signal-positive micronuclei after X-rays colchicine obsiexp. obs ./exp. 70/80 49/53 1.43/1.51 35/44 67/44 0.52/1.00 obs. = observed; exp. = expected (in the case of X-rays based on the relative DNA content of the chromosomes according to Mendelsohn er aL, 1973, in the case of colchicine based on the number of chromosomes). Table IL Classification of the signal-positive micronuclei with regard to their distribution per binucleated lymphocyte Chromosome 2 Chromosome 7 No. cases with 1 signal positive micronucleus X-rays colchicine X-rays colchicine X-rays colchicine 70 49 35 67 66 45 33 57 2 2 1 5 no. of signal-positive micronuclei per 1000 micronuclei X-rays 80 colchicine - HE- • 80 - - — 40 - -- 20 - — — n obs. axp. chromosome 2 2 signal positive micronuclei Total no. siginal-positive micronuclei after obs. exp. chromosome 7 obs. exp. obs. exp. chromosome 2 chromosome 7 Fig. 2. Comparison between the theoretically expected (exp.) and the experimentally observed (obs.) frequency of involvement of chromosome 2 and chromosome 7 in the formation of micronuclei induced by X-rays (2.5 Gy) or colchicine (0.04/0.06 Hg/ml). (Bars indicate the 95% confidence intervals). case for chromosome 7; comparing the frequency of observed (67) and expected (44) chromosome 7-positive micronuclei, a significant over-involvement of this chromosome in micronucleus formation described by a factor of 1.5 was detected (Figure 2). The ratio obtained from the signal-positive micronucleus values for chromosome 2 and chromosome 7 after colchicine treatment (35/67 = 0.52) (Table I) deviates strongly from the expected value of 1 which should be found if both chromosomes had been involved in micronucleus formation to the same degree. Table II shows the classification of only the signal-positive micronuclei with regard to their distribution per binucleated lymphocyte. The most frequent finding was that only one chromosome 2- or one chromosome 7-positive micronucleus per binucleated lymphocyte was observed. After X-ray exposure, two cases were found with two signal-positive micronuclei per binucleated lymphocyte for chromosome 2 and chromo- some 7 respectively. Also after colchicine treatment, one binucleated cell was observed with two signal-positive micronuclei for chromosome 2 and, interestingly, even five binucleated cells with two chromosome 7-positive micronuclei. Discussion In this study, we compared the frequency found experimentally of involvement of chromosome 2 and chromosome 7 in micronucleus formation with the one that would be expected theoretically, when a participation of each individual chromosome on the basis of randomness is assumed. As micronucleiinducing agents, we used the clastogen X-ray irradiation (2.5 Gy) and the aneuploidy-inducing spindle poison colchicine (0.04/0.06 ng/ml). We have specifically focused on chromosome 2 and chromosome 7 because: (i) the non-random occurrence of exchanges involving chromosome 7 in healthy control individuals (Tawn, 1988) hints at a special genetic fragility of this chromosome; (ii) the non-random chromosomal abnormalities of chromosome 7 have been shown to be of interest in the development of a number of different malignancies such as malignant gliomas, neuroblastoma, prostate cancer, transformed lymphoblasts and transformed liver epithelial cells (e.g. Steel et aL, 1980; Bigner et aL, 1986, 1987; Rey et aL, 1987; Cremer et aL, 1988; Llombart-Bosch et aL, 1989; Takahashi et aL, 1994; Steadman et aL, 1994). Steadman et aL (1994) showed that in transformed rat liver epithelial cells the non-random occurrence of the chromosomal abnormality involving chromosome 7 is located near the proto-oncogene c-myc; (iii) chromosome 2 is representative of the large sized chromosomes and thus is expected to be involved more frequently in damaging events than the shorter chromosomes. The results obtained after X-ray exposure clearly show that neither of the two analysed chromosomes is included preferentially into micronuclei, but that their participation in the expression of cytogenetic damage is merely random (Figure 2). The correlation between the relative DNA content (and thus the relative length) of the chromosomes and the probability of the chromosomes being damaged by X-rays is obvious (Table I). As X-rays are known to damage chromosomes 57 K-Wuttke et al mainly by inducing chromosome breakage resulting in two acentric fragments (e.g. Evans, 1988; WeiBenborn and Streffer, 1991), the occurrence of two micronuclei with a signal from the same chromosomes (Table II) within a binucleated lymphocyte is not surprising; their packing into different micronuclei can be expected because of the spatial location of the fragments belonging to the same damaged chromosome. The targets of the damaging action of colchicine are the microtubules which form the mitotic spindle. By this mechanism, lagging chromosomes appear during anaphase; these cannot be distributed into the daughter-cells and so the incorporation of these chromosomes into micronuclei during telophase is the result. Hence, micronuclei induced by colchicine are expected to be composed of whole chromosomes. The colchicine concentration chosen in this study (0.02-0.08 |i.g/ml) does not lead to a total disruption of the spindle; consequently, the cleavage of the cells was still possible and induced micronuclei could be expressed. Based on knowledge of the effects of colchicine, one would expect that a damaged spindle fibre would probably affect all chromosomes with equal frequency. However, the results of this study are not in accordance with this assumption; while the observed involvement of chromosome 2 in micronucleus formation corresponds with the theoretical frequency, we found a significant over-involvement of chromosome 7 (Figure 2, Table I). Why should chromosome 7 fail to align on the remaining spindle more frequently than other chromosomes? At present, we are not able to give a satisfactory explanation. Instead, we have developed three hypothetical explanations: (i) it is known that different parts of the cytoskeleton are linked together and that their functions are co-ordinated (for review see Alberts et al, 1983). There is evidence that cytoplasmic microtubules co-ordinate the various parts of the cytoskeleton responsible for complex intracelluar movements. They influence the distribution of actin filaments as well as of intermediate filaments, which play a structural and tensionbearing role in the cell. The principal filaments of the cytoskeleton are extensively interconnected by a three-dimensional network as well as by soluble, diffusing factors. One component of the nuclear matrix, in which the chromosomes are postulated to be anchored, is the non-chromatin network, a three-dimensional system of filaments with associated granular structures (Nickerson et al, 1987). There is no evidence, but it is imaginable, that these two filament systems interact (especially during mitosis when the nuclear envelope is disappearing and the spindle enters the nuclear area), so that the colchicineinduced disruption of the microtubules might affect the network of the nuclear matrix. Depending upon the anchorage location of chromosome 7 within the matrix, this chromosome may be predisposed for a failure in aligning on the spindle; (ii) the relative number of inserting microtubules per kinetochore of the different chromosomes has not been investigated up to now. If each chromosome is not attached by the same amount of spindle fibres, one explanation for our finding with chromosome 7 could be that this chromosome might contain a relatively low number of microtubules, so that the probability of this chromosome becoming a laggard as a result of colchicineinduced disruption of the spindle is relatively high; (iii) it may be possible that chromosome 7 has a number of nucleotide sequences similar to sequences within other chromosomes, so that this chromosome is especially prone to hybridize with areas of other chromosomes. With this mechanism, its chance 58 of aligning on a spindle with reduced numbers of microtubules might be modified. We are aware that our attempts to explain the principal finding of the over-involvement of chromosome 7 in micronucleus formation are speculative. Further investigations are in progress to clarify whether such a non-random participation in the expression of cytogenetic damage can also be found for other chromosomes, maybe for those with similar size as chromosome 7. The results of Ntisse et al (1992a,b) confirm our observations for chromosome 2 and chromosome 7 generally; the DNA distribution of radiation-induced micronuclei measured by flow cytometry did not show any peaks, leading to the conclusion that the composition of radiation-induced micronuclei can be explained by random breakage of chromosomes. In contrast, by using the spindle poison 2-chlorobenzylidene malonitrile, the authors described a DNA distribution of the induced micronuclei with several peaks, suggesting a correspondence with the DNA distribution of specific chromosomes which unfortunately have not been defined precisely. Looking at the classification of the signal-positive micronuclei with regard to their distribution per binucleated cell after colchicine-treatment (Table II), the occurrence of two signal-positive micronuclei per binucleated lymphocyte for chromosome 2 and chromosome 7 respectively, is surprising; after colchicine treatment, each micronucleus should be composed of entire chromosomes. For chromosome 7 the highest number of binucleates with two micronuclei per cell containing this chromosome was found. It can be excluded that each of the two micronuclei enclosed one of the homologues of chromosome 7, for the corresponding signal was also detected in the main nucleus. If the chromatids of the lagging chromosomes began to separate, one would then see two distinct signals that could have been included into different micronuclei. Another explanation can be derived from the similar results found by other investigators; according to Sternes and Vig (1989), there appears to be a loose correlation between the formation of acentric fragments and kinetochore-negative micronuclei after colchicine treatment. It is also possible that some of the kinetochore-negative micronuclei actually represent the whole chromosome-micronuclei, in which the antigen for the antikinetochore-antibody is altered, so that the kinetochore fluorescence is not detectable. However, studies using in situ hybridization for centromere identification showed only ~50-70% centromere-positive micronuclei (Adler, 1993; Hayashi et al, 1994). Thus, micronuclei induced by colchicine arose predominantly, but not exclusively, by whole lagging chromosomes. Whether or not the remaining fraction of micronuclei is due to chromosome breakage remains to be determined. In conclusion, our results (referring to chromosome 2 and chromosome 7) and those cited from the literature (referring to the entire genome of a cell) indicate a quite unexpected trend in the answer to our initial question about the comparative influence of clastogenic and aneuploidy-inducing agents on the participation of individual chromosomes in the expression of micronuclei. Whereas after exposure to ionizing radiation, chromosome-specific damage would be imaginable, and after treatment with a spindle poison an effect with equal frequency on the individual chromosomes would be expected, quite the opposite must be postulated now. However, further thorough studies on specific chromosomes and an investigation of Detection of chromosomes 2 and 7 In micronuclei using FISH possible spindle poison mediated damaging mechanisms on individual chromosomes are needed. References AdlerJ.-D. (1993) Synopsis of the in vivo results obtained with the 10 known or suspected aneugens tested in the CEC collaborative study. Mutat. Res., 287, 131-137. Alberts.B., BrayJD., LewisJ., Raff,M., RobertsJC. and WatsonJ.D. (eds) (1983) The Cytoskeleton. In Molecular Biology of the Cell Garland Publishing, New York. Bigner.S.H., MarkJ- Bullard,D.E., MahaleyJVl.S.Jr and Bigner.D.D. (1986) Chromosomal evolution in malignant human gliomas starts with specific and usually numerical deviations. Cancer Genet. Cytogenet., 22, 121-135. Bigner.S.H., WongAJ., MarkJ., Muhlbaier,L.H., Kinzler,K.W., Vbgelstein,B. and BignerJ).D. (1987) Relationship between gene amplification and chromosomal deviations in malignant gliomas. Cancer Genet. Cytogenet., 29, 165-170. Brogger,A- (1977) Non-random localization of chromosome damage in human cells and targets for clastogenic action. In de la Chapelle,A. and Sorsa>I. (eds), Chromosomes Today. Elsevier, New York, pp. 297-306. Cremer.T., Lichter.P., BordenJ., WardJD.C. and Manuelidis,L. (1988) Detection of chromosome aberrations in metaphase and interphase tumor cells by in situ hybridization using chromosome-specific library probes. Hum. Genet, 80, 235-246. Evans,H.J. (1988) Mutation cytogenetics: past, present and future. Mutat. Res., 204, 355-363. HandoJ.C, NathJ. and TuckerJ.D. (1994) Sex chromosomes, micronuclei and aging in women. Chromosoma, 103, 186-192. Hayashi,M., Maki-PaakkanenJ., Tanabe.H., Honmajvl, Suzuki.T., Matsuoka,A., Mizusawaji- and Sofuni.T. (1994) Isolation of micronuclei from mouse blood and fluorescence in situ hybridization with a mouse centromeric DNA probe. Mutat. Res., 307, 245-251. HayataJ. and Dutrillaux.B. (1985) Non-random involvement of rat and mouse chromosomes in the stable type chromosomal rearrangements observed in irradiated bone marrow cells. Proc. Jap. Acad., 61, 179—182. LJombart-Bosch.A., Carda,C, Peydro-Olaya, A., Noguei^R., BoixJ., and PellinA- (1989) Pigmented esthesioneuroblastoma showing dual differentiation following transplantation in nude mice. Virchows Archiv A. Pathol. Anat., 414, 199-208. Mendelsohn.M.L., Mayall,B.H., Bogart,E., Moore O.D.H. and Perry3.H. (1973) DNA content and DNA-based centromeric index of the 24 human chromosomes. Science, 179, 1126-1129. MUller.W.-U. and Streffer.C. (1994) Micronucleus Assays. Adv. Mutagen. Res., 5, 1-134. Natarajan,A.T., Darroudi,F., Vermeulen,S. and WiegantJ. (1992) Frequencies of X-ray and fast neutron induced chromosome translocations in human peripheral blood lymphocytes as detected by in situ hybridization using chromosome specific DNA libraries. In Sugahara,T., Sagan,L.A. and Aoyama,T. (eds), Low Dose Irradiation and Biological Defense Mechanisms. Elsevier Science Publishers, pp. 343-346. NickersonJ.A., He,D., Fey,G. and Penman.S. (1987) In Strauss,P.R. and Wilson,S.H. (eds), The Eukaryotic Nucleus. Molecular Biochemistry and Macromolecular Assemblies. Vol. 2. Telford Press, Caldwell, pp. 763-782. Nusse,M., KramerJ. and Miller.B.M. (1992a) Factors influencing the DNA content of radiation-induced micronuclei. Int. J. Radial. Biol, 62, 587-602. NUsse,M., Recknagel.S. and Beisker.W. (1992b) Micronuclei induced by 2chlorobenzylidene malonitrile contain single chromosomes as demonstrated by the combined use of flow cytometry and immunofluorescent staining with anti-kinetochore antibodies. Mutagenesis, 7, 57-67. ReyJ.A., BelloJ.M., de CamposJ.M., Kusak^.E. and Moreno.S. (1987) On trisomy of chromosome 7 in human gliomas. Cancer Genet. Cytogenet., 29, 323-326. RichardJ3., Mulerisjvl. and Dutrillaux,B. (1994) The frequency of micronuclei with X chromosome increases with age in human females. Mutat. Res., 316, 1-7. Sachs,L. (1984) Angewandte Statistik, 6th edn. Springer, Berlin. Steel.C.M., Shade,M. and Woodward,M.A. (1980) Chromosome aberrations acquired in vitro by human B-cell lines I. Gains and losses of material. J. Natl. Cancer Inst., 64, 95-98. Sternes,K.I. and Vig3-K. (1989) Micronuclei, kinetochores and hypoploidy. Mutagenesis, 4, 425-431. SteadmanJ.S., Lee,L.W., Smith.GJ. and GrishamJ.W. (1994) DNA contents and chromosomes of clonal lines of transformed rat liver epithelial cells and of cells from their derived tumors. Carcinogenesis, 15, 963-969. Takahashi.S., QianJ., BrownJ.A., AlcarazA, BostwickJXG., Lieber,M.M and Jenkins, R.B. (1994) Potential markers of prostate cancer aggressiveness detected by fluorescence in situ hybridization in needle biopsies. Cancer Res., 54, 3574-3579. TawnJ. (1988) The non-random occurrence of exchanges involving chromosomes 7 and 14 in human lymphocytes: a prospective study of control individuals. Mutat. Res., 199, 215-220. WeiBenborn.U. and Streffer.C. (1991) Micronuclei with kinetochores in human melanoma cells and rectal carcinomas. Int. J. Radial. Biol, 59, 373-383. Received on June 24, 1996; accepted on October 23, 1996 59