* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Assignment 5 Bioenergy/ Photosynthesis
Ultrasensitivity wikipedia , lookup
Western blot wikipedia , lookup
Microbial metabolism wikipedia , lookup
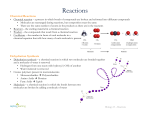
Metabolic network modelling wikipedia , lookup
Nicotinamide adenine dinucleotide wikipedia , lookup
Electron transport chain wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Magnesium in biology wikipedia , lookup
Multi-state modeling of biomolecules wikipedia , lookup
Catalytic triad wikipedia , lookup
NADH:ubiquinone oxidoreductase (H+-translocating) wikipedia , lookup
Citric acid cycle wikipedia , lookup
Adenosine triphosphate wikipedia , lookup
Biosynthesis wikipedia , lookup
Metalloprotein wikipedia , lookup
Biochemistry wikipedia , lookup
Photosynthesis wikipedia , lookup
Enzyme inhibitor wikipedia , lookup
Light-dependent reactions wikipedia , lookup
Oxidative phosphorylation wikipedia , lookup
Evolution of metal ions in biological systems wikipedia , lookup
Name Sec. . Score . Working in groups of two to four and armed with a textbook or some other reliable source, answer the following questions concerning enzyme function and photosynthesis (Chapters 5 & 6). This assignment is worth 30 points with the possible points for each question in parenthesis. The following sites may help you answer and understand the questions. http://www.biology-pages.info/E/Enzymes.html http://www.microtack.com/html/enzyme1.htm http://www.ucl.ac.uk/~ucbcdab/enzass/inhibition.htm 1. (5) Define the following terms that relate to an enzyme: activation energy, active site, conformation, denaturation, and coenzyme/cofactor binding site? Enzymes are proteins and therefore must be folded or conformed into a specific shape to be functional. Their function is to assist the occurrence of the necessary biological reactions within the parameters (limits) (pH, O2 concentrations, CO2 concentrations, temperature, etc.) of normal biological function. Enzyme folding or conformation includes the opening of a portion of the protein called the active site. This site is designed to bind to reactant molecules called substrate. Once the reactants are bound, forming an enzyme-substrate complex (or E-S C) via usually hydrogen bonds or ionic interactions, the enzyme can change its conformation to move the reactants into a position to react (e.g. split the reactant apart, put two reactants together, or change the reactivity or shape of the reactant). The amount of energy required for the reaction to proceed without an enzyme is called the activation energy and enzymes lower this amount of energy by being able to bind to the substrate and then change their conformation to force the reaction. Denaturation of a protein is a process by which some factors, like temperature or pH, affect the shape of a protein by breaking or disrupting the bonds (H-bonds, R-R interactions, or ionic interactions) of the protein. In the case of an enzyme, denaturation changes the shape of the active site. After denaturation the enzyme cannot react with the substrate and the reaction will not proceed. Another binding site on the enzyme located at a distance from the active site, called the cofactor/coenzyme site, binds either to a vitamin or a mineral. Once the vitamin or mineral is bound the active site shape is altered, either to open the site or to close the site. In this way vitamins and minerals are necessary for regulating normal enzyme function. 2. (5) How are enzymes inhibited by competing molecules and by non-competing molecules? If another molecule has a similar enough shape to the substrate it can partially bind to the active site of an enzyme and thereby compete with the actual substrate for binding to the active site. This is called competing (competitive) inhibition of an enzyme. At this point it comes down to concentration or a numbers game. If there are more competitors than substrate then the enzyme will be blocked from binding to the substrate but if more substrate is available then the reaction will proceed because the enzyme “sees” the substrate more often than the competitor (This is just random bumping into each other not a seek and destroy mission.). This problem can usually be rectified by providing an increase in the amount or concentration of the normal substrate molecule and increase the likelihood of the enzyme bumping into substrate rather than the inhibitor. If another molecule binds to the cofactor/coenzyme site then the situation is exceedingly dire to normal enzyme and organism function. This is called non-competing (non-competitive) Biology& 100 Mr. Brumbaugh 1 Class Assignment 5 inhibition. When something binds to the cofactor/coenzyme site it can bind tighter than the usual cofactor/coenzyme. This action blocks this regulatory site and shuts down the enzyme by altering the enzymes conformation and thus the active site so that the substrate can’t bind to the active site at all. Since it doesn’t deal with concentration of substrate then the only way to correct this situation is to destroy the enzyme and build a new enzyme and eliminating the non-competitive inhibitor. This takes time and could result in the death of the organism depending on how many of the enzymes for that reaction are affected. Some poisons have this effect and this is why they must be eliminated or neutralized as quickly as possible from the body before the enzymes of cells are affected. 3. (5) Draw a graph that would represent enzyme function at various temperatures and various pH levels? Explain why the graphs give the bell shaped appearance? Enzymes function best in an optimal pH or temperature environment causing the graph of activity to take on a bell shaped curve. The shape of the curve relates to the degree of denaturation of the enzyme caused by the environmental change. 4. (5) Describe the concept of energy coupling reactions in regards to ATP usage? Adenosine Tri-Phosphate (ATP) coupling reactions involve combining one reaction say A + B that is energetically unfavorable at normal biological parameters with a reaction that will add some energy to the reaction. To accomplish this reaction an ATP molecule donates a P (represents energy) to one reactant which energizes this reactant, we will call A, to react with another reactant called B forming a new molecule called AB. Once the reaction has taken place the P is released and the product AB is formed. The P (often written as Pi) can then be recombined with an Adenosine Di-Phosphate (ADP) to make another ATP with the addition of more energy. The AB that was produced can be used however the cell wants. These web sites may be helpful to your understanding of photosynthesis. http://www.ftexploring.com/me/photosyn1.html https://www.khanacademy.org/science/biology/photosynthesis-in-plants/introduction-to-stages-ofphotosynthesis/v/photosynthesis Biology& 100 Mr. Brumbaugh 2 Class Assignment 5 5. (5) Describe the light dependent reactions of photosynthesis in five steps or less. A. Sunlight strikes two separate chlorophyll based photosynthetic pigment systems (remarkably called Photosystem I (PSI) and Photosystem II (PSII)) which excite two electrons within a core magnesium atom of each pigment system (labelled P680 and P700 in the figure below the numbers refer to the wavelength of light that best stimulates electron excitation). The pigment chlorophyll houses the magnesium in both systems. B. The excited electrons of each system are passed through an Electron Transport Chain (ETC) associated with the different photosystems. The first chain associated with PSII generates an ATP molecule as the electrons are passed down the chain and the electrons are passed to the second pigment system (PSI). A second electron chain associated with PSI reduces a Nicotinamide Adenine Dinucleotide Phosphate (NADP+) molecule by giving the electrons from the second pigment system to NADP+ to form NADPH + H+. Where did the H+ come from to complete this transfer? C. The hydrogen that associates with the NADP+ comes from the splitting of a water molecule to replace the electrons lost from the first pigment system (PSII). Remember PSII gave its electrons to the second pigment system (PSI). The H+ ion moves across the thylakoid membrane where the cytochrome complexes are embedded to cause a build-up of potential energy. This gradient energy is allowed to dissipate back across the membrane via a protein complex called ATP Synthase (or I can make ATP doggoneitase) to yield an ATP molecule from ADP and Pi. D. The oxygen from the water is given off as waste to either be used by plant cells mitochondria in cellular respiration or to the atmosphere via leaf stomata. These reactions occur in pigments called chlorophyll embedded into the thylakoid membrane of a chloroplast’s grana. How does the oxygen get out of the plant? It can either be used by the plant cells mitochondria to act as the final electron acceptor in cell respiration or released out of the leaf through the stomata. Biology& 100 Mr. Brumbaugh 3 Class Assignment 5 6. (5) Describe the light independent reactions (Calvin cycle or dark reactions) of photosynthesis) in five steps or less. Divide the Calvin cycle into two halves: Molecule building and Rearrangement. A. An enzyme named “Rubisco” (see figure below) combines 3CO2 molecules from the atmosphere (via the leaf stomata) with 3 molecules of RuBP (Ribulose BiPhosphate) that is already inside the stroma of a chloroplast to form six molecules called 3-PGA or 3phosphoglycerate (carbon fixation). B. 12ATP’s from the light reactions are used to energize (open or break) the CO2’s. 2 ATP per each CO2 because two bonds need to be broken. C. This opens the CO to accept the H+ from 6NADPH + H+. These steps generate 6 molecules of Glyceraldehyde 3-Phosphate (6G3P) (Molecule building). D. The G3P’s are separated and one of the G3P’s is used to build other molecules like glucose, lipids, amino acids or anything the plant cell needs, while the rest (5G3P’s) are rearranged (Rearrangement) back into the 3 RuBP’ s that is needed to start the cycle again. The rearrangement steps cost 3 more ATP’s These reactions occur via enzymes embedded into the chloroplast’s stroma. What limits the rate of these reactions? The availability of CO2, RuBP, ATP, NADH + H+ (from the light dependent reactions), and rubisco activity would limit the rate of reaction by the Calvin cycle enzymes. Biology& 100 Mr. Brumbaugh 4 Class Assignment 5