* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
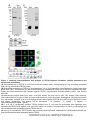
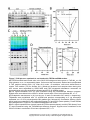
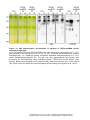
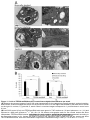
Download TEF30 interacts with photosystem II monomers and is involved in the
Survey
Document related concepts
Hedgehog signaling pathway wikipedia , lookup
Cellular differentiation wikipedia , lookup
Protein moonlighting wikipedia , lookup
Cell culture wikipedia , lookup
Organ-on-a-chip wikipedia , lookup
Extracellular matrix wikipedia , lookup
Protein phosphorylation wikipedia , lookup
Cytokinesis wikipedia , lookup
Cell encapsulation wikipedia , lookup
Signal transduction wikipedia , lookup
Endomembrane system wikipedia , lookup
Western blot wikipedia , lookup
Transcript
Plant Physiology Preview. Published on December 7, 2015, as DOI:10.1104/pp.15.01458 1 Running head: TEF30 is involved in PSII repair 2 3 4 Michael Schroda 5 Molekulare Biotechnologie & Systembiologie 6 TU Kaiserslautern 7 Erwin-Schrödinger Straße 70, D-67663 Kaiserslautern 8 Germany 9 Tel.: ++49 631 205 2697 10 Fax: ++49 631 2015 2699 11 e-mail: [email protected] 12 13 14 Most appropriate journal research area: Cell Biology 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 1 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. Copyright 2015 by the American Society of Plant Biologists 31 TEF30 interacts with photosystem II monomers and is involved in the repair of 32 photodamaged photosystem II in Chlamydomonas reinhardtii 33 34 Ligia Segatto Muranaka*1, Mark Rütgers*1, Sandrine Bujaldon2, Anja Heublein3, Stefan 35 Geimer3, Francis-André Wollman2, and Michael Schroda1 36 37 1 38 70, D-67663 Kaiserslautern, Germany 39 2 40 Biologie Physico-Chimique, UMR CNRS/UPMC 7141, Paris, France 41 3 Molekulare Biotechnologie & Systembiologie, TU Kaiserslautern, Erwin-Schrödinger Laboratoire de Physiologie Membranaire et Moléculaire du Chloroplaste, Institut de Zellbiologie/Elektronenmikroskopie, Universität Bayreuth, D-95440 Bayreuth, Germany 42 43 * Equal contribution 44 45 One-sentence Summary: 46 A protein conserved in the green lineage binds quantitatively to photosystem II 47 monomers at the stromal side of thylakoid membranes and facilitates supercomplex 48 assembly during photosystem II repair. 49 50 51 52 53 54 55 56 57 2 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. 58 Footnotes: 59 This work was supported by the Deutsche Forschungsgemeinschaft [Schr 617/9-1], the 60 DAAD/CNPq [290020/2010-7], and the Forschungsschwerpunkt BioComp. 61 62 Corresponding author: 63 Michael Schroda 64 e-mail: [email protected] 65 66 List of author contributions: 67 L.S.M. and M.R. performed most of the experiments and were supervised by M.S; S.B. 68 contributed to the biophysical experiments, which were supervised by F.A.W. A.H. and 69 S.G. performed the electron microscopy work; all authors contributed to the design of 70 the experiments and data analysis; M.S. conceived the project and wrote the article with 71 contributions of all the authors. 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 3 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. 89 Abstract 90 The remarkable capability of photosystem (PS) II to oxidize water comes along with its 91 vulnerability to oxidative damage. Accordingly, organisms harboring PSII have 92 developed strategies to protect PSII from oxidative damage and to repair damaged PSII. 93 Here we report on the characterization of the TEF30 protein in Chlamydomonas 94 reinhardtii that is conserved in the green lineage and induced by high light. Fractionation 95 studies revealed that TEF30 is associated with the stromal side of thylakoid membranes. 96 By using blue native / Deriphat-PAGE, sucrose density gradients, and isolated PSII 97 particles, we found TEF30 to quantitatively interact with monomeric PSII complexes. 98 Electron microscopy images revealed significantly reduced thylakoid membrane stacking 99 in TEF30-underexpressing cells when compared with control cells. Biophysical and 100 immunological data point to an impaired PSII repair cycle in TEF30-underexpressing 101 cells and a reduced ability to form PSII supercomplexes after high light exposure. Taken 102 together, our data suggest potential roles for TEF30 in facilitating the incorporation of a 103 new D1 protein and/or the reintegration of CP43 into repaired PSII monomers, protecting 104 repaired PSII monomers from undergoing repeated repair cycles, or facilitating the 105 migration of repaired PSII monomers back to stacked regions for supercomplex 106 reassembly. 107 4 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. 108 Introduction 109 Oxygenic photosynthesis is essential for almost all life on earth as it provides the 110 reduced carbon and the oxygen required for respiration. A key enzyme in oxygenic 111 photosynthesis is photosystem (PS) II, which catalyzes the light-driven oxidation of 112 water. The core of PSII in algae and land plants contains D1 (PsbA), D2 (PsbD), CP43 113 (PsbC), CP47 (PsbB), the α- (PsbE) and β-subunits (PsbF) of cytochrome b559 as well 114 as several intrinsic low molecular-mass subunits. The core monomer is associated with 115 the extrinsic oxygen evolving complex (OEC) consisting of OEE1 (PSBO), OEE2 116 (PSBP), and OEE3 (PSBQ) which stabilize the inorganic Mn4O5Ca cluster required for 117 water oxidation (reviewed in Pagliano et al., 2013). PSII core monomers assemble into 118 dimers to which at both sides light-harvesting proteins (LHCII) bind to form PSII 119 supercomplexes. In land plants, each PSII dimer binds two each of the monomeric minor 120 antenna proteins CP24, CP26, and CP29 in addition to up to four trimeric major antenna 121 (Caffarri et al., 2009; Kouril et al., 2011). Biochemical evidence suggests that in the 122 thylakoid membrane up to eight LHCII trimers can be present per PSII core dimer, 123 presumably because the existence of a pool of “extra” LHCII (Kouril et al., 2013). In 124 Chlamydomonas reinhardtii, lacking CP24, PSII dimers bind two each of CP26 and 125 CP29 monomers as well as up to six trimeric major antenna (Tokutsu et al., 2012). The 126 reaction center proteins D1 and D2 bind all the redox-active cofactors required for PSII 127 electron transport (Umena et al., 2011). Light captured by the internal antenna proteins 128 CP43 and CP47 and the outer antenna induces charge separation in PSII, which in turn 129 enables the OEC to oxidize water and provide electrons to the electron transfer chain. In 130 land plants and green algae PSII supercomplexes are localized to stacked regions of the 131 thylakoid membranes, while the synthesis of PSII cores is considered to take place in 132 stroma lamellae. 133 A particular feature of PSII is its vulnerability to light, with the D1 protein being a 134 target of light-induced damage and damage being proportional to the photon flux density 135 (PFD) applied (Tyystjarvi and Aro, 1996). To cope with this damage, an elaborate highly 136 conserved mechanism has evolved termed the PSII repair cycle during which damaged 137 PSII complexes are partially disassembled and the defective D1 protein is replaced by a 138 de novo synthesized copy (for recent reviews see Nixon et al., 2010; Komenda et al., 139 2012; Mulo et al., 2012; Nath et al., 2013a; Nickelsen and Rengstl, 2013; Tyystjarvi, 5 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. 140 2013; Jarvi et al., 2015). Photodamage occurs at all light intensities, but when the rate of 141 damage exceeds the capacity for repair, photoinhibition is manifested as a decrease in 142 photosynthetic activity and in the proportion of active PSII reaction centers (Aro et al., 143 1993). While PSII photodamage occurs in the supercomplexes in the stacked membrane 144 regions, the replacement of damaged D1 takes place in stroma lamellae (Aro et al., 145 2005). Thus, the PSII repair cycle requires lateral migration of PSII complexes, which is 146 impaired by the macromolecular crowding in stacked thylakoid membranes (Kirchhoff, 147 2013). Lateral migration of damaged PSII complexes is facilitated by thylakoid 148 membrane unfolding and PSII supercomplex disassembly. Both processes are 149 enhanced by the phosphorylation of PSII core subunits D1, D2, CP43, and PsbH, which 150 is mainly mediated by the protein kinase STN8 (Tikkanen et al., 2008; Fristedt et al., 151 2009; Herbstova et al., 2012; Nath et al., 2013b; Wunder et al., 2013). Efficient PSII 152 supercomplex disassembly also requires the THF1/NYC4/Psb29 protein (Huang et al., 153 2013; Yamatani et al., 2013). After migration of PSII monomers to unstacked thylakoid 154 regions, core subunits are dephosphorylated by the PSII core phosphatase PBCP 155 (Samol et al., 2012), which is required for the efficient degradation of D1 (Koivuniemi et 156 al., 1995; Rintamaki et al., 1996; Kato and Sakamoto, 2014). Degradation of D1 is 157 subsequently realized by the membrane-integral FtsH protease (Lindahl et al., 2000; 158 Silva et al., 2003) and by lumenal and stromal Deg proteases (Haussuhl et al., 2001; 159 Kapri-Pardes et al., 2007; Sun et al., 2010). Degradation is assisted by the lumenal 160 protein TLP18.3, presumably by its phosphatase activity and ability to interact with 161 lumenal Deg1 (Sirpio et al., 2007; Wu et al., 2011; Zienkiewicz et al., 2012). D1 162 proteolysis follows the partial disassembly of the PSII complex during which CP43 and 163 low molecular-mass subunits are released to generate a CP43-free PSII monomer (Aro 164 et al., 2005). Thereafter, a newly synthesized D1 copy is co-translationally inserted from 165 a plastidial 70S ribosome into the thylakoid membrane and processed by the CtpA 166 peptidase (Zhang et al., 1999, 2000; Che et al., 2013). In Arabidopsis thaliana, the D1 167 synthesis rate appears to be negatively regulated by the PDI6 protein (Wittenberg et al., 168 2014). Moreover, yet unknown steps during PSII repair require the stromal cyclophilin 169 ROC4 and stromal HSP70 (Schroda et al., 1999; Yokthongwattana et al., 2001; Cai et 170 al., 2008). The PSII repair cycle is completed by the reassembly of the CP43 protein, 171 ligation of the OEC, back-migration of PSII to stacked membrane regions and 6 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. 172 supercomplex formation. Except for CtpA all mentioned factors appear to be specific for 173 PSII repair, while many more auxiliary factors play roles in PSII de novo synthesis and 174 repair (reviewed in Jarvi et al., 2015). 175 In this study we report on the functional characterization of the TEF30 protein (for 176 thylakoid enriched fraction 30) in Chlamydomonas. In this organism, TEF30 was first 177 identified in a proteomics study on isolated thylakoid membranes (Allmer et al., 2006). It 178 attracted our attention because its abundance increased 1.7-fold in membrane-enriched 179 fractions of Chlamydomonas cells that had been shifted from 41 to 145 µmol photons m- 180 2 181 Arabidopsis (MET1) has been functionally characterized only recently (Bhuiyan et al., 182 2015). Both MET1 and TEF30 interact quantitatively with monomeric PSII core particles 183 at the stroma side of the thylakoid membranes and play a role in the assembly of PSII 184 monomers and/or their migration to stacked membrane regions for supercomplex 185 assembly. While MET1 appears to exert this function during PSII de novo biogenesis 186 and during the PSII repair cycle in Arabidopsis, TEF30 appears to function exclusively 187 during PSII repair in Chlamydomonas. s-1 for 8 h (Mettler et al., 2014) (Supplemental Figure S1). The TEF30 ortholog in 188 189 Results 190 TEF30 is an abundant chloroplast protein capable of self-oligomerization 191 As shown in Supplemental Figure S2A, TEF30 proteins are well conserved in the green 192 lineage. The N-termini of TEF30 proteins are predicted to qualify as chloroplast transit 193 peptides and all TEF30 homologs contain PDZ and TPR domains known to mediate 194 protein-protein interactions (Allan and Ratajczak, 2011; Ye and Zhang, 2013) 195 (Supplemental Figure S2B). Following the 36-amino acid chloroplast transit peptide 196 predicted by ChloroP (Emanuelsson et al., 1999), Chlamydomonas TEF30 (but not its 197 homologs from Arabidopsis or moss) harbors sequence motifs characteristic for 198 thylakoid TAT pathway targeting signals (i.e., a RRXXφ twin-arginine motif and an AXA 199 processing site, with X standing for any amino acid and φ for Leu, Val, Phe, or Met) 200 (Albiniak et al., 2012). The region between both motifs is supposed to be hydrophobic, 201 which however is not the case in Chlamydomonas TEF30. The calculated molecular 202 mass of Chlamydomonas TEF30 lacking the predicted 36-amino acids chloroplast transit 203 peptide is 33.3 kDa, while it is 30.4 kDa if the putative TAT targeting signal was 7 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. 204 processed. As shown in Figure 1A, mature TEF30 on SDS-gels with Chlamydomonas 205 whole-cell protein extracts migrated slightly above recombinant TEF30 having a 206 calculated molecular mass of 31.4 kDa (in recombinant TEF30 the predicted chloroplast 207 and TAT targeting signals were replaced with a hexahistine tag, see Supplemental 208 Figure S2B). This indicates that the predicted chloroplast transit peptide is cleaved off in 209 mature TEF30, while the predicted TAT signal is not. Perhaps the incomplete TAT signal 210 mediates TEF30’s targeting to the thylakoid periphery? Mature TEF30 was detected by 211 immunoblot analyses in isolated chloroplasts, but hardly in isolated mitochondria (Figure 212 1B). Moreover, immunofluorescence microscopy revealed TEF30 to be confined to the 213 cup-shaped chloroplast (Figure 1C), thus indicating that TEF30 is a chloroplast protein. 214 The routine detection of TEF30 in shotgun proteomics approaches on 215 Chlamydomonas indicates that it is a well-expressed protein (Hemme et al., 2014; 216 Mettler et al., 2014; Ramundo et al., 2014; Schmollinger et al., 2014). To get an estimate 217 of its cellular abundance, we separated Chlamydomonas whole-cell proteins next to 218 recombinant TEF30 and D2 proteins on SDS gels and quantified signals obtained from 219 immunoblot analyses (Figure 1D). These analyses revealed TEF30 and D2 to represent 220 0.05±0.007 ng/µg chlorophyll (n=4) and 0.59±0.016 ng/µg chlorophyll (n=3), 221 respectively. 222 The PDZ and TPR domains in TEF30 are likely to mediate protein-protein 223 interactions. To test whether TEF30 is capable of self-oligomerization, recombinant 224 TEF30 was subjected to EDC (1-ethyl-3-(3-dimethylaminopropyl)carbodiimide)-mediated 225 zero-length crosslinking and separated by SDS-PAGE. As shown in Figure 1E, 226 untreated recombinant TEF30 migrated with an apparent molecular mass of ~30 kDa, 227 while EDC-treated TEF30 also migrated with apparent masses of ~60, ~90, ~120, and 228 ~150 kDa, thus indicating that TEF30 is indeed capable of self-oligomerization. 229 230 TEF30 is localized to the stromal side of thylakoid membranes 231 TEF30 was previously detected in Chlamydomonas thylakoid membrane fractions 232 (Allmer et al., 2006). To verify TEF30’s thylakoid localization, Chlamydomonas cells 233 were separated into soluble and membrane-enriched fractions by freeze-thawing and 234 fractions were analyzed by immunoblotting. As shown in Figure 2A TEF30, like integral 235 membrane protein cytochrome f, was only detected in the membrane-enriched pellet, 8 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. 236 while stromal CGE1 was only found in the soluble fraction. Cell lysis by sonication had 237 no effect on CGE1’s exclusive localization to the soluble fraction, but resulted in a partial 238 localization of TEF30 and cytochrome f to the soluble fraction, presumably because 239 sonication created small thylakoid membrane vesicles that were not precipitated by 240 centrifugation. Incubation of cells lysed by freeze-thawing in the presence of NaCl, 241 Na2CO3, CaCl2, or urea had no effect on the exclusive localization of cytochrome f to the 242 pellet and of CGE1 to the soluble fraction. However, NaCl led to a partial, and Na2CO3, 243 CaCl2 and urea to a virtually complete shifting of TEF30 to the soluble fraction (Figure 244 2A). Hence, TEF30 appears to be associated with thylakoid membranes rather than 245 embedded into them, which is in agreement with the TMHMM algorithm (Sonnhammer 246 et al., 1998) predicting TEF30 not to contain any transmembrane helices. 247 To determine whether TEF30 is associated with the stromal or the lumenal side of 248 thylakoid membranes, thylakoid membranes from cells lysed by freeze/thawing were 249 purified by flotation on sucrose step gradients and incubated with trypsin (Figure 2B). 250 While the CF1β subunit of the ATP synthase – situated at the stromal side of thylakoid 251 membranes – was almost completely degraded by trypsin, lumenal plastocyanin was 252 degraded only to a small extent and integral membrane protein cytochrome f was not 253 degraded at all. TEF30 was degraded to a similar extent as CF1β, indicating that TEF30 254 is associated to the stromal side of thylakoid membranes. 255 To test, whether TEF30’s association to thylakoid membranes was due to its 256 interaction with one of the major thylakoid membrane protein complexes, we fractionated 257 Chlamydomonas mutants devoid or depleted of PSII, PSI, cytochrome b6/f, or the ATP 258 synthase by freeze/thawing into soluble and membrane-enriched fractions and analyzed 259 the localization of TEF30 by immunoblotting. Although cell-walled strains were used (in 260 contrast to the experiments shown in Figure 2A, which were done with a cell wall- 261 deficient strain), TEF30 in wild-type cells was again mainly detected in membrane- 262 enriched fractions (Figure 2C). Also in PSI, ATP synthase, and cytochrome b6/f mutants 263 most of the TEF30 protein was detected in membrane-enriched fractions, suggesting 264 that TEF30 was not associated with one of these complexes. In contrast, virtually all 265 TEF30 shifted to soluble fractions in mutants affected in the core subunits of PSII (PsbA- 266 D), while TEF30 retained its location to membrane-enriched fractions in a mutant lacking 267 the O subunit of the oxygen evolving complex (OEC) of PSII. As this mutant still 9 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. 268 accumulates considerable amounts of PSII core subunits (de Vitry et al., 1989), our 269 combined data indicate that TEF30 is associated with (a) PSII core subunit(s) exposed 270 to the stromal side of the thylakoid membranes. 271 272 TEF30 co-purifies with PSII particles 273 To verify the interaction of TEF30 with PSII, we took advantage of a Chlamydomonas 274 strain (D2His) in which the D2 subunit was genetically engineered to contain a 275 hexahistidine tag at its C-terminus (Sugiura et al., 1998). Detergent-solubilized thylakoid 276 membranes prepared from wild-type and D2His cells were subjected to nickel-affinity 277 chromatography and bound proteins were eluted with imidazole. Immunoblot analyses 278 revealed TEF30 and D1 to be amply present, PSAD to be present in small amounts, and 279 CF1β and cytochrome f not to be present in eluates from the D2His strain (Figure 3A). 280 None of these proteins was detected in control eluates from the wild-type strain. To 281 make sure that TEF30 was not interacting with PSI present as a contamination in the 282 PSII preparation, we immunoprecipitated TEF30 from purified PSII particles derived from 283 the D2His strain. As shown in Figure 3B and Figure 3C, several proteins co-precipitated 284 with TEF30 but not with preimmune serum. Among the co-precipitating proteins was D1 285 but not PSAD, indicating that TEF30 is tightly interacting with PSII particles and not with 286 PSI. 287 288 TEF30 co-migrates with PSII core subunits on sucrose density gradients and blue 289 native gels 290 To further substantiate our finding that TEF30 interacts with PSII core particles, we 291 isolated thylakoid membranes from cell wall-deficient Chlamydomonas cells by flotation 292 on sucrose step gradients and, following detergent solubilization, separated protein 293 complexes on sucrose density gradients. Immunoblot analyses on fractions taken from 294 these gradients revealed that sample processing resulted in the separation of PSII cores 295 from LHCII and the OEC. This has been observed before when sucrose density gradient 296 centrifugation was employed for analyzing PSII supercomplex compositions (Caffarri et 297 al., 2009; Tokutsu et al., 2012) and in our hands for unknown reasons occurred 298 predominantly with cells lacking cell walls. TEF30 clearly co-migrated with monomeric 10 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. 299 PSII core complexes (Drop et al., 2014) and not with LHCII, the OEC, PSI, the b6/f 300 complex, or the ATP synthase (Figure 4A). 301 Sucrose density gradients were also run with solubilized thylakoid membranes 302 isolated from cell-walled cells of wild-type, PSII mutant and PSI mutant strains. Here, 303 sample processing left most PSII supercomplexes intact (Figure 4B). On gradients run 304 with wild-type thylakoids, TEF30 co-migrated again quantitatively with monomeric PSII 305 core complexes and only traces of TEF30 co-migrated with larger PSII assemblies that 306 were still smaller than fully assembled PSII supercomplexes. TEF30 also did not co- 307 migrate with LHCII, PSI, the b6/f complex, or the ATP synthase. While the quantitative 308 co-migration of TEF30 with monomeric PSII core complexes was not altered in gradients 309 run with thylakoids from the F23 PSI mutant, TEF30 was found to migrate much less 310 deeply into the gradient, to a position even above LHCII, in gradients run with thylakoids 311 from the ΔpsbA/D PSII mutant (Figure 4B). Solubilized thylakoids from cell-walled wild- 312 type and mutant strains were also separated by BN-PAGE and analyzed by 313 immunoblotting. As shown in Figure 4C, TEF30 was again found to co-migrate 314 quantitatively with PSII monomers, but not with PSII supercomplexes, in gel lanes 315 containing solubilized thylakoids from wild-type and PSI mutant strains, while TEF30 316 was not detected in lanes with thylakoids from the nac2 PSII mutant strain. In summary, 317 these results confirm that TEF30 is specifically associated with monomeric PSII core 318 complexes. 319 320 PSII in TEF30-amiRNA cells is more susceptible to photoinhibition than PSII in 321 control cells 322 To get insights into the function of TEF30 in Chlamydomonas chloroplasts, we 323 generated strains constitutively expressing artificial microRNAs (amiRNAs) targeting the 324 coding region of TEF30 transcripts based on the pChlamiRNA2 vector (Molnar et al., 325 2009). Strains cw15-325 and cw15-302 were used as recipients, and in both we could 326 routinely identify transformants in which TEF30 protein levels were reduced to less than 327 20% of wild-type levels (Supplemental Figure S3). When comparing cells from control 328 and TEF30-amiRNA strains we observed no obvious morphological differences (as 329 judged from light microscopy) and no differences in growth under hetero-, mixo- or 11 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. 330 photoautotrophic conditions at various light and temperature regimes (Supplemental 331 Figure S4). 332 As we found membrane-associated TEF30 to accumulate in response to higher 333 light intensities and to be associated with PSII core complexes (Supplemental Figure S1; 334 Figure 2, Figure 3, and Figure 4), we reasoned that TEF30 might play a role in the 335 protection/repair of PSII from photoinhibition. To test this idea, we monitored PSII 336 maximum quantum efficiency (Fv/Fm) in cells from control and TEF30-amiRNA strains 337 grown at ~30 µmol photons m-2 s-1 that had been shifted to high light (HL) at a PFD of 338 1800 µmol photons m-2 s-1 for 60 min and allowed to recover at ~30 µmol photons m-2 s-1 339 for 3 h. As shown in Figure 5A, PSII maximum quantum efficiency declined more 340 strongly and recovered more slowly in TEF30-amiRNA cells as compared to control 341 cells. The same was observed by 77K fluorescence emission spectroscopy 342 (Supplemental Figure S5). To test, whether this apparent higher sensitivity of PSII to HL 343 in TEF30-amiRNA cells was also reflected at the level of PSII subunit accumulation, we 344 shifted cells from control and TEF30-amiRNA strains grown at ~30 µmol photons m-2 s-1 345 to HL of 800 µmol photons m-2 s-1 for 6 h and monitored the accumulation of subunits of 346 all four major thylakoid membrane complexes, as well as of TEF30, VIPP1, and 347 LHCSR3 by immunoblotting (Figure 5B). VIPP1 and LHCSR3 were monitored as 348 controls because they are strongly induced by HL (Supplemental Figure S1; Naumann 349 et al., 2005; Nordhues et al., 2012). VIPP1 was suggested to play a role in the 350 organization of lipid microdomains in thylakoid membranes (Rutgers and Schroda, 351 2013), whereas LHCSR3 was shown to be essential for the induction of non- 352 photochemical quenching (NPQ) in Chlamydomonas (Peers et al., 2009). 353 While TEF30 was virtually absent in the TEF30-amiRNA strain used, it 354 accumulated during HL exposure in cells of the control strain, thus confirming the 355 proteomics data (Supplemental Figure S1). The exposure to HL in neither strain affected 356 the accumulation of cytochrome f, CF1β, LHCII, or the CP26/CP29 minor antenna. In 357 contrast, levels of OEE1/2 proteins declined only in TEF30-amiRNA cells. Moreover, 358 levels of D1, D2, and CP43 declined in control and TEF30-amiRNA cells, but to a much 359 larger extent in the latter, indeed indicating a role of TEF30 in the protection/repair of 360 PSII from photoinhibition. We noted elevated levels of PsaA in TEF30-amiRNA cells that 361 declined during HL treatment to similar levels as did PsaA in control cells. This 12 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. 362 observation appears to be specific to the strain used here, as we did not detect elevated 363 PsaA levels in other TEF30-amiRNA strains. 364 Significant photoinhibition of PSII is known to occur also at low light intensities 365 under chilling conditions (Greer et al., 1988; Li et al., 2004). This is because 366 temperature-dependent reactions during the PSII repair cycle occur with reduced pace 367 at low temperatures, while photochemical reactions are temperature-independent 368 (Tyystjarvi, 2013). To test, whether TEF30-amiRNA cells are also more prone to 369 photoinhibition under chilling conditions, we shifted control and TEF30-amiRNA cells 370 grown at ~30 µmol photons m-2 s-1 from 25°C to 10°C for 18 h and monitored levels of 371 TEF30 and of marker subunits of the four major thylakoid membrane complexes by 372 immunoblotting. As shown in Figure 5C, chilling led to a slight increase in TEF30 protein 373 levels in control cells, but in cells of neither strain did chilling affect levels of PSAD, 374 cytochrome f, and CF1β. However, chilling did induce a decline of D1 levels in TEF30- 375 amiRNA cells, while it did not affect D1 levels in control cells, therefore corroborating the 376 notion that TEF30 plays a role in the protection/repair of PSII from photoinhibition. 377 We made the interesting observation that the accumulation of the LHCSR3 378 protein in HL was impaired in TEF30-amiRNA cells when compared to control cells, 379 while that of VIPP1 was unaffected (Figure 5B). The reduced accumulation of the 380 LHCSR3 protein in TEF30-amiRNA cells was only observed at time points later than 1 h 381 of HL exposure and, as revealed by RT-qPCR, was due to a reduced induction of 382 LHCSR3 gene expression (Figure 6A and Supplemental Figure S6A). Moreover, this 383 effect was only observed in cells grown under mixotrophic conditions with ammonium as 384 nitrogen source (TAP-NH4 medium), while it was not observed in cells grown 385 mixotrophically on nitrate (TAP-NO3) or photoautotrophically on ammonium (TMP-NH4). 386 Accordingly, the induction of NPQ was similar in photoautotrophically grown control and 387 TEF30-amiRNA cells (Supplemental Figure S6B). 388 The induction of LHCSR3 gene expression was shown to correlate with the PFD 389 applied and to be abolished in the presence of photosystem II inhibitor 3-(3,4- 390 dichlorophenyl)1,1-dimethylurea (DCMU) and therefore appears to correlate with the 391 rate of linear electron flow (LEF) (Naumann et al., 2007; Petroutsos et al., 2011; 392 Maruyama et al., 2014). Therefore, a smaller LEF in TEF30-amiRNA cells versus control 393 cells grown mixotrophically in the presence of ammonium and exposed to HL could 13 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. 394 provide an explanation for the reduced induction of LHCSR3 gene expression in TEF30- 395 amiRNA cells. This decrease in LEF might be a consequence of an increased fraction of 396 photodamaged PSII in TEF30-amiRNA cells. To test this idea, we measured oxygen 397 evolution as a direct measure for LEF rates on thylakoid membranes isolated from 398 TEF30-amiRNA and control cells grown at LL (~30 µmol photons m-2 s-1) or HL 399 (~800 µmol photons m-2 s-1) for 4 h. As shown in Figure 6B, the oxygen evolution 400 capacity at 600 µmol photons m-2 s-1 was indeed ~25% lower in HL-grown TEF30- 401 amiRNA cells compared with control cells, while no difference was seen in LL-grown 402 cells. 403 404 Neither PSII de novo synthesis nor PSII stability, but efficient repair of 405 photodamaged PSII is impaired in TEF30-amiRNA cells 406 We observed that levels of the D1 protein of PSII were sometimes slightly lower in LL- 407 grown TEF30-amiRNA cells than in control cells (Figure 5B and Figure 5C). We 408 envisaged two explanations for this finding, namely that TEF30 plays a role in D1 409 synthesis during de novo biogenesis of PSII reaction centers, or that a role of TEF30 in 410 counteracting photoinhibition leads to increased photodamage of PSII in TEF30-amiRNA 411 cells also under LL conditions. To distinguish between these two possibilities, we 412 induced PSII degradation in control and TEF30-amiRNA cells by sulfur starvation for 48 413 h and monitored the re-accumulation of PSII subunits through de novo synthesis upon 414 sulfur repletion for another 24 h (Kosourov et al., 2005; Malnoe et al., 2014). As shown 415 in Figure 7A and Figure 7B, levels of D1, D2, and CP43 declined faster during sulfur 416 starvation in TEF30-amiRNA cells when compared to control cells, but recovery kinetics 417 were the same in both strains. As PSII subunits re-accumulating during sulfur repletion 418 may not necessarily be incorporated into PSII supercomplexes, we analyzed PSII 419 supercomplexes by BN-PAGE during sulfur starvation (for 24 h) and sulfur repletion (for 420 6 h) (Figure 7C and Figure 7D). As judged from D1 and CP43 signals from immunoblots 421 on native gels, PSII supercomplexes in TEF30-amiRNA cells declined and re- 422 accumulated during sulfur starvation and repletion as fast as those in control cells. 423 Interestingly, PSII dimers declined faster in TEF30-amiRNA cells during sulfur 424 starvation, but during sulfur repletion re-accumulated with the same kinetics as in control 14 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. 425 cells. Moreover, levels of unassembled CP43 were higher in TEF30-amiRNA cells than 426 in control cells under control conditions and during sulfur repletion (Figure 7C). 427 Although these data do not support a role of TEF30 in PSII de novo biosynthesis, 428 it was still possible that the absence of TEF30 during PSII de novo synthesis rendered 429 PSII more unstable. To test this, we grew TEF30-amiRNA and control cells under LL 430 conditions, added chloramphenicol (CAP) to inhibit de novo synthesis of plastid-encoded 431 proteins, and left the cultures at LL, placed them into the dark, or exposed them to 200 432 µmol photons m-2 s-1 and monitored levels of PSII subunits. We found D1 levels to 433 decline faster in TEF30-amiRNA cells than in control cells only if CAP-treated cultures 434 were placed in the light (Figure 8). Hence, PSII complexes assembled in TEF30- 435 amiRNA cells are not generally more unstable than those assembled in control cells. 436 We could, however, still not rule out the possibility that improper de novo 437 assembly of PSII in TEF30-amiRNA cells increased the sensitivity of PSII to 438 photodamage. To address this possibility, we exposed cells from control and TEF30- 439 amiRNA strains to HL of 1800 µmol photons m-2 s-1 in the presence of CAP for 1 h. CAP 440 was added to prevent the repair of photodamaged PSII during HL exposure, and the 441 exposure time of 1 h was chosen because after 1 h HL treatment LHCSR3 levels were 442 still comparable in both strains (Figure 6A). After HL exposure, CAP was washed out to 443 allow PSII repair to proceed. As there was no significant difference regarding the decline 444 in PSII maximum quantum efficiency (Fv/Fm) and D1 protein levels between both strains 445 after HL exposure, we can conclude that PSII is equally light sensitive in TEF30-amiRNA 446 and control cells (Figure 9). However, Fv/Fm values and D1 protein levels recovered 447 significantly faster in control than in TEF30-amiRNA cells, strongly suggesting that 448 TEF30 plays a role in PSII repair from photodamage. 449 450 TEF30-amiRNA cells appear to be impaired in PSII supercomplex formation during 451 PSII repair 452 To get an idea at which step of the PSII repair cycle TEF30 might be involved, we 453 analyzed the partitioning of the D1 protein into PSII core and supercomplexes in cells 454 from control and TEF30-amiRNA cells by native Deriphat-PAGE (Formighieri et al., 455 2012). As shown in Figure 10, there was no difference in the partitioning of D1 in cells of 456 both strains grown under LL conditions. However, much less D1 was found to 15 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. 457 accumulate in PSII supercomplexes than in core complexes in TEF30-amiRNA cells 458 compared to control cells when cells had been exposed to HL of 800 µmol photons m-2 459 s-1 for 4 h. TEF30 in native extracts from both strains co-migrated quantitatively with PSII 460 monomers, again corroborating that TEF30 specifically interacts with PSII monomers, 461 but not with PSII supercomplexes. While TEF30 levels were clearly reduced in TEF30- 462 amiRNA cells, HL inducibility and co-migration of TEF30 with monomers were retained 463 in mutant cells. Over all, these data indicate a possible role of TEF30 in redirecting 464 repaired PSII monomers from stroma exposed regions back to appressed regions for 465 PSII supercomplex reassembly after repair. 466 467 Thylakoid membrane stacks in TEF30-amiRNA cells contain fewer lamellae than 468 control cells 469 Finally, to investigate whether depletion of TEF30 had any phenotypes on thylakoid 470 membrane ultrastructure, we analyzed a total of 500 images by transmission electron 471 microscopy from cells of two control and three independent TEF30-amiRNA strains 472 grown at LL or at HL of ~800 µmol photons m-2 s-1 for 4 h. In ~66% of cw15-325 control 473 cells grown in LL we found stacks containing at least 10 thylakoid lamellae (Figure 11) 474 (Nordhues et al., 2012). However, only ~22% of LL-grown TEF30-amiRNA cells 475 contained stacks with more than 10 lamellae, whereas thylakoids were much more 476 loosely organized in ~78% of the mutant cells (Figure 11). While the fraction of cells 477 containing thylakoid stacks with more than 10 lamellae remained unchanged in HL- 478 exposed cells (~61% in control and ~21% in TEF30-amiRNA cells), 26% of control and 479 mutant cells contained swollen thylakoids. The fraction of cells with loosely organized 480 thylakoids declined in control and mutant cells (to 13% and 53%, respectively). These 481 data suggest that the depletion of TEF30 results in less pronounced thylakoid 482 membrane stacking in cells grown in both, LL and HL intensities. 483 484 Discussion 485 We report here on the functional characterization of the TEF30 protein in 486 Chlamydomonas reinhardtii. As a protein present in the green algal, flowering and 487 nonflowering plant lineages, but not detected in nonphotosynthetic organisms, TEF30 is 488 one out of 597 GreenCut proteins. Moreover, because TEF30 is not present in 16 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. 489 cyanobacteria, C. merolae, P. tricornutum, or T. pseudonana, it is part of the ViridiCut 490 subcategory comprising 312 proteins (Supplemental Figure S1A; Merchant et al., 2007; 491 Heinnickel and Grossman, 2013). ViridiCut proteins are likely enriched in green lineage- 492 specific functions, such as mechanisms of chlorophyll a/b protein regulation (Karpowicz 493 et al., 2011) – a prediction that according to the results of this study turned out to be 494 correct. 495 496 TEF30 is a chloroplast protein associated with PSII monomers at the stromal side 497 of thylakoid membranes 498 Immunological data on isolated organelles and immunofluorescence data clearly 499 indicate that TEF30 is localized in the chloroplast (Figure 1B and Figure 1C). Cell 500 fractionation and trypsin digestion experiments revealed TEF30 to be largely associated 501 with the stromal side of thylakoid membranes, although no transmembrane helices were 502 predicted in TEF30. Accordingly, fractionations done in the presence of (chaotropic) 503 salts indicated that the membrane association of TEF30 is realized via electrostatic 504 interactions (Figure 2). The following evidence indicates that TEF30’s association with 505 thylakoid membranes is realized by its direct interaction with PSII monomers: (i) TEF30 506 was largely recovered in soluble fractions when mutant cells deficient in D1, D2, CP43, 507 or CP47 proteins were fractionated into membrane and soluble fractions (Figure 2C). (ii) 508 TEF30 co-eluted with affinity-purified PSII particles (Figure 3). (iii) TEF30 co-migrated 509 quantitatively with PSII monomers on sucrose density gradients, on blue-native gels, 510 and on native Deriphat gels (Figure 4 and Figure 10). (iv) Co-migration of TEF30 with 511 PSII monomers was abolished if thylakoid membrane proteins were derived from PSII 512 mutants (Figure 4B and Figure 4C). 513 With TEF30 and D2 representing ~0.05 and ~0.59 ng protein per ng chlorophyll 514 (Figure 1D) and exhibiting molecular masses of 33.3 and 39.45 kDa, respectively, 515 TEF30 on a molar basis is ~10 times less abundant than D2. Given that some TEF30 516 was also recovered in soluble fractions and that TEF30 was found to be able to form 517 oligomers (Figure 1E, Figure 2A and Figure 2C), the stoichiometry of TEF30 518 (oligomers?) to PSII is likely below 1:10. PSII monomers are not the main active form of 519 PSII and are mainly located to stroma thylakoids (Danielsson et al., 2006). Therefore, 520 TEF30’s association with PSII monomers is in agreement with the calculated 17 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. 521 stoichiometries between TEF30 and D2. A localization of TEF30 to stroma lamellae was 522 also indicated by the finding that TEF30 and CF1β with a known location to stroma 523 lamellae shared the same sensitivity to trypsin digestion of isolated thylakoid 524 membranes. Consistently, proteomics approaches on thylakoid membranes separated 525 into grana and stroma lamellae identified the Arabidopsis TEF30 homolog (AT1G55480) 526 exclusively in stroma-lamellae (Tomizioli et al., 2014). 527 528 TEF30 is involved in PSII repair 529 TEF30 shares its ability to associate with PSII components at the stroma side with a 530 number of other stromal proteins, including Deg7 (Sun et al., 2010), THF1/Psb29/NYC4 531 (Wang et al., 2004; Huang et al., 2013), AtPDI6 (Wittenberg et al., 2014), LPA3 (Cai et 532 al., 2010), and ROC4 (Lippuner et al., 1994; Cai et al., 2008). While Deg7, 533 THF1/Psb29/NYC4, AtPDI6, and ROC4 play roles in the PSII repair cycle (Keren et al., 534 2005; Cai et al., 2008; Sun et al., 2010; Huang et al., 2013; Yamatani et al., 2013; 535 Wittenberg et al., 2014), LPA3 appears to be required for coordinating the membrane 536 insertion of de novo synthesized CP43 (Cai et al., 2010). 537 Of these possible functions of TEF30 (PSII de novo synthesis or repair), our data 538 argue against a role of TEF30 in PSII de novo synthesis. This is because control and 539 TEF30-amiRNA cells, in which PSII degradation was induced by sulfur starvation 540 (Kosourov et al., 2005; Malnoe et al., 2014), re-accumulated de novo synthesized PSII 541 core components, dimers and supercomplexes with the same kinetics after sulfur 542 repletion (Figure 7). As judged from blue native and native Deriphat gels, PSII 543 (super)complexes in TEF30-underexpressing cells also appear to be correctly 544 assembled (Figure 7C and Figure 10). This is supported by the observation that in the 545 presence of chloroplast protein synthesis inhibitors, PSII stability in the dark and PSII 546 sensitivity to HL exposure were comparable with those of control cells (Figure 8C and 547 Figure 9). Moreover, O2 evolution capacities of LL-grown cells were the same in control 548 and TEF30-underexpressing strains (Figure 6B). 549 Rather, our results point to a role of TEF30 in the PSII repair cycle: most 550 importantly, PSII activity in the absence of protein synthesis inhibitors was more 551 sensitive to HL and recovered more slowly in TEF30-underexpressing cells than in 552 control cells (Figure 5A; Supplemental Figure S5). This was reflected by the 18 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. 553 accumulation kinetics of PSII core subunits, which after HL exposure or in LL under 554 chilling conditions (known to enhance photoinhibition (Greer et al., 1988; Li et al., 2004)) 555 declined more pronouncedly in TEF30-underexpressing than in control cells, while minor 556 and major antenna were unaffected (Figure 5B and Figure 5C). Moreover, PSII stability 557 in the presence of chloroplast protein synthesis inhibitors was impaired in TEF30- 558 underexpressing cells only when they were grown in the light, but not when they were 559 grown in the dark (Figure 8). Finally, like TEF30-underexpressing cells, mutants in 560 factors known to be involved in PSII repair hardly display any phenotypes if not grown 561 under high / fluctuating light (Supplemental Figure S4) (Keren et al., 2005; Sirpio et al., 562 2007; Cai et al., 2008; Sun et al., 2010). 563 At which step of the PSII repair cycle may TEF30 play a role? Native Deriphat 564 gels indicated a strongly reduced accumulation of PSII supercomplexes in TEF30- 565 underexpressing versus control cells following an exposure to HL for 4 h (Figure 10). 566 This observation, combined with the finding that TEF30 did not co-migrate with PSII- 567 LHCII supercomplexes on sucrose density gradients, Deriphat- and BN-PAGE (Figure 4 568 and Figure 10), points to a role of TEF30 in directing repaired PSII monomers from 569 stroma-exposed regions back to stacked regions for PSII dimer/supercomplex re- 570 assembly, and/or in stabilizing/protecting repaired PSII monomers (Figure 12). In 571 Arabidopsis, grana margins (when compared with grana cores and stroma membranes) 572 were shown to represent the membrane regions in which PSII core subunits are mainly 573 dephosphorylated, D1 levels are lowest and FtsH levels highest and were therefore 574 identified as the main thylakoid subdomain for proteolytic degradation of D1 (Tietz et al., 575 2015). Hence, if unprotected PSII monomers in TEF30-underexpressing cells would 576 linger too long in such proteolytically active domains after repair, they may undergo 577 further cycles of D1 degradation and replacement, thus explaining the inefficient PSII 578 repair in TEF30-underexpressing cells. However, TEF30 may also facilitate complex re- 579 assembly (e.g. of temporally released CP43) or the incorporation of de novo synthesized 580 D1 during repair (Figure 12). In the absence of de novo D1 synthesis during PSII repair, 581 PSII core monomer complexes completely disappeared from stroma membranes (Aro et 582 al., 2005). PSII monomers whose re-assembly is delayed during repair may therefore be 583 more prone to proteolytic attack. It is important to note that TEF30 apparently facilitates 584 the PSII repair cycle but is not essential for it. The PSII repair cycle exists in all 19 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. 585 organisms capable of performing oxygenic photosynthesis but TEF30 is found only in 586 the Viridiplantae. Therefore, its role in facilitating PSII repair may have evolved to help 587 overcome constraints to the PSII repair cycle that are specific to the Viridiplantae. 588 589 Reduced accumulation of LHCSR3 and reduced thylakoid membrane stacking 590 may be pleiotrophic effects of impaired PSII repair in TEF30-underexpressing cells 591 Upon HL exposure, the LHCSR3 protein accumulated to lower levels in TEF30- 592 underexpressing cells than in control cells (Figure 5B). LHCSR3 becomes activated by 593 an increased acidification of the thylakoid lumen and mediates the dissipation of excess 594 light energy by a protonation-induced interaction with PSII supercomplexes which is 595 dependent on the PSBR protein (Bonente et al., 2011; Tokutsu and Minagawa, 2013; 596 Xue et al., 2015). The reduced accumulation of LHCSR3 in TEF30-underexpressing 597 cells was observed only when cells were exposed for more than 1 h to HL and was 598 realized at the level of gene expression (Figure 6A and Supplemental Figure S6A). 599 Moreover, the effect was only observed in cells grown mixotrophically with ammonium 600 as nitrogen source and not in cells grown under photoautotrophic conditions with 601 ammonium, or grown mixotrophically with nitrate as nitrogen source (Figure 6A). Hence, 602 it appear as if the impaired induction of the LHCSR3 gene in TEF30-underexpressing 603 cells is limited to conditions in which sinks for products of the light reactions are limiting 604 (nitrate reduction and carbon fixation are highly ATP- and NADPH-demanding). 605 LHCSR3 gene expression was shown to correlate with the PFD applied and thus with 606 linear electron flow (LEF) rates (Naumann et al., 2007; Peers et al., 2009; Petroutsos et 607 al., 2011; Maruyama et al., 2014). Accordingly, in cells grown mixotrophically in the 608 presence of ammonium and exposed to HL for >1 h, LEF rates are supposed to be 609 lower if TEF30 levels are low. This indeed appeared to be the case, as the oxygen 610 evolution capacity was ~25% lower in HL-exposed TEF30-underexpressing cells than in 611 control cells, but unaltered in LL-grown cells (Figure 6B). Therefore, we propose that the 612 reduced accumulation of LHCSR3 in TEF30-underexpressing cells grown under 613 apparently sink-limited conditions is a consequence of increased PSII photoinhibition 614 and thus of reduced LEF under such conditions rather than the cause for increased 615 photoinhibition. In support of this idea, blocking the Calvin-Benson cycle was shown to 20 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. 616 inhibit the repair of photodamaged PSII, 617 Chlamydomonas (Takahashi and Murata, 2005). thus enhancing photoinhibition in 618 Using transmission electron microscopy we observed that ~66% of the cells of 619 TEF30-underexpressing strains contained fewer than 10 lamellae in thylakoid 620 membrane stacks, whereas this was true for only ~22% of the cells in control strains 621 (Figure 11). Therefore, the thylakoid organization in control cells under our growth 622 conditions (LL of ~30 µmol photons m-2 s-1) resembles that of shade leaves with many 623 thylakoids per stack, while that of TEF30-underexpressing cells resembles the 624 organization found in sun leaves, with fewer lamellae per stack (Guillot-Salomon et al., 625 1978; Lichtenthaler et al., 1981). One possible explanation for this phenotype is that 626 TEF30 directly affects thylakoid stacking and impaired PSII repair is a consequence of 627 this. The number of membrane layers in grana stacks was shown to correlate with the 628 abundance of the CURT1 protein, which is located to grana margins and induces 629 membrane curvature (Armbruster et al., 2013). Hence, TEF30 might positively modulate 630 the function of CURT1, leading to reduced membrane stacking in TEF30- 631 underexpressing cells. An argument against this idea is that reduced thylakoid 632 membrane stacking was shown to improve PSII repair by facilitating lateral migration of 633 damaged PSII supercomplexes and therefore increasing their accessibility to the FtsH 634 protease (Tikkanen et al., 2008; Fristedt et al., 2009; Kirchhoff et al., 2013; Nath et al., 635 2013b). Indeed, reduced membrane stacking may account for the faster degradation 636 rates of PSII dimers and supercomplexes upon sulfur starvation in TEF30- 637 underexpressing cells (Figure 7), but conflicts with their impaired capacity for PSII repair. 638 A second, more plausible explanation for the reduced thylakoid membrane stacking in 639 TEF30-underexpressing cells is that it represents a countermeasure of these cells to 640 improve PSII repair, e.g. by reducing CURT1 function at the levels of CURT1 protein 641 accumulation or phosphorylation state, or by increasing the phosphorylation state of PSII 642 core subunits (Tikkanen et al., 2008; Fristedt et al., 2009; Goral et al., 2010; Samol et 643 al., 2012; Armbruster et al., 2013; Kirchhoff et al., 2013; Tietz et al., 2015). 644 645 Did an additional function for TEF30 in supercomplex assembly of de novo 646 synthesized PSII evolve in land plants? 21 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. 647 The Arabidopsis TEF30 homolog (AT1G55480) was first described by Ishikawa et al. 648 (2005), who termed it ZKT for protein containing a PDZ domain, a K-box region, and a 649 TPR region and proposed a role for the protein in wounding responses. Very recently, 650 the same protein was named MET1, for M-enriched thylakoid protein 1 (where M stands 651 for mesophyll cells) (Bhuiyan et al., 2015). Our results on Chlamydomonas TEF30 are 652 largely congruent with those from the excellent study by Bhuiyan et al. with respect to 653 the inducibility of TEF30/MET1 by HL, the interaction of the proteins with the stroma side 654 of PSII complexes in thylakoid membranes, the lack of obvious phenotypes in mutants 655 under standard growth conditions, as well as the increased sensitivity of PSII and the 656 reduced accumulation of PSII supercomplexes in TEF30/MET1-depleted mutants 657 compared to controls upon HL exposure. Therefore we agree with Bhuiyan et al. that a 658 function of TEF30/MET1 in wounding is unlikely. 659 However, there are also differences between traits observed for TEF30/MET1 in 660 Chlamydomonas and Arabidopsis, which led to slightly different conclusions regarding 661 the function of this protein. By immunoblot analysis of Arabidopsis thylakoid membrane 662 protein complexes separated by BN-PAGE, Bhuiyan et al. found MET1 to co-migrate 663 with several PSII (sub)complexes including dimers, monomers, CP43-less monomers, 664 reaction centers, and unassembled proteins. With the yeast 2-hybrid system they also 665 found MET1 to directly interact with stromal loops of CP43 and CP47. Based on these 666 data Bhuiyan et al. concluded that MET1 is involved in the folding and/or stabilization of 667 these stroma-exposed CP43/CP47 domains, thereby ensuring their correct configuration 668 upon assembly of the PSII core during de novo PSII biogenesis and during the PSII 669 repair cycle. By analyzing Chlamydomonas thylakoid membrane protein complexes on 670 sucrose density gradients, BN-PAGE and Deriphat-PAGE, we observed TEF30 to 671 quantitatively interact with PSII monomers and observed only traces of TEF30 to co- 672 migrate with larger PSII assemblies (sucrose gradients, Figure 4B) or smaller ones 673 (Deriphat-PAGE, Figure 10). Moreover, based on the finding that PSII supercomplex 674 formation was only impaired in TEF30-underexpressing cells pushed towards PSII repair 675 by HL exposure (Figure 10) and not in cells pushed towards de novo PSII synthesis 676 during recovery from sulfur starvation (Figure 7), we concluded that TEF30 is rather 677 involved in PSII repair and not in the assembly of de novo synthesized PSII complexes. 678 Whether these differences are due to experimental setups or whether MET1 has 22 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. 679 acquired an additional role in PSII de novo assembly during land plant evolution remains 680 to be elucidated. The latter idea may be supported by the finding that PSII de novo 681 synthesis and repair appear to be spatially separated in Chlamydomonas (Uniacke and 682 Zerges, 2007), while both processes appear to be only temporally separated during cell 683 differentiation in land plants (Nickelsen and Rengstl, 2013). 684 685 Materials and Methods 686 Chlamydomonas strains and culture conditions 687 Chlamydomonas strain cw15-325 (cwd, mt+, arg7-) and cw15-302 (cwd, mt+, arg7-, nit-), 688 kindly provided by R. Matagne, University of Liège, Belgium, were used as recipient 689 strains for transformation with plasmid pCB412 (containing only the ARG7 gene) 690 (Schroda et al., 1999) to generate control strains, or with pMS632 (containing the ARG7 691 gene and the TEF30-amiRNA construct) to downregulate TEF30. Transformations were 692 carried out using 1 µg of HindIII-linearized plasmids and vortexing with glass beads 693 (Kindle, 1990). Photosynthesis mutants used were F23 (lacking PsaA) (Girard et al., 694 1980), FUD34 (lacking CP43) (Rochaix et al., 1989), FUD44 (lacking OEE1) (Mayfield et 695 al., 1987), ΔbB/ ΔpsbB (lacking CP47) (Minai et al., 2006), FUD47 and nac2 (both 696 lacking D2) (Erickson et al., 1986; Kuchka et al., 1989), FUD50 (lacking CF1β) 697 (Woessner et al., 1986), ΔpetA (lacking cyt f) (Rimbault et al., 2000), and a double 698 mutant ΔpsbA/ΔpsbD (lacking D1 and D2). Strain cc124 (mt-) was employed as a wild- 699 type control for experiments with photosynthesis mutants. Strain D2-H (Sugiura et al., 700 1998) was a gift from J. Minagawa, National Institute for Basic Biology, Japan. Unless 701 specified otherwise, all strains were grown mixotrophically in Tris-acetate-phosphate 702 (TAP) medium (Harris, 2008) supplemented with 7.5 mM NH4Cl and a revised 703 micronutrient composition (Kropat et al., 2011) on a rotatory shaker at 25°C and 704 ~30 μmol m-2 s-1. Minimal medium (TMP) was prepared in the same way, but lacking 705 acetate. Cultures were diluted the day before experiments were conducted and grown to 706 mid-log phase. Cell densities were determined using a Z2 Coulter Counter (Beckman 707 Coulter). 708 709 Phylogenetic analyses 23 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. 710 Multiple sequence alignments 711 Chlamydomonas 712 (XP_001416302), Micromonas 713 (XP_002955399), Physcomitrella 714 moellendorffii 715 (AP004670.4), 716 (XP_003629926), and Glycine max (XP_001235451) using the ClustalW2.1 program 717 ((Larkin et al., 2007). Transit peptide sequences were predicted with ChloroP 718 (Emanuelsson et al., 1999) or TargetP (Emanuelsson et al., 2000) programs. The 719 phylogenetic tree was inferred by the neighbor-joining method using the MEGA5.1 720 program (Tamura et al., 2011) and statistical support was assessed with 1000 bootstrap 721 replicates. reinhardtii were with (Cre01.g031100.t1.2), (XP_002988351), Arabidopsis performed patens thaliana mays sequences Ostreococcus (XP_003059792), pusilla Zea protein (XP_008652074), (AT1G55480.1), lucimarinus Volvox (Phpat.019G052300.1), carteri Selaginella Oryza Medicago from sativa truncatula 722 723 Cloning, expression and purification of recombinant TEF30 724 The TEF30 coding sequence was amplified from Chlamydomonas cDNA with primers 5’- 725 tactggatccgccgaatacatcgagctgga-3’ 726 The resulting 817-bp PCR product was digested with BamHI and HindIII and cloned in 727 frame into the PetDuet vector (Novagen) cleaved with the same enzymes, yielding 728 pMS649. TEF30 was overexpressed in E. coli BL21 and purified by nickel-nitrilotriacetic 729 acid agarose (Ni-NTA) according to the manufacturer’s instructions (Qiagen). Imidazole 730 was largely removed by three successive runs of concentration/dilution with KMH buffer 731 (20 mM HEPES-KOH pH 7.2, 80 mM KCl, 2.5 mM MgCl2) using Amicon Ultra-4 tubes 732 (Millipore). After that, proteins were supplemented with 10% glycerol and 1 mM 733 dithiothreitol (DTT). For antibody generation TEF30 was purified under denaturing 734 conditions with guanidine-HCl and eluted with 200 mM imidazole in 6 M urea. 1 mg of 735 purified protein was used for a 3-months rabbit immunization protocol (SeqLab) for 736 antibody generation. and 5’-gcaacatgttcaacaaggagaagtaagcttactca-3’. 737 738 Protein analyses and thylakoid membrane isolation 739 Protein extractions, SDS–PAGE, semi-dry blotting, and immunodetections were carried 740 out as described before (Liu et al., 2005; Schulz-Raffelt et al., 2007). Proteins on SDS- 741 polyacrylamide gels were loaded on the basis of chlorophyll concentrations or protein 24 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. 742 concentrations determined according to (Lowry et al., 1951). Immunodetection was done 743 using enhanced chemiluminescence (ECL) and the FUSION-FX7 Advance™ imaging 744 system (PEQLAB). Antisera described previously were against mitochondrial carbonic 745 anhydrase (Agrisera AS11 1737), HSP70B (Schroda et al., 1999), HSP70A (Schulz- 746 Raffelt et al., 2007), VIPP1 (Liu et al., 2005), LHCSR3 (Naumann et al., 2007), CF1β 747 (Lemaire and Wollman, 1989), PSAD (Naumann et al., 2005), LHCII (Vallon et al., 748 1986), CP43 (Vallon et al., 1985), Cyt f (Pierre and Popot, 1993), D1 (Agrisera AS05 749 084), OEE33 (Agrisera AS06 142-33), OEE23 (Agrisera AS06 167), PsaA (Agrisera 750 AS06 172), PsaB (Agrisera AS10 695), CP26 (Agrisera AS09 407), CP29 (Agrisera 751 AS06 117), mtCA (Agrisera AS11 1737), D2 (Agrisera AS06 146), and CGE1 (Schroda 752 et al., 2001). Densitometric quantification of the bands after immunodetection was 753 calculated by the FUSIONCapt Advance program for image capture and analysis 754 (PEQLAB). Thylakoids were isolated as described previously (Chua and Bennoun, 755 1975). Briefly, cells were lysed using a BioNebulizer (Glas-Col) with an operating N2 756 pressure of 1.5-2 bar and lysates centrifuged through a sucrose step gradient (0.6, 1.3, 757 and 1.8 M sucrose) at 40,000 rpm for 1 h. Thylakoids floating between the 1.8 and 1.3 M 758 transition were collected, diluted in a buffer containing 5 mM HEPES-KOH pH 7.5 and 759 10 mM EDTA, and pelleted by a centrifugation at 40,000 rpm for 15 min. 760 761 EDC-crosslinking of recombinant TEF30 762 100 ng of recombinant TEF30 was incubated in KMH buffer with 2.5 mM EDC (1-Ethyl- 763 3[3-dimethylaminopropyl]carbodiimid hydrochlorid) and the double concentration of NHS 764 (N-hydroxysuccinimid). The mixture was incubated for 30 min at 25°C. The crosslinking 765 reaction was stopped by the addition of 0.2 M ammonium acetate and samples were 766 analyzed by immunoblotting. 767 768 Cell fractionations 769 Isolation of chloroplasts was performed as described previously (Zerges and Rochaix, 770 1998). Mitochondria were isolated according to (Eriksson et al., 1995), but using a 771 BioNebulizer (Glas-Col) for cell disruption. For the separation of soluble and membrane 772 enriched fractions, 2 x 107 cells were harvested by a 2-min centrifugation at 1300 g and 773 4°C and resuspended in lysis buffer (10 mM Tris-HCl pH 8.0, 1 mM EDTA, 0.25 x 25 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. 774 protease inhibitor cocktail (Roche)). An aliquot was taken for whole-cell proteins. The 775 remainder was frozen in liquid nitrogen and thawed at 25°C. This cycle was repeated 776 two times. Disrupted cells were centrifuged for 30 min at 18000 g and 4°C. The 777 supernatant was collected as soluble protein fraction. Pellets were resuspended by brief 778 sonication with the same volume of lysis buffer. When indicated, a final concentration of 779 250 mM NaCl, 200 mM Na2CO3, 1 M CaCl2, or 6 M urea was added to the lysis buffer 780 before freeze/thawing the cells. For localization experiments, isolated thylakoids were 781 incubated on ice with 0.1 mg/mL trypsin. After the addition of 0.25 x proteinase inhibitor 782 cocktail (Roche), proteins were precipitated with 80% acetone. Pellets were 783 resuspended in Laemmli buffer for SDS-PAGE (Laemmli, 1970). 784 785 Vector construction and screening for TEF30-underexpressing transformants 786 The amiRNA targeting TEF30 was designed according to (Molnar et al., 2009) using the 787 WMD2 web platform (http://wmd2.weigelworld.org) (Ossowski et al., 2008). The resulting 788 oligonucleotides 789 ctagtAAGTTCGCTCGCGGTGGCGAAtctcgctgatcggcaccatgggggtggtggtgatcagcgctaTTC 790 GTTACCGCGAGCGAACTTg-3’ 791 ctagcAAGTTCGCTCGCGGTAACGAAtagcgctgatcaccaccacccccatggtgccgatcagcgagaTT 792 CGCCACCGCGAGCGAACTTa -3’ were annealed by a gradual temperature reduction 793 from 100°C to 60°C (0.3°C/min) in a thermocycler and ligated into the pChlamiRNA2 794 (Molnar et al., 2009), yielding pMS632. Correct constructs were screened as described 795 by Molnar et al. and sequenced. After transformation of this construct into 796 Chlamydomonas cells, 50 arginine-prototrophic transformants of each recipient strain 797 (cw15-325 or cw15-302) were screened by immunoblotting. 5’and 5’- 798 799 Cold stress, high light, photoinhibition and sulfur starvation experiments 800 For cold stress, 150 mL of 2 x106 cells grown at 25°C were incubated under continuous 801 agitation in a water bath at 10°C and ~30 μmol photons m−2 s−1 for 18 h. High light 802 experiments were performed with 150 mL of 2 x106 cells in 400-mL beakers placed onto 803 spots on a rotary shaker that were illuminated (~800 μmol photons m−2 s−1) by Osram 804 HLX 250W 64663 Xenophot bulbs. A glass plate non-permissive to infrared and UV 805 irradiation was placed between light sources and beakers. The position of the beakers 26 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. 806 was altered in regular intervals and the cell culture temperature maintained at 25°C. The 807 same setup was used for photoinhibition, but cultures were exposed to ~1800 μmol 808 photons m−2 s−1. For recovery after photoinhibition, cultures were placed on a rotatory 809 shaker at 25°C and ~30 μmol photons m–2 s–1. When indicated, 500 μg mL−1 lincomycin 810 and/or 100 μg mL−1 chloramphenicol were added before photoinhibiton or in the 811 beginning of the recovery phase. To wash out the inhibitors, cells were centrifuged for 812 2 min at 956 g and 25°C and cell pellets were resuspended in fresh TAP-NH4 medium. 813 Photon flux densities (PFD) were determined with a light meter (LI-COR LI-250A). Sulfur 814 starvation experiments were performed in TAP-NH4 according to (Malnoe et al., 2014), 815 but cultures were maintained at ~50 μmol photons m–2 s–1. To follow de novo synthesis 816 of photosystem II, cells were centrifuged and resuspended in TAP-NH4 medium and 817 kept in the light (~30 μmol photons m-2 s–1) for 24 hours. 818 819 Isolation of PSII particles and immunoprecipitation 820 Isolation of PSII particles was performed with ca. 2 L of 3 x106 cells/mL of the D2His 821 strain according to Cullen et al. (2007) with a ProBondTM resin column (Invitrogen). After 822 their elution, PSII particles were mixed with an equal volume of 20 mM MES, pH 6.3, 823 15 mM NaCl, 20% (w/v) PEG 8000, 10 mM ascorbic acid and 10 mM EDTA, and 824 centrifuged at 11,000 g for 10 min. The pellet was resuspended in 20 mM MES pH 6.3, 825 15 mM NaCl, 10 mM ascorbic acid, 10% PEG, 10 mM CaCl2 and centrifuged as before 826 to decrease the imidazole concentration. The pellet containing isolated PSII particles 827 was resuspended in 250 µL of buffer A (20 mM MES, pH 6.3, 15 mM NaCl, 333 mM 828 sucrose, and 10 mM CaCl2). The same procedure was followed with cc124 cells as 829 control. The coupling of antibodies against TEF30 and of preimmune serum to protein A- 830 Sepharose beads for immunoprecipitation was carried out as described previously 831 (Schroda et al., 2001; Liu et al., 2005). Coupled beads were washed twice in buffer A 832 and incubated with 250 µL of isolated PSII particles for one hour at 4°C. Beads were 833 then washed three times in buffer B (buffer A with the addition of 0.1% (v/v) of Triton 834 X-100) and two times in 10 mM Tris-HCl, pH 7.5. Immunopreciptated proteins were 835 collect by boiling the beads in 2% SDS and 12% sucrose for 40 sec. After addition of 0.1 836 M DTT, eluates were subjected to SDS-PAGE. 837 27 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. 838 Blue native-PAGE, Deriphat-PAGE and sucrose density centrifugation 839 Blue native-PAGE (BN-PAGE) was performed with isolated thylakoids, crude membrane 840 fractions or whole-cell proteins according to (Schagger and von Jagow, 1991; Schagger 841 et al., 1994). Native Deriphat-PAGE was performed according to (Peter and Thornber, 842 1991) with a 4% acrylamide stacking gel poured upon the 7.5% acrylamide running gel. 843 Thylakoid membranes (0.25 mg chlorophyll/mL) were solubilized for 20 min on ice in 5 844 mM HEPES-KOH pH 7.5 and 1 mM EDTA containing 1% α-dodecyl maltoside (α-DDM). 845 For sucrose density centrifugation, isolated thylakoids were solubilized with 1% β- 846 dodecyl maltoside (β-DDM) and loaded onto a linear 0.2 to 0.5 M sucrose gradient 847 containing 10 mM HEPES-KOH pH 7.5 and 0.05% β-DDM. On the bottom of the 848 gradient a 2 M sucrose cushion was present. The gradient was centrifuged at 205,000 g 849 for 16 h at 4°C and 25-26 fractions were collected from bottom to top. 850 851 Immunofluorescence staining and transmission electron microscopy 852 Immunofluorescence microscopy was performed with control transformants in the cw15- 853 325 strain background. Cells were fixed and stained as described previously (Uniacke et 854 al., 2011). Primary antibodies were against TEF30, HSP70B and HSP70A, used in 855 1:300, 1:8000, 1:4000 dilutions, respectively. As a secondary antibody we used a 856 fluorescein isothiocyanate–labeled goat anti-rabbit antibody (Sigma- Aldrich) in 1:500 857 dilution. After incubation with the secondary antibody, slides were washed in PBS and a 858 drop of mounting solution containing DAPI (Vectashield, Vector Laboratories, United 859 Kingdom) was applied at the center of each the slide. Images were obtained with an 860 OLYMPUS BX53 microscope with a violet filter for DAPI and a green filter for FITC and 861 an OLYMPUS DP26 color camera. For transmission electron microscopy, cells were 862 collected, fixed and embedded as described previously (Nordhues et al., 2012). Images 863 were analyzed with a JEM-2100 (Jeol) or EM 902A (Carl Zeiss) transmission electron 864 microscope (both operated at 80 kV). Micrographs were taken using a 4080x4080 pixel 865 or 1350x1040 pixel charge-coupled device camera (UltraScan 4000 or Erlangshen 866 ES500W, respectively; Gatan) and the Gatan Digital Micrograph software (version 867 1.70.16). 868 869 RNA extraction and qRT-PCR 28 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. 870 2 x107 cells were collected for RNA extraction with RNeasy Plant Mini kit (Qiagen) 871 following the manufacturer’s instructions. DNA contaminations were removed with 872 DNAse (RNase-free Turbo DNase, Ambion). cDNA synthesis was done with the MULV 873 reverse transcriptase (Promega), deoxynucleotide triphosphate and oligo(dT)18 primers. 874 qRT-PCR was performed using the StepOnePlus RT-PCR system (Applied Biosystems) 875 and the Maxima SYBR Green kit from Fermentas as described in (Strenkert et al., 876 2011). Primers for LHCSR3.1 were the same as described in (Peers et al., 2009). 877 878 Biophysical measurements 879 Fluorescence induction measurements were performed at 25°C with a DeepGreen 880 Fluorometer (JBeamBio). The maximum quantum efficiency of PSII, Fv/Fm, was 881 determined after incubating cells in the dark for 1 min according to the equation Fv = Fm - 882 F0, where F0 represents the fluorescence when the electron acceptor of PSII is fully 883 oxidized and Fm the maximum PSII fluorescence. Fm was measured during 250 ms 884 saturating flash of light or in the presence of 10 mM DCMU. Low-temperature 885 fluorescence emission spectra were performed with a laboratory built setup. Samples 886 were submerged in liquid nitrogen and excited with a LED source (LS-450, Ocean Optics 887 - blue LED, 450 nm). The emission spectra were measured by a CCD spectrophotometer 888 (QE6500, Ocean Optics). The device was calibrated by the manufacturer and calibration 889 was verified using standard band-pass filters. For the fluorescence quenching 890 measurements, cells were grown mixotrophically in TAP-NH4 medium at 50 µmol photons 891 m-2 s-1, harvested in the exponential phase and resuspended in TMP-NH4 at a 892 concentration of 2 x 107 cells per mL. Cells were agitated and exposed to HL intensities of 893 500 µmol photons m-2 s-1 provided by a WH1200 Phyto (JBeambio) to induce the 894 accumulation of LHCSR3. The NPQ response was measured using a chlorophyll 895 fluorescence imaging system (Johnson et al., 2009). Samples were incubated in the dark 896 under strong agitation for 15 min before each measurement. NPQ was calculated as (Fm 897 - Fm’) / Fm’, where Fm’ is the maximum PSII fluorescence in light (after 10 sec at 898 700 μmol photons m-2 s-1). Fm and Fm’ were measured after exposure to a brief 899 saturating pulse of 2100 μmol photons m-2 s-1 for 250 ms. Oxygen evolution was 900 measured with a Clark-type oxygen electrode in the presence of 0.5 mM 2,6- 901 dichlorobenzoquinone (2,6-DCBQ) and 2 mM potassium hexacyanoferrate 29 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. (III) 902 (K3Fe(CN)6) as electron acceptors. The measurements were carried out with crude 903 membrane fractions in TMP-NH4 medium at 25°C. 904 905 Supplemental Figure S1. TEF30 protein accumulates in Chlamydomonas cells 906 exposed to high light. 907 908 Supplemental Figure S2. Phylogenetic tree and alignment of TEF30 homologs. 909 910 Supplemental Figure S3. Screening of TEF30-amiRNA transformants. 911 912 Supplemental Figure S4. TEF30-underexpressing transformants exhibit no 913 obvious growth phenotype under a variety of growth conditions. 914 915 Supplemental Figure S5. PSII fluorescence is more affected by high light in 916 TEF30-amiRNA cells than in control cells. 917 918 Supplemental Figure S6. Reduced induction of LHCSR3 expression in TEF30- 919 amiRNA cells is realized at the transcriptional level and NPQ is not affected under 920 photoautotrophic conditions. 921 922 Acknowledgments 923 We would like to thank Jun Minagawa for providing the D2His strain, Michael Hippler for 924 providing antisera against LHCSR3 and PSAD, and Fabrice Rappaport for helping with 925 the biophysical measurements. 926 927 Figure legends 928 Figure 1. Antibody characterization and analysis of TEF30 oligomer formation, 929 cellular abundance and intracellular localization. 930 (A) Immunodetection of TEF30 in cw15-325 whole-cell proteins (WC) corresponding to 2 931 µg chlorophyll compared with 0.25 ng recombinant TEF30 protein. 932 (B) Subcellular localization of TEF30 by immunoblotting. 10 or 3 µg (depending on the 933 antiserum used) protein from whole cells (WC), chloroplasts (cp) and mitochondria (mt) 30 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. 934 isolated from strain cw15-302 were separated by SDS-PAGE and immunodetected with 935 antisera against TEF30, mitochondrial carboanhydrase (mtCA), and stromal HSP70B. 936 (C) Microscopy images taken from strain cw15-325. Shown are from left to right: DIC 937 images, DAPI staining, immunofluorescence, merge of DAPI and immunofluorescence, 938 and a schematic drawing that combines information from the images. Antisera used for 939 immunofluorescence were against HSP70B and HSP70A as markers for stroma and 940 cytosol, respectively, and against TEF30. Chloroplast – cp; pyrenoid – P; cytosol – C; 941 flagella – f; immunofluorescence signal – green. 942 (D) 1 to 8 ng of recombinant proteins TEF30 (purified from E. coli) and D2 (purchased 943 from Agrisera) were separated by SDS-PAGE together with cw15-325 whole-cell 944 proteins corresponding to 0.125-2 µg chlorophyll and immunodetected with antibodies 945 against TEF30 and D2. 946 (E) 100 ng of recombinant TEF30 protein was incubated with EDC/NHS, separated on a 947 SDS-polyacrylamide gel, and immunodetected with antibodies against TEF30. 948 949 Figure 2. Localization of TEF30 in thylakoids of Chlamydomonas wild-type and 950 mutant cells. 951 (A) Salt treatments during cell fractionation. Whole cells (WC) of strain cw15-325 were 952 resuspended in lysis buffer (LB) containing the indicated salts, subjected to 953 freeze/thawing, and separated into soluble (S) and pellet (P) fractions. One sample 954 resuspended in LB only was subjected to sonication instead of freeze/thawing. Proteins 955 were separated by SDS-PAGE and immunodetected with antibodies against TEF30 and 956 against integral membrane protein cytochrome f (Cytf) and stromal CGE1 as controls. 957 (B) Trypsin treatment of thylakoid membranes. Isolated thylakoids from strain cw15-325 958 were incubated on ice with trypsin, separated by SDS-PAGE and immunodetected with 959 antibodies against TEF30 and the peripheral CF1β subunit of the ATPase complex, 960 lumenal plastocyanin (PCY1) and integral membrane protein cytochrome f (Cytf) as 961 controls. 962 (C) Cell fractionation of various photosynthesis mutants. Whole cells (WC) of wild type 963 (WT) and thylakoid membrane protein mutants were separated into soluble (S) and 964 pellet (P) fractions via freeze/thawing. Proteins were separated by SDS-PAGE and 31 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. 965 immunodetected with antibodies against TEF30 and against integral membrane protein 966 cytochrome f (Cytf) or D1 and stromal CGE1 as controls. 967 968 Figure 3. Co-purification of TEF30 with PSII particles. 969 (A) Nickel-affinity purification of PSII particles. Detergent-solubilized thylakoid 970 membranes from a Chlamydomonas strain harboring a hexahistidine-tagged D2 protein 971 (D2His) and from wild-type control strain cc124 (WT) were subjected to nickel-affinity 972 purification. Thylakoid membrane input proteins corresponding to 1 µg chlorophyll and 973 4% of the imidazole eluate were separated by SDS-PAGE and immunodetected with 974 antibodies against TEF30 and proteins representative for the major thylakoid membrane 975 complexes. 976 (B) Silver staining of proteins immunoprecipitated from hexahistidine-tagged PSII 977 particles with preimmune serum (Pre) and antibodies against TEF30. 978 (C) Immunodetection of proteins immunopreciptated from PSII particles. Purified PSII 979 particles equivalent to ~1 µg chlorophyll (Input) and 20% of the immunoprecipitates 980 obtained with preimmune serum and antibodies against TEF30 (IP) were separated by 981 SDS-PAGE and immunodecorated with the antisera indicated. 982 983 Figure 4. Co-migration of TEF30 with PSII core subunits as revealed by sucrose 984 density gradient centrifugation and BN-PAGE. 985 (A) Thylakoid membranes from strain cw15-325 (1 mg chlorophyll/mL) were solubilized 986 with 1% β-DDM and separated on a linear 0.2 to 0.5 M sucrose gradient at 205,000 g for 987 16 h. Proteins were collected in 25 fractions, separated by SDS-PAGE and detected 988 with colloidal blue staining (upper panel) or immunologically with the antisera indicated 989 (lower panels). 990 (B) Analysis of thylakoid membrane protein complexes by sucrose density gradient 991 centrifugation as described in (A) was done on wild-type strain cc124, PSI mutant F23, 992 and PSII mutant ΔpsbA/D. 993 (C) Analysis of thylakoid membrane protein complexes by BN-PAGE. Thylakoids from 994 wild-type strain cc124, PSI mutant F23, and PSII mutant nac2 were solubilized with 1% 995 β-DDM and proteins corresponding to 5 µg chlorophyll separated on a 5-15% 996 polyacrylamide BN gradient gel, followed by Coomassie staining (left panel) and 32 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. 997 immunodetection with antibodies against TEF30 (right panel). Protein bands were 998 assigned to LHCII monomers (#1), LHCII trimers (#2), PSII monomers (#3), PSII II 999 dimers (#4), PSI supercomplexes (#5), and PSII supercomplexes (#6) according to Jarvi 1000 et al. (2011). 1001 1002 Figure 5. PSII activity and levels of PSII core subunits are more sensitive to high 1003 light intensities or low temperatures in TEF30-amiRNA cells than in control cells. 1004 (A) cw15-325 control (Con) and TEF30-amiRNA strains were photoinhibited (PI) for 60 1005 min at 1800 μmol photons m–2 s–1, and allowed to recover at 30 μmol photons m–2 s–1. 1006 PSII fluorescence was measured by pulse amplitude modulated fluorometry. Values 1007 represent the mean from three biological replicates, error bars indicate SD. 1008 (B) Control (Con) and TEF30-amiRNA #1.48 cells in the cw15-325 strain background 1009 were exposed to high PFDs (HL) of ~800 µmol photons m-2 s-1 for 6 h. Whole-cell 1010 proteins corresponding to 0.2, 1 or 2 µg chlorophyll (depending on the antiserum used) 1011 were separated on 10 or 14% SDS-polyacrylamide gels and analyzed by 1012 immunoblotting. 1013 (C) Control (Con) and TEF30-amiRNA #2.48 cells in the cw15-325 strain background 1014 were grown at a PFD of ~30 μmol photons m–2 s–1 and shifted from a growth 1015 temperature of 25°C to 10°C for 18 h. Samples were analyzed as in (B). 1016 1017 Figure 6. Comparison of LHCSR3 expression and oxygen evolution in control and 1018 TEF30-amiRNA cells. 1019 (A) Time course of LHCSR3 accumulation at 600 μmol photons m–2 s–1 (HL) in cw15- 1020 325 control and TEF30-amiRNA cells grown in TAP-NH4, TAP-NO3, or TMP-NH4. 1021 Whole-cell proteins were separated by SDS-PAGE und immunodetected with antibodies 1022 against LHCSR3 and CF1β as loading control. 1023 (B) Oxygen evolution from thylakoid membranes isolated from cw15-325 control and 1024 TEF30-amiRNA cells grown in TAP-NH4 medium at LL of ~30 µmol photons m-2 s-1 or 1025 HL of ~800 µmol photons m-2 s-1 for 4 h. Values represent the mean from experiments 1026 on two independent control and TEF30-amiRNA lines. Error bars indicate SD. (Note that 1027 the same thylakoid preparation was used for all PFDs applied, thus explaining the 1028 decline in O2 evolution at 1600 µmol photons m-2 s-1). 33 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. 1029 1030 Figure 7. PSII de novo synthesis is not impaired in TEF30-amiRNA strains. 1031 (A) TEF30-amiRNA and control cells in the cw15-325 background were grown in TAP- 1032 NH4 at ~30 µmol photons m-2 s-1 to exponential phase. Cells were then harvested, 1033 washed and resuspended in TAP medium lacking sulfur (-S). After 48 h under –S 1034 conditions, cells were shifted back to regular TAP medium (+S) for another 24 h. Whole- 1035 cell proteins from samples taken during this time course were separated by SDS-PAGE 1036 and PSII polypeptide abundance measured via immunoblotting using the antibodies 1037 indicated and CF1β as loading control. 1038 (B) D1 signal intensities from (A) were quantified using the FUSIONCapt Advance 1039 program. Signals were normalized to the initial D1 levels in each strain. Error bars 1040 represent SD, n = 2. 1041 (C) S-depletion was done as in (A), but time on –S and +S reduced to 24 h and 6 h, 1042 respectively. Thylakoids were solubilized with 1% α-DDM and proteins corresponding to 1043 5 µg chlorophyll separated on a 5-15% polyacrylamide BN gradient gel, followed by 1044 Coomassie staining (upper panel) and immunodetection with antibodies against D1 and 1045 CP43 (lower panels). Protein bands were assigned as in Figure 4C, the asterisk 1046 designates free CP43. 1047 (D) D1 signal intensities from protein bands #6 (PSII supercomplexes) and #4 (PSII 1048 dimers) from (C) were quantified using the FUSIONCapt Advance program. Signals 1049 were normalized to the initial D1 levels in each strain. Error bars represent SD, n = 4. 1050 1051 Figure 8. PSII stability is not impaired in TEF30-amiRNA strains. 1052 (A) TEF30-amiRNA and control cells in the cw15-325 background were grown at ~30 1053 µmol photons m-2 s-1 in TAP-medium in the presence or absence of 100 µg/mL CAP for 1054 12 h. Proteins from whole cells harvested during the time course corresponding to 0.5 or 1055 1 µg chlorophyll (depending on the antiserum used) were analyzed by immunoblotting. 1056 D1 levels were quantified with the FUSIONCapt Advance program. D1 levels were 1057 normalized to the values before the addition of CAP (at 0 h) and the ratio between D1 1058 levels from TEF30-amiRNA strains and controls calculated. Values represent the mean 1059 from four independent replicates, errors bars indicate SD. 34 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. 1060 (B) TEF30-amiRNA and control cells in the cw15-325 background were grown in TMP 1061 minimal medium at 200 μmol photons m–2 s–1 in the presence of 100 µg/mL CAP for 12 1062 h. Analyses were done as in (A) with two biological replicates. 1063 (C) TEF30-amiRNA and control cells in the cw15-325 background were grown in TAP- 1064 medium in the dark in the presence of 100 µg/mL CAP for 72 h. Analyses were done as 1065 in (A) with two biological replicates. 1066 1067 Figure 9. Not PSII sensitivity to high light, but PSII repair is impaired in TEF30- 1068 amiRNA strains. 1069 (A) Cells of cw15-325 control (Con) and TEF30-amiRNA strains were photoinhibited (PI) 1070 for 60 min at 1800 μmol photons m–2 s–1 in the presence of chloroplast translation 1071 inhibitors chloramphenicol and lincomycin, and allowed to recover at 30 μmol photons 1072 m–2 s–1 (LL) without inhibitors. PSII fluorescence was measured by pulse amplitude 1073 modulated fluorometry. Values represent the mean from two biological replicates, error 1074 bars indicate SD. 1075 (B) Immunoblot analysis of whole-cell proteins collected from the experiments described 1076 in (A). Results from a typical experiment are shown in the upper panel, the mean from 1077 quantified D1 signals (normalized relative to the initial D1 signal) are shown in the lower 1078 panel. Values represent the mean from five biological replicates, error bars indicate SD. 1079 1080 Figure 10. PSII supercomplex accumulation is impaired in TEF30-amiRNA strains 1081 exposed to high light. 1082 cw15-325 control (Con) and TEF30-amiRNA cells were grown at 30 µmol photons m-2 s-1 1083 (LL) and exposed to 800 µmol photons m-2 s-1 for 4 h (HL). Thylakoid membranes were 1084 isolated, 1085 electrophoretically on a 7.5% native Deriphat-polyacrylamide gel. The gel was then 1086 photographed (left panels) and processed for immunoblotting using antibodies against 1087 TEF30 and the D1 protein (right panels). Bands were assigned to free pigments (#0), 1088 antenna 1089 supercomplexes (#4) according to Formighieri et al. (2012). solubilized monomers with (#1), 1% LHCII α-DDM trimers and (#2), protein PSI/II complexes cores 1090 35 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. (#3), separated and PSI/II 1091 Figure 11. Cells of TEF30-amiRNA strains contain fewer thylakoid membranes per 1092 stack 1093 (A) Electron microscopy images of cw15-325 cells characteristic for the categories 1094 normally stacked, reduced stacking, and swollen. Overview images are shown on the 1095 left and a zoom-in of the region demarcated by the black box is shown on the right. N, 1096 nucleus; P, pyrenoid; S, starch. Bars in overview images correspond to 1 µm and those 1097 in zoom-ins to 0.2 µm. 1098 (B) cw15-325 control (Con) and TEF30-amiRNA cells were grown in TAP-medium at 1099 ~30 µmol photons m-2 s-1 (LL) and exposed to ~800 µmol photons m-2 s-1 (HL) for 4 h. 50 1100 electron micrographs each for two control strains and three independent TEF30-amiRNA 1101 strains were taken for each condition and thylakoid phenotypes assessed according to 1102 the categories shown in (A). Values are given in percent and represent mean values 1103 from the two control and three TEF30-amiRNA strains. Error bars indicate SD. 1104 Significance was tested using a t-test (*P < 0.05 and **P < 0.01). 1105 1106 Figure 12. A model for TEF30 function during PSII repair 1107 When PSII is damaged e.g. at high light intensities, the PSII complex enters the repair 1108 cycle. For this, PSII supercomplexes are disassembled and PSII monomers migrate 1109 from stacked into stroma-exposed membrane regions. After PSII monomers are partially 1110 disassembled to generate CP43-less forms, damaged D1 is degraded by FTSH and 1111 DEG proteases and a newly synthesized D1 copy is inserted into PSII monomers. At this 1112 step TEF30 may facilitate incorporation of the new D1 protein and/or reassembly of 1113 CP43, bind to repaired PSII monomers to protect them from undergoing repeated repair 1114 cycles, or facilitate the migration of repaired PSII monomers back to stacked regions for 1115 supercomplex reassembly. These functions are not mutually exclusive. 1116 1117 Literature Cited 1118 1119 1120 1121 Albiniak AM, Baglieri J, Robinson C (2012) Targeting of lumenal proteins across the thylakoid membrane. J Exp Bot 63: 1689-1698 Allan RK, Ratajczak T (2011) Versatile TPR domains accommodate different modes of target protein recognition and function. Cell Stress Chaperones 16: 353-367 36 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. 1122 1123 1124 1125 1126 1127 1128 1129 1130 1131 1132 1133 1134 1135 1136 1137 1138 1139 1140 1141 1142 1143 1144 1145 1146 1147 1148 1149 1150 1151 1152 1153 1154 1155 1156 1157 1158 1159 1160 1161 1162 1163 1164 1165 1166 1167 Allmer J, Naumann B, Markert C, Zhang M, Hippler M (2006) Mass spectrometric genomic data mining: Novel insights into bioenergetic pathways in Chlamydomonas reinhardtii. Proteomics 6: 6207-6220 Armbruster U, Labs M, Pribil M, Viola S, Xu W, Scharfenberg M, Hertle AP, Rojahn U, Jensen PE, Rappaport F, Joliot P, Dormann P, Wanner G, Leister D (2013) Arabidopsis CURVATURE THYLAKOID1 proteins modify thylakoid architecture by inducing membrane curvature. Plant Cell 25: 2661-2678 Aro EM, Suorsa M, Rokka A, Allahverdiyeva Y, Paakkarinen V, Saleem A, Battchikova N, Rintamaki E (2005) Dynamics of photosystem II: a proteomic approach to thylakoid protein complexes. J Exp Bot 56: 347-356 Aro EM, Virgin I, Andersson B (1993) Photoinhibition of Photosystem II. Inactivation, protein damage and turnover. Biochim Biophys Acta 1143: 113-134 Bhuiyan NH, Friso G, Poliakov A, Ponnala L, van Wijk KJ (2015) MET1 is a thylakoid-associated TPR protein involved in photosystem II supercomplex formation and repair in Arabidopsis. Plant Cell 27: 262-285 Bonente G, Ballottari M, Truong TB, Morosinotto T, Ahn TK, Fleming GR, Niyogi KK, Bassi R (2011) Analysis of LhcSR3, a protein essential for feedback deexcitation in the green alga Chlamydomonas reinhardtii. PLoS Biol 9: e1000577 Caffarri S, Kouril R, Kereiche S, Boekema EJ, Croce R (2009) Functional architecture of higher plant photosystem II supercomplexes. EMBO J 28: 3052-3063 Cai W, Ma J, Chi W, Zou M, Guo J, Lu C, Zhang L (2010) Cooperation of LPA3 and LPA2 is essential for photosystem II assembly in Arabidopsis. Plant Physiol 154: 109-120 Cai W, Ma J, Guo J, Zhang L (2008) Function of ROC4 in the efficient repair of photodamaged photosystem II in Arabidopsis. Photochem Photobiol 84: 13431348 Che YF, Fu AG, Hou X, McDonald K, Buchanan BB, Huang WD, Luan S (2013) Cterminal processing of reaction center protein D1 is essential for the function and assembly of photosystem II in Arabidopsis. Proc Natl Acad Sci U S A 110: 1624716252 Chua NH, Bennoun P (1975) Thylakoid membrane polypeptides of Chlamydomonas reinhardtii: wild-type and mutant strains deficient in photosystem II reaction center. Proc Natl Acad Sci U S A 72: 2175-2179 Cullen M, Ray N, Husain S, Nugent J, Nield J, Purton S (2007) A highly active histidine-tagged Chlamydomonas reinhardtii Photosystem II preparation for structural and biophysical analysis. Photochem Photobiol Sci 6: 1177-1183 Danielsson R, Suorsa M, Paakkarinen V, Albertsson PA, Styring S, Aro EM, Mamedov F (2006) Dimeric and monomeric organization of photosystem II. Distribution of five distinct complexes in the different domains of the thylakoid membrane. J Biol Chem 281: 14241-14249 de Vitry C, Olive J, Drapier D, Recouvreur M, Wollman FA (1989) Posttranslational events leading to the assembly of photosystem II protein complex: a study using photosynthesis mutants from Chlamydomonas reinhardtii. J Cell Biol 109: 9911006 Drop B, Webber-Birungi M, Yadav SK, Filipowicz-Szymanska A, Fusetti F, Boekema EJ, Croce R (2014) Light-harvesting complex II (LHCII) and its 37 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. 1168 1169 1170 1171 1172 1173 1174 1175 1176 1177 1178 1179 1180 1181 1182 1183 1184 1185 1186 1187 1188 1189 1190 1191 1192 1193 1194 1195 1196 1197 1198 1199 1200 1201 1202 1203 1204 1205 1206 1207 1208 1209 1210 1211 1212 1213 1214 supramolecular organization in Chlamydomonas reinhardtii. Biochim Biophys Acta 1837: 63-72 Emanuelsson O, Nielsen H, Brunak S, von Heijne G (2000) Predicting subcellular localization of proteins based on their N-terminal amino acid sequence. J Mol Biol 300: 1005-1016 Emanuelsson O, Nielsen H, von Heijne G (1999) ChloroP, a neural network-based method for predicting chloroplast transit peptides and their cleavage sites. Protein Sci 8: 978-984 Erickson JM, Rahire M, Malnoe P, Girard-Bascou J, Pierre Y, Bennoun P, Rochaix JD (1986) Lack of the D2 protein in a Chlamydomonas reinhardtii psbD mutant affects photosystem II stability and D1 expression. EMBO J 5: 1745-1754 Eriksson M, Gardestrom P, Samuelsson G (1995) Isolation, purification, and characterization of mitochondria from Chlamydomonas reinhardtii. Plant Physiol 107: 479-483 Formighieri C, Ceol M, Bonente G, Rochaix JD, Bassi R (2012) Retrograde signaling and photoprotection in a gun4 mutant of Chlamydomonas reinhardtii. Mol Plant 5: 1242-1262 Fristedt R, Willig A, Granath P, Crevecoeur M, Rochaix JD, Vener AV (2009) Phosphorylation of photosystem II controls functional macroscopic folding of photosynthetic membranes in Arabidopsis. Plant Cell 21: 3950-3964 Girard J, Chua NH, Bennoun P, Schmidt G, Delosme M (1980) Studies on mutants deficient in the photosystem I reaction centers in Chlamydomonas reinhardtii. Curr Genet 2: 215-221 Goral TK, Johnson MP, Brain AP, Kirchhoff H, Ruban AV, Mullineaux CW (2010) Visualizing the mobility and distribution of chlorophyll proteins in higher plant thylakoid membranes: effects of photoinhibition and protein phosphorylation. Plant J 62: 948-959 Greer DH, Laing WA, Kipnis T (1988) Photoinhibition of photosynthesis in intact kiwifruit (Actinidia deliciosa) leaves: Effect of temperature. Planta 174: 152-158 Guillot-Salomon T, Tuquet C, De Lubac M, Hallais MF, Signol M (1978) [Comparative analysis of ultrastructure and lipid composition of plastids from sun and shade plants (author's transl)]. Cytobiologie 17: 442-452 Harris EH (2008) The Chlamydomonas Sourcebook: Introduction to Chlamydomonas and Its Laboratory Use. In HH Elizabeth, Ph.D, BS David, Ph.D, PD George B. Witman, eds, The Chlamydomonas Sourcebook (Second Edition). Elsevier/ Academic Press, San Diego, CA Haussuhl K, Andersson B, Adamska I (2001) A chloroplast DegP2 protease performs the primary cleavage of the photodamaged D1 protein in plant photosystem II. EMBO J 20: 713-722 Heinnickel ML, Grossman AR (2013) The GreenCut: re-evaluation of physiological role of previously studied proteins and potential novel protein functions. Photosynth Res 116: 427-436 Hemme D, Veyel D, Muhlhaus T, Sommer F, Juppner J, Unger AK, Sandmann M, Fehrle I, Schonfelder S, Steup M, Geimer S, Kopka J, Giavalisco P, Schroda M (2014) Systems-wide analysis of acclimation responses to long-term heat stress and recovery in the photosynthetic model organism Chlamydomonas reinhardtii. Plant Cell 26: 4270-4297 38 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. 1215 1216 1217 1218 1219 1220 1221 1222 1223 1224 1225 1226 1227 1228 1229 1230 1231 1232 1233 1234 1235 1236 1237 1238 1239 1240 1241 1242 1243 1244 1245 1246 1247 1248 1249 1250 1251 1252 1253 1254 1255 1256 1257 1258 1259 1260 Herbstova M, Tietz S, Kinzel C, Turkina MV, Kirchhoff H (2012) Architectural switch in plant photosynthetic membranes induced by light stress. Proc Natl Acad Sci U S A 109: 20130-20135 Huang W, Chen Q, Zhu Y, Hu F, Zhang L, Ma Z, He Z, Huang J (2013) Arabidopsis thylakoid formation 1 is a critical regulator for dynamics of PSII-LHCII complexes in leaf senescence and excess light. Mol Plant 6: 1673-1691 Ishikawa A, Tanaka H, Kato C, Iwasaki Y, Asahi T (2005) Molecular characterization of the ZKT gene encoding a protein with PDZ, K-Box, and TPR motifs in Arabidopsis. Biosci Biotechnol Biochem 69: 972-978 Jarvi S, Suorsa M, Aro EM (2015) Photosystem II repair in plant chloroplasts Regulation, assisting proteins and shared components with photosystem II biogenesis. Biochim Biophys Acta Jarvi S, Suorsa M, Paakkarinen V, Aro EM (2011) Optimized native gel systems for separation of thylakoid protein complexes: novel super- and mega-complexes. Biochem J 439: 207-214 Johnson X, Vandystadt G, Bujaldon S, Wollman FA, Dubois R, Roussel P, Alric J, Beal D (2009) A new setup for in vivo fluorescence imaging of photosynthetic activity. Photosynth Res 102: 85-93 Kapri-Pardes E, Naveh L, Adam Z (2007) The thylakoid lumen protease Deg1 is involved in the repair of photosystem II from photoinhibition in Arabidopsis. Plant Cell 19: 1039-1047 Karpowicz SJ, Prochnik SE, Grossman AR, Merchant SS (2011) The GreenCut2 resource, a phylogenomically derived inventory of proteins specific to the plant lineage. J Biol Chem 286: 21427-21439 Kato Y, Sakamoto W (2014) Phosphorylation of photosystem II core proteins prevents undesirable cleavage of D1 and contributes to the fine-tuned repair of photosystem II. Plant J 79: 312-321 Keren N, Ohkawa H, Welsh EA, Liberton M, Pakrasi HB (2005) Psb29, a conserved 22-kD protein, functions in the biogenesis of Photosystem II complexes in Synechocystis and Arabidopsis. Plant Cell 17: 2768-2781 Kindle KL (1990) High-frequency nuclear transformation of Chlamydomonas reinhardtii. Proc Natl Acad Sci U S A 87: 1228-1232 Kirchhoff H (2013) Diffusion of molecules and macromolecules in thylakoid membranes. Biochim Biophys Acta Kirchhoff H, Sharpe RM, Herbstova M, Yarbrough R, Edwards GE (2013) Differential mobility of pigment-protein complexes in granal and agranal thylakoid membranes of C(3) and C(4) plants. Plant Physiol 161: 497-507 Koivuniemi A, Aro EM, Andersson B (1995) Degradation of the D1- and D2-proteins of photosystem II in higher plants is regulated by reversible phosphorylation. Biochemistry 34: 16022-16029 Komenda J, Sobotka R, Nixon PJ (2012) Assembling and maintaining the Photosystem II complex in chloroplasts and cyanobacteria. Curr Opin Plant Biol 15: 245-251 Kosourov S, Makarova V, Fedorov AS, Tsygankov A, Seibert M, Ghirardi ML (2005) The effect of sulfur re-addition on H(2) photoproduction by sulfur-deprived green algae. Photosynth Res 85: 295-305 39 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. 1261 1262 1263 1264 1265 1266 1267 1268 1269 1270 1271 1272 1273 1274 1275 1276 1277 1278 1279 1280 1281 1282 1283 1284 1285 1286 1287 1288 1289 1290 1291 1292 1293 1294 1295 1296 1297 1298 1299 1300 1301 1302 1303 1304 1305 Kouril R, Oostergetel GT, Boekema EJ (2011) Fine structure of granal thylakoid membrane organization using cryo electron tomography. Biochim Biophys Acta 1807: 368-374 Kouril R, Wientjes E, Bultema JB, Croce R, Boekema EJ (2013) High-light vs. lowlight: effect of light acclimation on photosystem II composition and organization in Arabidopsis thaliana. Biochim Biophys Acta 1827: 411-419 Kropat J, Hong-Hermesdorf A, Casero D, Ent P, Castruita M, Pellegrini M, Merchant SS, Malasarn D (2011) A revised mineral nutrient supplement increases biomass and growth rate in Chlamydomonas reinhardtii. Plant J 66: 770-780 Kuchka MR, Goldschmidt-Clermont M, van Dillewijn J, Rochaix JD (1989) Mutation at the Chlamydomonas nuclear NAC2 locus specifically affects stability of the chloroplast psbD transcript encoding polypeptide D2 of PS II. Cell 58: 869-876 Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227: 680-685 Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23: 2947-2948 Lemaire C, Wollman FA (1989) The chloroplast ATP synthase in Chlamydomonas reinhardtii. I. Characterization of its nine constitutive subunits. J Biol Chem 264: 10228-10234 Li XG, Duan W, Meng QW, Zou Q, Zhao SJ (2004) The function of chloroplastic NAD(P)H dehydrogenase in tobacco during chilling stress under low irradiance. Plant Cell Physiol 45: 103-108 Lichtenthaler HK, Buschmann C, Doll M, Fietz HJ, Bach T, Kozel U, Meier D, Rahmsdorf U (1981) Photosynthetic activity, chloroplast ultrastructure, and leaf characteristics of high-light and low-light plants and of sun and shade leaves. Photosynth Res 2: 115-141 Lindahl M, Spetea C, Hundal T, Oppenheim AB, Adam Z, Andersson B (2000) The thylakoid FtsH protease plays a role in the light-induced turnover of the photosystem II D1 protein. Plant Cell 12: 419-431 Lippuner V, Chou IT, Scott SV, Ettinger WF, Theg SM, Gasser CS (1994) Cloning and characterization of chloroplast and cytosolic forms of cyclophilin from Arabidopsis thaliana. J Biol Chem 269: 7863-7868 Liu C, Willmund F, Whitelegge JP, Hawat S, Knapp B, Lodha M, Schroda M (2005) J-domain protein CDJ2 and HSP70B are a plastidic chaperone pair that interacts with vesicle-inducing protein in plastids 1. Mol Biol Cell 16: 1165-1177 Lowry OH, Rosebrough NJ, Farr AL, Randall RJ (1951) Protein measurement with the Folin phenol reagent. J Biol Chem 193: 265-275 Malnoe A, Wang F, Girard-Bascou J, Wollman FA, de Vitry C (2014) Thylakoid FtsH protease contributes to photosystem II and cytochrome b6f remodeling in Chlamydomonas reinhardtii under stress conditions. Plant Cell 26: 373-390 Maruyama S, Tokutsu R, Minagawa J (2014) Transcriptional regulation of the stressresponsive light harvesting complex genes in Chlamydomonas reinhardtii. Plant Cell Physiol 55: 1304-1310 40 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. 1306 1307 1308 1309 1310 1311 1312 1313 1314 1315 1316 1317 1318 1319 1320 1321 1322 1323 1324 1325 1326 1327 1328 1329 1330 1331 1332 1333 1334 1335 1336 1337 1338 1339 1340 1341 1342 1343 1344 1345 1346 1347 1348 1349 1350 1351 1352 Mayfield SP, Bennoun P, Rochaix JD (1987) Expression of the nuclear encoded Oee1 protein is required for oxygen evolution and stability of Photosystem-II particles in Chlamydomonas reinhardtii. EMBO J 6: 313-318 Merchant SS, Prochnik SE, Vallon O, Harris EH, Karpowicz SJ, Witman GB, Terry A, Salamov A, Fritz-Laylin LK, Marechal-Drouard L, Marshall WF, Qu LH, Nelson DR, Sanderfoot AA, Spalding MH, Kapitonov VV, Ren Q, Ferris P, Lindquist E, Shapiro H, Lucas SM, Grimwood J, Schmutz J, Cardol P, Cerutti H, Chanfreau G, Chen CL, Cognat V, Croft MT, Dent R, Dutcher S, Fernandez E, Fukuzawa H, Gonzalez-Ballester D, Gonzalez-Halphen D, Hallmann A, Hanikenne M, Hippler M, Inwood W, Jabbari K, Kalanon M, Kuras R, Lefebvre PA, Lemaire SD, Lobanov AV, Lohr M, Manuell A, Meier I, Mets L, Mittag M, Mittelmeier T, Moroney JV, Moseley J, Napoli C, Nedelcu AM, Niyogi K, Novoselov SV, Paulsen IT, Pazour G, Purton S, Ral JP, RianoPachon DM, Riekhof W, Rymarquis L, Schroda M, Stern D, Umen J, Willows R, Wilson N, Zimmer SL, Allmer J, Balk J, Bisova K, Chen CJ, Elias M, Gendler K, Hauser C, Lamb MR, Ledford H, Long JC, Minagawa J, Page MD, Pan J, Pootakham W, Roje S, Rose A, Stahlberg E, Terauchi AM, Yang P, Ball S, Bowler C, Dieckmann CL, Gladyshev VN, Green P, Jorgensen R, Mayfield S, Mueller-Roeber B, Rajamani S, Sayre RT, Brokstein P, Dubchak I, Goodstein D, Hornick L, Huang YW, Jhaveri J, Luo Y, Martinez D, Ngau WC, Otillar B, Poliakov A, Porter A, Szajkowski L, Werner G, Zhou K, Grigoriev IV, Rokhsar DS, Grossman AR (2007) The Chlamydomonas genome reveals the evolution of key animal and plant functions. Science 318: 245-250 Mettler T, Muhlhaus T, Hemme D, Schottler MA, Rupprecht J, Idoine A, Veyel D, Pal SK, Yaneva-Roder L, Winck FV, Sommer F, Vosloh D, Seiwert B, Erban A, Burgos A, Arvidsson S, Schonfelder S, Arnold A, Gunther M, Krause U, Lohse M, Kopka J, Nikoloski Z, Mueller-Roeber B, Willmitzer L, Bock R, Schroda M, Stitt M (2014) Systems analysis of the response of photosynthesis, metabolism, and growth to an increase in irradiance in the photosynthetic model organism Chlamydomonas reinhardtii. Plant Cell 26: 2310-2350 Minai L, Wostrikoff K, Wollman FA, Choquet Y (2006) Chloroplast biogenesis of photosystem II cores involves a series of assembly-controlled steps that regulate translation. Plant Cell 18: 159-175 Molnar A, Bassett A, Thuenemann E, Schwach F, Karkare S, Ossowski S, Weigel D, Baulcombe D (2009) Highly specific gene silencing by artificial microRNAs in the unicellular alga Chlamydomonas reinhardtii. Plant J 58: 165-174 Mulo P, Sakurai I, Aro EM (2012) Strategies for psbA gene expression in cyanobacteria, green algae and higher plants: from transcription to PSII repair. Biochim Biophys Acta 1817: 247-257 Nath K, Jajoo A, Poudyal RS, Timilsina R, Park YS, Aro EM, Nam HG, Lee CH (2013a) Towards a critical understanding of the photosystem II repair mechanism and its regulation during stress conditions. FEBS Lett 587: 3372-3381 Nath K, Poudyal RS, Eom JS, Park YS, Zulfugarov IS, Mishra SR, Tovuu A, Ryoo N, Yoon HS, Nam HG, An G, Jeon JS, Lee CH (2013b) Loss-of-function of OsSTN8 suppresses the photosystem II core protein phosphorylation and interferes with the photosystem II repair mechanism in rice (Oryza sativa). Plant J 76: 675-686 41 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. 1353 1354 1355 1356 1357 1358 1359 1360 1361 1362 1363 1364 1365 1366 1367 1368 1369 1370 1371 1372 1373 1374 1375 1376 1377 1378 1379 1380 1381 1382 1383 1384 1385 1386 1387 1388 1389 1390 1391 1392 1393 1394 1395 1396 1397 1398 1399 Naumann B, Busch A, Allmer J, Ostendorf E, Zeller M, Kirchhoff H, Hippler M (2007) Comparative quantitative proteomics to investigate the remodeling of bioenergetic pathways under iron deficiency in Chlamydomonas reinhardtii. Proteomics 7: 3964-3979 Naumann B, Stauber EJ, Busch A, Sommer F, Hippler M (2005) N-terminal processing of Lhca3 Is a key step in remodeling of the photosystem I-lightharvesting complex under iron deficiency in Chlamydomonas reinhardtii. J Biol Chem 280: 20431-20441 Nickelsen J, Rengstl B (2013) Photosystem II assembly: From cyanobacteria to plants. Annu Rev Plant Biol 64: 609-635 Nixon PJ, Michoux F, Yu J, Boehm M, Komenda J (2010) Recent advances in understanding the assembly and repair of photosystem II. Ann Bot 106: 1-16 Nordhues A, Schottler MA, Unger AK, Geimer S, Schonfelder S, Schmollinger S, Rutgers M, Finazzi G, Soppa B, Sommer F, Muhlhaus T, Roach T, KriegerLiszkay A, Lokstein H, Crespo JL, Schroda M (2012) Evidence for a role of VIPP1 in the structural organization of the photosynthetic apparatus in Chlamydomonas. Plant Cell 24: 637-659 Ossowski S, Schwab R, Weigel D (2008) Gene silencing in plants using artificial microRNAs and other small RNAs. Plant J 53: 674-690 Pagliano C, Saracco G, Barber J (2013) Structural, functional and auxiliary proteins of photosystem II. Photosynth Res 116: 167-188 Peers G, Truong TB, Ostendorf E, Busch A, Elrad D, Grossman AR, Hippler M, Niyogi KK (2009) An ancient light-harvesting protein is critical for the regulation of algal photosynthesis. Nature 462: 518-521 Peter GF, Thornber JP (1991) Biochemical composition and organization of higherplant Photosystem-II light-harvesting pigment-proteins. J Biol Chem 266: 1674516754 Petroutsos D, Busch A, Janssen I, Trompelt K, Bergner SV, Weinl S, Holtkamp M, Karst U, Kudla J, Hippler M (2011) The chloroplast calcium sensor CAS is required for photoacclimation in Chlamydomonas reinhardtii. Plant Cell 23: 29502963 Pierre Y, Popot JL (1993) Identification of two 4-kDa miniproteins in the cytochrome b6f complex from Chlamydomonas reinhardtii. C R Acad Sci III 316: 1404-1409 Ramundo S, Casero D, Muhlhaus T, Hemme D, Sommer F, Crevecoeur M, Rahire M, Schroda M, Rusch J, Goodenough U, Pellegrini M, Perez-Perez ME, Crespo JL, Schaad O, Civic N, Rochaix JD (2014) Conditional depletion of the Chlamydomonas chloroplast ClpP protease activates nuclear genes involved in autophagy and plastid protein quality control. Plant Cell 26: 2201-2222 Rimbault B, Esposito D, Drapier D, Choquet Y, Stern D, Wollman FA (2000) Identification of the initiation codon for the atpB gene in Chlamydomonas chloroplasts excludes translation of a precursor form of the beta subunit of the ATP synthase. Mol Gen Genet 264: 486-491 Rintamaki E, Kettunen R, Aro EM (1996) Differential D1 dephosphorylation in functional and photodamaged photosystem II centers. Dephosphorylation is a prerequisite for degradation of damaged D1. J Biol Chem 271: 14870-14875 Rochaix JD, Kuchka M, Mayfield S, Schirmerrahire M, Girardbascou J, Bennoun P (1989) Nuclear and chloroplast mutations affect the synthesis or stability of the 42 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. 1400 1401 1402 1403 1404 1405 1406 1407 1408 1409 1410 1411 1412 1413 1414 1415 1416 1417 1418 1419 1420 1421 1422 1423 1424 1425 1426 1427 1428 1429 1430 1431 1432 1433 1434 1435 1436 1437 1438 1439 1440 1441 1442 1443 1444 1445 chloroplast psbC gene-product in Chlamydomonas reinhardtii. EMBO J 8: 10131021 Rutgers M, Schroda M (2013) A role of VIPP1 as a dynamic structure within thylakoid centers as sites of photosystem biogenesis? Plant Signal Behav 8: e27037 Samol I, Shapiguzov A, Ingelsson B, Fucile G, Crevecoeur M, Vener AV, Rochaix JD, Goldschmidt-Clermont M (2012) Identification of a photosystem II phosphatase involved in light acclimation in Arabidopsis. Plant Cell 24: 25962609 Schagger H, Cramer WA, von Jagow G (1994) Analysis of molecular masses and oligomeric states of protein complexes by blue native electrophoresis and isolation of membrane protein complexes by two-dimensional native electrophoresis. Anal Biochem 217: 220-230 Schagger H, von Jagow G (1991) Blue native electrophoresis for isolation of membrane protein complexes in enzymatically active form. Anal Biochem 199: 223-231 Schmollinger S, Muhlhaus T, Boyle NR, Blaby IK, Casero D, Mettler T, Moseley JL, Kropat J, Sommer F, Strenkert D, Hemme D, Pellegrini M, Grossman AR, Stitt M, Schroda M, Merchant SS (2014) Nitrogen-sparing mechanisms in Chlamydomonas affect the transcriptome, the proteome, and photosynthetic metabolism. Plant Cell 26: 1410-1435 Schroda M, Vallon O, Whitelegge JP, Beck CF, Wollman FA (2001) The chloroplastic GrpE homolog of Chlamydomonas: Two isoforms generated by differential splicing. Plant Cell 13: 2823-2839 Schroda M, Vallon O, Wollman FA, Beck CF (1999) A chloroplast-targeted heat shock protein 70 (HSP70) contributes to the photoprotection and repair of photosystem II during and after photoinhibition. Plant Cell 11: 1165-1178 Schulz-Raffelt M, Lodha M, Schroda M (2007) Heat shock factor 1 is a key regulator of the stress response in Chlamydomonas. Plant J 52: 286-295 Silva P, Thompson E, Bailey S, Kruse O, Mullineaux CW, Robinson C, Mann NH, Nixon PJ (2003) FtsH is involved in the early stages of repair of photosystem II in Synechocystis sp PCC 6803. Plant Cell 15: 2152-2164 Sirpio S, Allahverdiyeva Y, Suorsa M, Paakkarinen V, Vainonen J, Battchikova N, Aro EM (2007) TLP18.3, a novel thylakoid lumen protein regulating photosystem II repair cycle. Biochem J 406: 415-425 Sonnhammer EL, von Heijne G, Krogh A (1998) A hidden Markov model for predicting transmembrane helices in protein sequences. Proc Int Conf Intell Syst Mol Biol 6: 175-182 Strenkert D, Schmollinger S, Sommer F, Schulz-Raffelt M, Schroda M (2011) Transcription factor dependent chromatin remodeling at heat shock and copper responsive promoters in Chlamydomonas reinhardtii. Plant Cell 23: 2285-2301 Sugiura M, Inoue Y, Minagawa J (1998) Rapid and discrete isolation of oxygenevolving His-tagged photosystem II core complex from Chlamydomonas reinhardtii by Ni2+ affinity column chromatography. FEBS Lett 426: 140-144 Sun X, Fu T, Chen N, Guo J, Ma J, Zou M, Lu C, Zhang L (2010) The stromal chloroplast Deg7 protease participates in the repair of photosystem II after photoinhibition in Arabidopsis. Plant Physiol 152: 1263-1273 43 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. 1446 1447 1448 1449 1450 1451 1452 1453 1454 1455 1456 1457 1458 1459 1460 1461 1462 1463 1464 1465 1466 1467 1468 1469 1470 1471 1472 1473 1474 1475 1476 1477 1478 1479 1480 1481 1482 1483 1484 1485 1486 1487 1488 1489 1490 1491 Takahashi S, Murata N (2005) Interruption of the Calvin cycle inhibits the repair of Photosystem II from photodamage. Biochim Biophys Acta 1708: 352-361 Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28: 2731-2739 Tietz S, Puthiyaveetil S, Enlow HM, Yarbrough R, Wood M, Semchonok DA, Lowry T, Li Z, Jahns P, Boekema EJ, Lenhert S, Niyogi KK, Kirchhoff H (2015) Functional implications of Photosystem II crystal formation in photosynthetic membranes. J Biol Chem 290: 14091-14106 Tikkanen M, Nurmi M, Kangasjarvi S, Aro EM (2008) Core protein phosphorylation facilitates the repair of photodamaged photosystem II at high light. Biochim Biophys Acta 1777: 1432-1437 Tokutsu R, Kato N, Bui KH, Ishikawa T, Minagawa J (2012) Revisiting the supramolecular organization of photosystem II in Chlamydomonas reinhardtii. J Biol Chem 287: 31574-31581 Tokutsu R, Minagawa J (2013) Energy-dissipative supercomplex of photosystem II associated with LHCSR3 in Chlamydomonas reinhardtii. Proc Natl Acad Sci U S A 110: 10016-10021 Tomizioli M, Lazar C, Brugiere S, Burger T, Salvi D, Gatto L, Moyet L, Breckels LM, Hesse AM, Lilley KS, Seigneurin-Berny D, Finazzi G, Rolland N, Ferro M (2014) Deciphering thylakoid sub-compartments using a mass spectrometrybased approach. Mol Cell Proteomics 13: 2147-2167 Tyystjarvi E (2013) Photoinhibition of Photosystem II. Int Rev Cell Mol Biol 300: 243303 Tyystjarvi E, Aro EM (1996) The rate constant of photoinhibition, measured in lincomycin-treated leaves, is directly proportional to light intensity. Proc Natl Acad Sci U S A 93: 2213-2218 Umena Y, Kawakami K, Shen JR, Kamiya N (2011) Crystal structure of oxygenevolving photosystem II at a resolution of 1.9 A. Nature 473: 55-60 Uniacke J, Colon-Ramos D, Zerges W (2011) FISH and immunofluorescence staining in Chlamydomonas. Methods Mol Biol 714: 15-29 Uniacke J, Zerges W (2007) Photosystem II assembly and repair are differentially localized in Chlamydomonas. Plant Cell 19: 3640-3654 Vallon O, Wollman FA, Olive J (1985) Distribution of intrinsic and extrinsic subunits of the PSII protein complex between appressed and non-appressed regions of the thylakoid membrane: an immunocytochemical study. FEBS Lett 183: 245-250 Vallon O, Wollman FA, Olive J (1986) Lateral distribution of the main protein complexes of the photosynthetic appartus in Chlamydomonas reinhardttii and in spinach: an immunocytochemical study using inteact thylakoid membranes and PS II enriched membrane preparation. Photobiochem Photobiophys 12: 203-220 Wang Q, Sullivan RW, Kight A, Henry RL, Huang J, Jones AM, Korth KL (2004) Deletion of the chloroplast-localized Thylakoid formation1 gene product in Arabidopsis leads to deficient thylakoid formation and variegated leaves. Plant Physiol 136: 3594-3604 Wittenberg G, Levitan A, Klein T, Dangoor I, Keren N, Danon A (2014) Knockdown of the Arabidopsis thaliana chloroplast protein disulfide isomerase 6 results in 44 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. 1492 1493 1494 1495 1496 1497 1498 1499 1500 1501 1502 1503 1504 1505 1506 1507 1508 1509 1510 1511 1512 1513 1514 1515 1516 1517 1518 1519 1520 1521 1522 1523 1524 1525 reduced levels of photoinhibition and increased D1 synthesis in high light. Plant J 78: 1003-1013 Woessner JP, Gillham NW, Boynton JE (1986) The sequence of the chloroplast atpB gene and its flanking regions in Chlamydomonas reinhardtii. Gene 44: 17-28 Wu HY, Liu MS, Lin TP, Cheng YS (2011) Structural and functional assays of AtTLP18.3 identify its novel acid phosphatase activity in thylakoid lumen. Plant Physiol 157: 1015-1025 Wunder T, Xu W, Liu Q, Wanner G, Leister D, Pribil M (2013) The major thylakoid protein kinases STN7 and STN8 revisited: effects of altered STN8 levels and regulatory specificities of the STN kinases. Front Plant Sci 4: 417 Xue H, Tokutsu R, Bergner SV, Scholz M, Minagawa J, Hippler M (2015) Photosystem II subunit R is required for efficient binding of light-harvesting complex stress-related protein 3 to Photosystem II-light-harvesting supercomplexes in Chlamydomonas reinhardtii. Plant Physiol 167: 1566-1578 Yamatani H, Sato Y, Masuda Y, Kato Y, Morita R, Fukunaga K, Nagamura Y, Nishimura M, Sakamoto W, Tanaka A, Kusaba M (2013) NYC4, the rice ortholog of Arabidopsis THF1, is involved in the degradation of chlorophyll protein complexes during leaf senescence. Plant J 74: 652-662 Ye F, Zhang M (2013) Structures and target recognition modes of PDZ domains: recurring themes and emerging pictures. Biochem J 455: 1-14 Yokthongwattana K, Chrost B, Behrman S, Casper-Lindley C, Melis A (2001) Photosystem II damage and repair cycle in the green alga Dunaliella salina: involvement of a chloroplast-localized HSP70. Plant Cell Physiol 42: 1389-1397 Zerges W, Rochaix JD (1998) Low density membranes are associated with RNAbinding proteins and thylakoids in the chloroplast of Chlamydomonas reinhardtii. J Cell Biol 140: 101-110 Zhang LX, Paakkarinen V, van Wijk KJ, Aro EM (1999) Co-translational assembly of the D1 protein into photosystem II. J Biol Chem 274: 16062-16067 Zhang LX, Paakkarinen V, van Wijk KJ, Aro EM (2000) Biogenesis of the chloroplastencoded D1 protein: Regulation of translation elongation, insertion, and assembly into photosystem II. Plant Cell 12: 1769-1781 Zienkiewicz M, Ferenc A, Wasilewska W, Romanowska E (2012) High light stimulates Deg1-dependent cleavage of the minor LHCII antenna proteins CP26 and CP29 and the PsbS protein in Arabidopsis thaliana. Planta 235: 279-288 1526 45 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. Figure 1. Antibody characterization and analysis of TEF30 oligomer formation, cellular abundance and intracellular localization. (A) Immunodetection of TEF30 in cw15-325 whole-cell proteins (WC) corresponding to 2 µg chlorophyll compared with 0.25 ng recombinant TEF30 protein. (B) Subcellular localization of TEF30 by immunoblotting. 10 or 3 µg (depending on the antiserum used) protein from whole cells (WC), chloroplasts (cp) and mitochondria (mt) isolated from strain cw15-302 were separated by SDSPAGE and immunodetected with antisera against TEF30, mitochondrial carboanhydrase (mtCA), and stromal HSP70B. (C) Microscopy images taken from strain cw15-325. Shown are from left to right: DIC images, DAPI staining, immunofluorescence, merge of DAPI and immunofluorescence, and a schematic drawing that combines information from the images. Antisera used for immunofluorescence were against HSP70B and HSP70A as markers for stroma and cytosol, respectively, and against TEF30. Chloroplast – cp; pyrenoid – P; cytosol – C; flagella – f; immunofluorescence signal – green. (D) 1 to 8 ng of recombinant proteins TEF30 (purified from E. coli) and D2 (purchased from Agrisera) were separated by SDS-PAGE together with cw15-325 whole-cell proteins corresponding to 0.125-2 µg chlorophyll and immunodetected with antibodies against TEF30 and D2. (E) 100 ng of recombinant TEF30 protein was incubated with EDC/NHS, separated on a SDS-polyacrylamide gel, and immunodetected with antibodies against TEF30. Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. Figure 2. Localization of TEF30 in thylakoids of Chlamydomonas wild-type and mutant cells. (A) Salt treatments during cell fractionation. Whole cells (WC) of strain cw15-325 were resuspended in lysis buffer (LB) containing the indicated salts, subjected to freeze/thawing, and separated into soluble (S) and pellet (P) fractions. One sample resuspended in LB only was subjected to sonication instead of freeze/thawing. Proteins were separated by SDS-PAGE and immunodetected with antibodies against TEF30 and against integral membrane protein cytochrome f (Cytf) and stromal CGE1 as controls. (B) Trypsin treatment of thylakoid membranes. Isolated thylakoids from strain cw15-325 were incubated on ice with trypsin, separated by SDS-PAGE and immunodetected with antibodies against TEF30 and the peripheral CF1β subunit of the ATPase complex, lumenal plastocyanin (PCY1) and integral membrane protein cytochrome f (Cytf) as controls. (C) Cell fractionation of various photosynthesis mutants. Whole cells (WC) of wild type (WT) and thylakoid membrane protein mutants were separated into soluble (S) and pellet (P) fractions via freeze/thawing. Proteins were separated by SDS-PAGE and immunodetected with antibodies against TEF30 and against integral membrane protein cytochrome f (Cytf) or D1 and stromal CGE1 as controls. Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. Figure 3. Co-purification of TEF30 with PSII particles. (A) Nickel-affinity purification of PSII particles. Detergentsolubilized thylakoid membranes from a Chlamydomonas strain harboring a hexahistidine-tagged D2 protein (D2His) and from wild-type control strain cc124 (WT) were subjected to nickel-affinity purification. Thylakoid membrane input proteins corresponding to 1 µg chlorophyll and 4% of the imidazole eluate were separated by SDS-PAGE and immunodetected with antibodies against TEF30 and proteins representative for the major thylakoid membrane complexes. (B) Silver staining of proteins immunoprecipitated from hexahistidine-tagged PSII particles with preimmune serum (Pre) and antibodies against TEF30. (C) Immunodetection of proteins immunopreciptated from PSII particles. Purified PSII particles equivalent to ~1 µg chlorophyll (Input) and 20% of the immunoprecipitates obtained with preimmune serum and antibodies against TEF30 (IP) were separated by SDS-PAGE and immunodecorated with the antisera indicated. Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. Figure 4. Co-migration of TEF30 with PSII core subunits as revealed by sucrose density gradient centrifugation and BN-PAGE. (A) Thylakoid membranes from strain cw15-325 (1 mg chlorophyll/mL) were solubilized with 1% β-DDM and separated on a linear 0.2 to 0.5 M sucrose gradient at 205,000 g for 16 h. Proteins were collected in 25 fractions, separated by SDS-PAGE and detected with colloidal blue staining (upper panel) or immunologically with the antisera indicated (lower panels). (B) Analysis of thylakoid membrane protein complexes by sucrose density gradient centrifugation as described in (A) was done on wild-type strain cc124, PSI mutant F23, and PSII mutant ΔpsbA/D. (C) Analysis of thylakoid membrane protein complexes by BN-PAGE. Thylakoids from wild-type strain cc124, PSI mutant F23, and PSII mutant nac2 were solubilized with 1% β-DDM and proteins corresponding to 5 µg chlorophyll separated on a 5-15% polyacrylamide BN gradient gel, followed by Coomassie staining (left panel) and immunodetection with antibodies against TEF30 (right panel). Protein bands were assigned to LHCII monomers (#1), LHCII trimers (#2), PSII monomers (#3), PSII II dimers (#4), PSI supercomplexes (#5), and PSII supercomplexes (#6) according to Jarvi et al. (2011) Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. Figure 5. PSII activity and levels of PSII core subunits are more sensitive to high light intensities or low temperatures in TEF30-amiRNA cells than in control cells. (A) cw15-325 control (Con) and TEF30-amiRNA strains were photoinhibited (PI) for 60 min at 1800 μmol photons m–2 s–1, and allowed to recover at 30 μmol photons m–2 s–1. PSII fluorescence was measured by pulse amplitude modulated fluorometry. Values represent the mean from three biological replicates, error bars indicate SD. (B) Control (Con) and TEF30-amiRNA #1.48 cells in the cw15-325 strain background were exposed to high PFDs (HL) of ~800 µmol photons m-2 s-1 for 6 h. Whole-cell proteins corresponding to 0.2, 1 or 2 µg chlorophyll (depending on the antiserum used) were separated on 10 or 14% SDS-polyacrylamide gels and analyzed by immunoblotting. (C) Control (Con) and TEF30-amiRNA #2.48 cells in the cw15-325 strain background were grown at a PFD of ~30 μmol photons m–2 s–1 and shifted from a growth temperature of 25°C to 10°C for 18 h. Samples were analyzed as in (B). Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. Figure 6. Comparison of LHCSR3 expression and oxygen evolution in control and TEF30-amiRNA cells. (A) Time course of LHCSR3 accumulation at 600 μmol photons m–2 s–1 (HL) in cw15-325 control and TEF30amiRNA cells grown in TAP-NH4, TAP-NO3, or TMP-NH4. Whole-cell proteins were separated by SDS-PAGE und immunodetected with antibodies against LHCSR3 and CF1β as loading control. (B) Oxygen evolution from thylakoid membranes isolated from cw15-325 control and TEF30-amiRNA cells grown in TAPNH4 medium at LL of ~30 µmol photons m-2 s-1 or HL of ~800 µmol photons m-2 s-1 for 4 h. Values represent the mean from experiments on two independent control and TEF30-amiRNA lines. Error bars indicate SD. (Note that the same thylakoid preparation was used for all PFDs applied, thus explaining the decline in O2 evolution at 1600 µmol photons m-2 s-1). Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. Figure 7. PSII de novo synthesis is not impaired in TEF30-amiRNA strains. (A) TEF30-amiRNA and control cells in the cw15-325 background were grown in TAP-NH4 at ~30 µmol photons m-2 s-1 to exponential phase. Cells were then harvested, washed and resuspended in TAP medium lacking sulfur (-S). After 48 h under –S conditions, cells were shifted back to regular TAP medium (+S) for another 24 h. Whole-cell proteins from samples taken during this time course were separated by SDS-PAGE and PSII polypeptide abundance measured via immunoblotting using the antibodies indicated and CF1β as loading control. (B) D1 signal intensities from (A) were quantified using the FUSIONCapt Advance program. Signals were normalized to the initial D1 levels in each strain. Error bars represent SD, n = 2. (C) S-depletion was done as in (A), but time on –S and +S reduced to 24 h and 6 h, respectively. Thylakoids were solubilized with 1% α-DDM and proteins corresponding to 5 µg chlorophyll separated on a 5-15% polyacrylamide BN gradient gel, followed by Coomassie staining (upper panel) and immunodetection with antibodies against D1 and CP43 (lower panels). Protein bands were assigned as in Figure 4C, the asterisk designates free CP43. (D) D1 signal intensities from protein bands #6 (PSII supercomplexes) and #4 (PSII dimers) from (C) were quantified using the FUSIONCapt Advance program. Signals were normalized to the initial D1 levels in each strain. Error bars represent SD, n = 4. Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. Figure 8. PSII stability is not impaired in TEF30amiRNA strains. (A) TEF30-amiRNA and control cells in the cw15-325 background were grown at ~30 µmol photons m-2 s-1 in TAP-medium in the presence or absence of 100 µg/mL CAP for 12 h. Proteins from whole cells harvested during the time course corresponding to 0.5 or 1 µg chlorophyll (depending on the antiserum used) were analyzed by immunoblotting. D1 levels were quantified with the FUSIONCapt Advance program. D1 levels were normalized to the values before the addition of CAP (at 0 h) and the ratio between D1 levels from TEF30-amiRNA strains and controls calculated. Values represent the mean from four independent replicates, errors bars indicate SD. (B) TEF30-amiRNA and control cells in the cw15-325 background were grown in TMP minimal medium at 200 μmol photons m–2 s–1 in the presence of 100 µg/mL CAP for 12 h. Analyses were done as in (A) with two biological replicates. (C) TEF30-amiRNA and control cells in the cw15-325 background were grown in TAP-medium in the dark in the presence of 100 µg/mL CAP for 72 h. Analyses were done as in (A) with two biological replicates. Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. Figure 9. Not PSII sensitivity to high light, but PSII repair is impaired in TEF30-amiRNA strains. (A) Cells of cw15-325 control (Con) and TEF30-amiRNA strains were photoinhibited (PI) for 60 min at 1800 μmol photons m–2 s–1 in the presence of chloroplast translation inhibitors chloramphenicol and lincomycin, and allowed to recover at 30 μmol photons m–2 s–1 (LL) without inhibitors. PSII fluorescence was measured by pulse amplitude modulated fluorometry. Values represent the mean from two biological replicates, error bars indicate SD. (B) Immunoblot analysis of whole-cell proteins collected from the experiments described in (A). Results from a typical experiment are shown in the upper panel, the mean from quantified D1 signals (normalized relative to the initial D1 signal) are shown in the lower panel. Values Downloaded fromreplicates, on June 15, 2017 by www.plantphysiol.org represent the mean fromCopyright five biological error- Published bars indicate SD. © 2015 American Society of Plant Biologists. All rights reserved. Figure 10. PSII supercomplex accumulation is impaired in TEF30-amiRNA strains exposed to high light. cw15-325 control (Con) and TEF30-amiRNA cells were grown at 30 µmol photons m-2 s-1 (LL) and exposed to 800 µmol photons m-2 s-1 for 4 h (HL). Thylakoid membranes were isolated, solubilized with 1% -DDM and protein complexes separated electrophoretically on a 7.5% native Deriphat-polyacrylamide gel. The gel was then photographed (left panels) and processed for immunoblotting using antibodies against TEF30 and the D1 protein (right panels). Bands were assigned to free pigments (#0), antenna monomers (#1), LHCII trimers (#2), PSI/II cores (#3), and PSI/II supercomplexes (#4) according to Formighieri et al. (2012). Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. Figure 11. Cells of TEF30-amiRNA strains contain fewer thylakoid membranes per stack (A) Electron microscopy images of cw15-325 cells characteristic for the categories normally stacked, reduced stacking, and swollen. Overview images are shown on the left and a zoom-in of the region demarcated by the black box is shown on the right. N, nucleus; P, pyrenoid; S, starch. Bars in overview images correspond to 1 µm and those in zoom-ins to 0.2 µm. (B) cw15-325 control (Con) and TEF30-amiRNA cells were grown in TAP-medium at ~30 µmol photons m-2 s-1 (LL) and exposed to ~800 µmol photons m-2 s-1 (HL) for 4 h. 50 electron micrographs each for two control strains and three Downloaded Junecondition 15, 2017 - Published by www.plantphysiol.org independent TEF30-amiRNA strains were taken from for on each and thylakoid phenotypes assessed according to Copyright © 2015 American Society of Plant Biologists. All rights reserved. the categories shown in (A). Values are given in percent and represent mean values from the two control and three TEF30-amiRNA strains. Error bars indicate SD. Significance was tested using a t-test (*P < 0.05 and **P < 0.01). Figure 12. A model for TEF30 function during PSII repair When PSII is damaged e.g. at high light intensities, the PSII complex enters the repair cycle. For this, PSII supercomplexes are disassembled and PSII monomers migrate from stacked into stroma-exposed membrane regions. After PSII monomers are partially disassembled to generate CP43-less forms, damaged D1 is degraded by FTSH and DEG proteases and a newly synthesized D1 copy is inserted into PSII monomers. At this step TEF30 may facilitate incorporation of the new D1 protein and/or reassembly of CP43, bind to repaired PSII monomers to protect them from undergoing repeated repair cycles, or facilitate the migration of repaired PSII monomers back to stacked regions for supercomplex reassembly. These functions are not mutually exclusive. Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. Parsed Citations Albiniak AM, Baglieri J, Robinson C (2012) Targeting of lumenal proteins across the thylakoid membrane. J Exp Bot 63: 1689-1698 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Allan RK, Ratajczak T (2011) Versatile TPR domains accommodate different modes of target protein recognition and function. Cell Stress Chaperones 16: 353-367 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Allmer J, Naumann B, Markert C, Zhang M, Hippler M (2006) Mass spectrometric genomic data mining: Novel insights into bioenergetic pathways in Chlamydomonas reinhardtii. Proteomics 6: 6207-6220 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Armbruster U, Labs M, Pribil M, Viola S, Xu W, Scharfenberg M, Hertle AP, Rojahn U, Jensen PE, Rappaport F, Joliot P, Dormann P, Wanner G, Leister D (2013) Arabidopsis CURVATURE THYLAKOID1 proteins modify thylakoid architecture by inducing membrane curvature. Plant Cell 25: 2661-2678 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Aro EM, Suorsa M, Rokka A, Allahverdiyeva Y, Paakkarinen V, Saleem A, Battchikova N, Rintamaki E (2005) Dynamics of photosystem II: a proteomic approach to thylakoid protein complexes. J Exp Bot 56: 347-356 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Aro EM, Virgin I, Andersson B (1993) Photoinhibition of Photosystem II. Inactivation, protein damage and turnover. Biochim Biophys Acta 1143: 113-134 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Bhuiyan NH, Friso G, Poliakov A, Ponnala L, van Wijk KJ (2015) MET1 is a thylakoid-associated TPR protein involved in photosystem II supercomplex formation and repair in Arabidopsis. Plant Cell 27: 262-285 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Bonente G, Ballottari M, Truong TB, Morosinotto T, Ahn TK, Fleming GR, Niyogi KK, Bassi R (2011) Analysis of LhcSR3, a protein essential for feedback de-excitation in the green alga Chlamydomonas reinhardtii. PLoS Biol 9: e1000577 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Caffarri S, Kouril R, Kereiche S, Boekema EJ, Croce R (2009) Functional architecture of higher plant photosystem II supercomplexes. EMBO J 28: 3052-3063 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Cai W, Ma J, Chi W, Zou M, Guo J, Lu C, Zhang L (2010) Cooperation of LPA3 and LPA2 is essential for photosystem II assembly in Arabidopsis. Plant Physiol 154: 109-120 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Cai W, Ma J, Guo J, Zhang L (2008) Function of ROC4 in the efficient repair of photodamaged photosystem II in Arabidopsis. Photochem Photobiol 84: 1343-1348 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Che YF, Fu AG, Hou X, McDonald K, Buchanan BB, Huang WD, Luan S (2013) C-terminal processing of reaction center protein D1 is essential for the function and assembly of photosystem II in Arabidopsis. Proc Natl Acad Sci U S A 110: 16247-16252 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Chua NH, Bennoun P (1975) Thylakoid membrane polypeptides of Chlamydomonas reinhardtii: wild-type and mutant strains deficient in photosystem II reaction center. Proc Natl Acad Sci U S A 72: 2175-2179 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. Cullen M, Ray N, Husain S, Nugent J, Nield J, Purton S (2007) A highly active histidine-tagged Chlamydomonas reinhardtii Photosystem II preparation for structural and biophysical analysis. Photochem Photobiol Sci 6: 1177-1183 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Danielsson R, Suorsa M, Paakkarinen V, Albertsson PA, Styring S, Aro EM, Mamedov F (2006) Dimeric and monomeric organization of photosystem II. Distribution of five distinct complexes in the different domains of the thylakoid membrane. J Biol Chem 281: 14241-14249 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title de Vitry C, Olive J, Drapier D, Recouvreur M, Wollman FA (1989) Posttranslational events leading to the assembly of photosystem II protein complex: a study using photosynthesis mutants from Chlamydomonas reinhardtii. J Cell Biol 109: 991-1006 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Drop B, Webber-Birungi M, Yadav SK, Filipowicz-Szymanska A, Fusetti F, Boekema EJ, Croce R (2014) Light-harvesting complex II (LHCII) and its supramolecular organization in Chlamydomonas reinhardtii. Biochim Biophys Acta 1837: 63-72 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Emanuelsson O, Nielsen H, Brunak S, von Heijne G (2000) Predicting subcellular localization of proteins based on their N-terminal amino acid sequence. J Mol Biol 300: 1005-1016 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Emanuelsson O, Nielsen H, von Heijne G (1999) ChloroP, a neural network-based method for predicting chloroplast transit peptides and their cleavage sites. Protein Sci 8: 978-984 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Erickson JM, Rahire M, Malnoe P, Girard-Bascou J, Pierre Y, Bennoun P, Rochaix JD (1986) Lack of the D2 protein in a Chlamydomonas reinhardtii psbD mutant affects photosystem II stability and D1 expression. EMBO J 5: 1745-1754 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Eriksson M, Gardestrom P, Samuelsson G (1995) Isolation, purification, and characterization of mitochondria from Chlamydomonas reinhardtii. Plant Physiol 107: 479-483 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Formighieri C, Ceol M, Bonente G, Rochaix JD, Bassi R (2012) Retrograde signaling and photoprotection in a gun4 mutant of Chlamydomonas reinhardtii. Mol Plant 5: 1242-1262 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Fristedt R, Willig A, Granath P, Crevecoeur M, Rochaix JD, Vener AV (2009) Phosphorylation of photosystem II controls functional macroscopic folding of photosynthetic membranes in Arabidopsis. Plant Cell 21: 3950-3964 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Girard J, Chua NH, Bennoun P, Schmidt G, Delosme M (1980) Studies on mutants deficient in the photosystem I reaction centers in Chlamydomonas reinhardtii. Curr Genet 2: 215-221 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Goral TK, Johnson MP, Brain AP, Kirchhoff H, Ruban AV, Mullineaux CW (2010) Visualizing the mobility and distribution of chlorophyll proteins in higher plant thylakoid membranes: effects of photoinhibition and protein phosphorylation. Plant J 62: 948959 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Greer DH, Laing WA, Kipnis T (1988) Photoinhibition of photosynthesis in intact kiwifruit (Actinidia deliciosa) leaves: Effect of temperature. Planta 174: 152-158 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. Guillot-Salomon T, Tuquet C, De Lubac M, Hallais MF, Signol M (1978) [Comparative analysis of ultrastructure and lipid composition of plastids from sun and shade plants (author's transl)]. Cytobiologie 17: 442-452 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Harris EH (2008) The Chlamydomonas Sourcebook: Introduction to Chlamydomonas and Its Laboratory Use. In HH Elizabeth, Ph.D, BS David, Ph.D, PD George B. Witman, eds, The Chlamydomonas Sourcebook (Second Edition). Elsevier/ Academic Press, San Diego, CA Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Haussuhl K, Andersson B, Adamska I (2001) A chloroplast DegP2 protease performs the primary cleavage of the photodamaged D1 protein in plant photosystem II. EMBO J 20: 713-722 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Heinnickel ML, Grossman AR (2013) The GreenCut: re-evaluation of physiological role of previously studied proteins and potential novel protein functions. Photosynth Res 116: 427-436 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Hemme D, Veyel D, Muhlhaus T, Sommer F, Juppner J, Unger AK, Sandmann M, Fehrle I, Schonfelder S, Steup M, Geimer S, Kopka J, Giavalisco P, Schroda M (2014) Systems-wide analysis of acclimation responses to long-term heat stress and recovery in the photosynthetic model organism Chlamydomonas reinhardtii. Plant Cell 26: 4270-4297 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Herbstova M, Tietz S, Kinzel C, Turkina MV, Kirchhoff H (2012) Architectural switch in plant photosynthetic membranes induced by light stress. Proc Natl Acad Sci U S A 109: 20130-20135 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Huang W, Chen Q, Zhu Y, Hu F, Zhang L, Ma Z, He Z, Huang J (2013) Arabidopsis thylakoid formation 1 is a critical regulator for dynamics of PSII-LHCII complexes in leaf senescence and excess light. Mol Plant 6: 1673-1691 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Ishikawa A, Tanaka H, Kato C, Iwasaki Y, Asahi T (2005) Molecular characterization of the ZKT gene encoding a protein with PDZ, K-Box, and TPR motifs in Arabidopsis. Biosci Biotechnol Biochem 69: 972-978 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Jarvi S, Suorsa M, Aro EM (2015) Photosystem II repair in plant chloroplasts - Regulation, assisting proteins and shared components with photosystem II biogenesis. Biochim Biophys Acta Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Jarvi S, Suorsa M, Paakkarinen V, Aro EM (2011) Optimized native gel systems for separation of thylakoid protein complexes: novel super- and mega-complexes. Biochem J 439: 207-214 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Johnson X, Vandystadt G, Bujaldon S, Wollman FA, Dubois R, Roussel P, Alric J, Beal D (2009) A new setup for in vivo fluorescence imaging of photosynthetic activity. Photosynth Res 102: 85-93 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Kapri-Pardes E, Naveh L, Adam Z (2007) The thylakoid lumen protease Deg1 is involved in the repair of photosystem II from photoinhibition in Arabidopsis. Plant Cell 19: 1039-1047 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Karpowicz SJ, Prochnik SE, Grossman AR, Merchant SS (2011) The GreenCut2 resource, a phylogenomically derived inventory of proteins specific to the plant lineage. J Biol Chem 286: 21427-21439 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. Kato Y, Sakamoto W (2014) Phosphorylation of photosystem II core proteins prevents undesirable cleavage of D1 and contributes to the fine-tuned repair of photosystem II. Plant J 79: 312-321 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Keren N, Ohkawa H, Welsh EA, Liberton M, Pakrasi HB (2005) Psb29, a conserved 22-kD protein, functions in the biogenesis of Photosystem II complexes in Synechocystis and Arabidopsis. Plant Cell 17: 2768-2781 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Kindle KL (1990) High-frequency nuclear transformation of Chlamydomonas reinhardtii. Proc Natl Acad Sci U S A 87: 1228-1232 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Kirchhoff H (2013) Diffusion of molecules and macromolecules in thylakoid membranes. Biochim Biophys Acta Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Kirchhoff H, Sharpe RM, Herbstova M, Yarbrough R, Edwards GE (2013) Differential mobility of pigment-protein complexes in granal and agranal thylakoid membranes of C(3) and C(4) plants. Plant Physiol 161: 497-507 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Koivuniemi A, Aro EM, Andersson B (1995) Degradation of the D1- and D2-proteins of photosystem II in higher plants is regulated by reversible phosphorylation. Biochemistry 34: 16022-16029 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Komenda J, Sobotka R, Nixon PJ (2012) Assembling and maintaining the Photosystem II complex in chloroplasts and cyanobacteria. Curr Opin Plant Biol 15: 245-251 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Kosourov S, Makarova V, Fedorov AS, Tsygankov A, Seibert M, Ghirardi ML (2005) The effect of sulfur re-addition on H(2) photoproduction by sulfur-deprived green algae. Photosynth Res 85: 295-305 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Kouril R, Oostergetel GT, Boekema EJ (2011) Fine structure of granal thylakoid membrane organization using cryo electron tomography. Biochim Biophys Acta 1807: 368-374 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Kouril R, Wientjes E, Bultema JB, Croce R, Boekema EJ (2013) High-light vs. low-light: effect of light acclimation on photosystem II composition and organization in Arabidopsis thaliana. Biochim Biophys Acta 1827: 411-419 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Kropat J, Hong-Hermesdorf A, Casero D, Ent P, Castruita M, Pellegrini M, Merchant SS, Malasarn D (2011) A revised mineral nutrient supplement increases biomass and growth rate in Chlamydomonas reinhardtii. Plant J 66: 770-780 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Kuchka MR, Goldschmidt-Clermont M, van Dillewijn J, Rochaix JD (1989) Mutation at the Chlamydomonas nuclear NAC2 locus specifically affects stability of the chloroplast psbD transcript encoding polypeptide D2 of PS II. Cell 58: 869-876 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227: 680-685 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23: 2947-2948 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. Lemaire C, Wollman FA (1989) The chloroplast ATP synthase in Chlamydomonas reinhardtii. I. Characterization of its nine constitutive subunits. J Biol Chem 264: 10228-10234 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Li XG, Duan W, Meng QW, Zou Q, Zhao SJ (2004) The function of chloroplastic NAD(P)H dehydrogenase in tobacco during chilling stress under low irradiance. Plant Cell Physiol 45: 103-108 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Lichtenthaler HK, Buschmann C, Doll M, Fietz HJ, Bach T, Kozel U, Meier D, Rahmsdorf U (1981) Photosynthetic activity, chloroplast ultrastructure, and leaf characteristics of high-light and low-light plants and of sun and shade leaves. Photosynth Res 2: 115-141 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Lindahl M, Spetea C, Hundal T, Oppenheim AB, Adam Z, Andersson B (2000) The thylakoid FtsH protease plays a role in the lightinduced turnover of the photosystem II D1 protein. Plant Cell 12: 419-431 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Lippuner V, Chou IT, Scott SV, Ettinger WF, Theg SM, Gasser CS (1994) Cloning and characterization of chloroplast and cytosolic forms of cyclophilin from Arabidopsis thaliana. J Biol Chem 269: 7863-7868 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Liu C, Willmund F, Whitelegge JP, Hawat S, Knapp B, Lodha M, Schroda M (2005) J-domain protein CDJ2 and HSP70B are a plastidic chaperone pair that interacts with vesicle-inducing protein in plastids 1. Mol Biol Cell 16: 1165-1177 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Lowry OH, Rosebrough NJ, Farr AL, Randall RJ (1951) Protein measurement with the Folin phenol reagent. J Biol Chem 193: 265275 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Malnoe A, Wang F, Girard-Bascou J, Wollman FA, de Vitry C (2014) Thylakoid FtsH protease contributes to photosystem II and cytochrome b6f remodeling in Chlamydomonas reinhardtii under stress conditions. Plant Cell 26: 373-390 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Maruyama S, Tokutsu R, Minagawa J (2014) Transcriptional regulation of the stress-responsive light harvesting complex genes in Chlamydomonas reinhardtii. Plant Cell Physiol 55: 1304-1310 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Mayfield SP, Bennoun P, Rochaix JD (1987) Expression of the nuclear encoded Oee1 protein is required for oxygen evolution and stability of Photosystem-II particles in Chlamydomonas reinhardtii. EMBO J 6: 313-318 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Merchant SS, Prochnik SE, Vallon O, Harris EH, Karpowicz SJ, Witman GB, Terry A, Salamov A, Fritz-Laylin LK, Marechal-Drouard L, Marshall WF, Qu LH, Nelson DR, Sanderfoot AA, Spalding MH, Kapitonov VV, Ren Q, Ferris P, Lindquist E, Shapiro H, Lucas SM, Grimwood J, Schmutz J, Cardol P, Cerutti H, Chanfreau G, Chen CL, Cognat V, Croft MT, Dent R, Dutcher S, Fernandez E, Fukuzawa H, Gonzalez-Ballester D, Gonzalez-Halphen D, Hallmann A, Hanikenne M, Hippler M, Inwood W, Jabbari K, Kalanon M, Kuras R, Lefebvre PA, Lemaire SD, Lobanov AV, Lohr M, Manuell A, Meier I, Mets L, Mittag M, Mittelmeier T, Moroney JV, Moseley J, Napoli C, Nedelcu AM, Niyogi K, Novoselov SV, Paulsen IT, Pazour G, Purton S, Ral JP, Riano-Pachon DM, Riekhof W, Rymarquis L, Schroda M, Stern D, Umen J, Willows R, Wilson N, Zimmer SL, Allmer J, Balk J, Bisova K, Chen CJ, Elias M, Gendler K, Hauser C, Lamb MR, Ledford H, Long JC, Minagawa J, Page MD, Pan J, Pootakham W, Roje S, Rose A, Stahlberg E, Terauchi AM, Yang P, Ball S, Bowler C, Dieckmann CL, Gladyshev VN, Green P, Jorgensen R, Mayfield S, Mueller-Roeber B, Rajamani S, Sayre RT, Brokstein P, Dubchak I, Goodstein D, Hornick L, Huang YW, Jhaveri J, Luo Y, Martinez D, Ngau WC, Otillar B, Poliakov A, Porter A, Szajkowski L, Werner G, Zhou K, Grigoriev IV, Rokhsar DS, Grossman AR (2007) The Chlamydomonas genome reveals the evolution of key animal and plant functions. Science 318: 245-250 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Mettler T, Muhlhaus T, Hemme D, Schottler MA, Rupprecht J, Idoine A, Veyel D, Pal SK, Yaneva-Roder L, Winck FV, Sommer F, Vosloh D, Seiwert B, Erban A, Burgos A, Arvidsson S, Schonfelder S, Arnold A, Gunther M, Krause U, Lohse M, Kopka J, Nikoloski Downloaded from M, on June 2017 - Systems Publishedanalysis by www.plantphysiol.org Z, Mueller-Roeber B, Willmitzer L, Bock R, Schroda Stitt 15, M (2014) of the response of photosynthesis, Copyright © 2015 American Society of Plant Biologists. All rights reserved. metabolism, and growth to an increase in irradiance in the photosynthetic model organism Chlamydomonas reinhardtii. Plant Cell 26: 2310-2350 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Minai L, Wostrikoff K, Wollman FA, Choquet Y (2006) Chloroplast biogenesis of photosystem II cores involves a series of assemblycontrolled steps that regulate translation. Plant Cell 18: 159-175 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Molnar A, Bassett A, Thuenemann E, Schwach F, Karkare S, Ossowski S, Weigel D, Baulcombe D (2009) Highly specific gene silencing by artificial microRNAs in the unicellular alga Chlamydomonas reinhardtii. Plant J 58: 165-174 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Mulo P, Sakurai I, Aro EM (2012) Strategies for psbA gene expression in cyanobacteria, green algae and higher plants: from transcription to PSII repair. Biochim Biophys Acta 1817: 247-257 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Nath K, Jajoo A, Poudyal RS, Timilsina R, Park YS, Aro EM, Nam HG, Lee CH (2013a) Towards a critical understanding of the photosystem II repair mechanism and its regulation during stress conditions. FEBS Lett 587: 3372-3381 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Nath K, Poudyal RS, Eom JS, Park YS, Zulfugarov IS, Mishra SR, Tovuu A, Ryoo N, Yoon HS, Nam HG, An G, Jeon JS, Lee CH (2013b) Loss-of-function of OsSTN8 suppresses the photosystem II core protein phosphorylation and interferes with the photosystem II repair mechanism in rice (Oryza sativa). Plant J 76: 675-686 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Naumann B, Busch A, Allmer J, Ostendorf E, Zeller M, Kirchhoff H, Hippler M (2007) Comparative quantitative proteomics to investigate the remodeling of bioenergetic pathways under iron deficiency in Chlamydomonas reinhardtii. Proteomics 7: 3964-3979 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Naumann B, Stauber EJ, Busch A, Sommer F, Hippler M (2005) N-terminal processing of Lhca3 Is a key step in remodeling of the photosystem I-light-harvesting complex under iron deficiency in Chlamydomonas reinhardtii. J Biol Chem 280: 20431-20441 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Nickelsen J, Rengstl B (2013) Photosystem II assembly: From cyanobacteria to plants. Annu Rev Plant Biol 64: 609-635 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Nixon PJ, Michoux F, Yu J, Boehm M, Komenda J (2010) Recent advances in understanding the assembly and repair of photosystem II. Ann Bot 106: 1-16 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Nordhues A, Schottler MA, Unger AK, Geimer S, Schonfelder S, Schmollinger S, Rutgers M, Finazzi G, Soppa B, Sommer F, Muhlhaus T, Roach T, Krieger-Liszkay A, Lokstein H, Crespo JL, Schroda M (2012) Evidence for a role of VIPP1 in the structural organization of the photosynthetic apparatus in Chlamydomonas. Plant Cell 24: 637-659 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Ossowski S, Schwab R, Weigel D (2008) Gene silencing in plants using artificial microRNAs and other small RNAs. Plant J 53: 674690 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Pagliano C, Saracco G, Barber J (2013) Structural, functional and auxiliary proteins of photosystem II. Photosynth Res 116: 167-188 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Peers G, Truong TB, Ostendorf E, Busch A, Elrad D, Grossman AR, Hippler M, Niyogi KK (2009) An ancient light-harvesting protein is critical for the regulation of algal photosynthesis. Nature 462: 518-521 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Peter GF, Thornber JP (1991) Biochemical composition and organization of higher-plant Photosystem-II light-harvesting pigmentproteins. J Biol Chem 266: 16745-16754 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Petroutsos D, Busch A, Janssen I, Trompelt K, Bergner SV, Weinl S, Holtkamp M, Karst U, Kudla J, Hippler M (2011) The chloroplast calcium sensor CAS is required for photoacclimation in Chlamydomonas reinhardtii. Plant Cell 23: 2950-2963 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Pierre Y, Popot JL (1993) Identification of two 4-kDa miniproteins in the cytochrome b6f complex from Chlamydomonas reinhardtii. C R Acad Sci III 316: 1404-1409 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Ramundo S, Casero D, Muhlhaus T, Hemme D, Sommer F, Crevecoeur M, Rahire M, Schroda M, Rusch J, Goodenough U, Pellegrini M, Perez-Perez ME, Crespo JL, Schaad O, Civic N, Rochaix JD (2014) Conditional depletion of the Chlamydomonas chloroplast ClpP protease activates nuclear genes involved in autophagy and plastid protein quality control. Plant Cell 26: 22012222 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Rimbault B, Esposito D, Drapier D, Choquet Y, Stern D, Wollman FA (2000) Identification of the initiation codon for the atpB gene in Chlamydomonas chloroplasts excludes translation of a precursor form of the beta subunit of the ATP synthase. Mol Gen Genet 264: 486-491 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Rintamaki E, Kettunen R, Aro EM (1996) Differential D1 dephosphorylation in functional and photodamaged photosystem II centers. Dephosphorylation is a prerequisite for degradation of damaged D1. J Biol Chem 271: 14870-14875 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Rochaix JD, Kuchka M, Mayfield S, Schirmerrahire M, Girardbascou J, Bennoun P (1989) Nuclear and chloroplast mutations affect the synthesis or stability of the chloroplast psbC gene-product in Chlamydomonas reinhardtii. EMBO J 8: 1013-1021 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Rutgers M, Schroda M (2013) A role of VIPP1 as a dynamic structure within thylakoid centers as sites of photosystem biogenesis? Plant Signal Behav 8: e27037 Samol I, Shapiguzov A, Ingelsson B, Fucile G, Crevecoeur M, Vener AV, Rochaix JD, Goldschmidt-Clermont M (2012) Identification of a photosystem II phosphatase involved in light acclimation in Arabidopsis. Plant Cell 24: 2596-2609 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Schagger H, Cramer WA, von Jagow G (1994) Analysis of molecular masses and oligomeric states of protein complexes by blue native electrophoresis and isolation of membrane protein complexes by two-dimensional native electrophoresis. Anal Biochem 217: 220-230 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Schagger H, von Jagow G (1991) Blue native electrophoresis for isolation of membrane protein complexes in enzymatically active form. Anal Biochem 199: 223-231 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Schmollinger S, Muhlhaus T, Boyle NR, Blaby IK, Casero D, Mettler T, Moseley JL, Kropat J, Sommer F, Strenkert D, Hemme D, Pellegrini M, Grossman AR, Stitt M, Schroda M, Merchant SS (2014) Nitrogen-sparing mechanisms in Chlamydomonas affect the transcriptome, the proteome, and photosynthetic metabolism. Plant Cell 26: 1410-1435 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Schroda M, Vallon O, Whitelegge JP, Beck CF, Wollman FA (2001) The chloroplastic GrpE homolog of Chlamydomonas: Two isoforms generated by differential splicing. Plant Cell 13: 2823-2839 Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Schroda M, Vallon O, Wollman FA, Beck CF (1999) A chloroplast-targeted heat shock protein 70 (HSP70) contributes to the photoprotection and repair of photosystem II during and after photoinhibition. Plant Cell 11: 1165-1178 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Schulz-Raffelt M, Lodha M, Schroda M (2007) Heat shock factor 1 is a key regulator of the stress response in Chlamydomonas. Plant J 52: 286-295 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Silva P, Thompson E, Bailey S, Kruse O, Mullineaux CW, Robinson C, Mann NH, Nixon PJ (2003) FtsH is involved in the early stages of repair of photosystem II in Synechocystis sp PCC 6803. Plant Cell 15: 2152-2164 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Sirpio S, Allahverdiyeva Y, Suorsa M, Paakkarinen V, Vainonen J, Battchikova N, Aro EM (2007) TLP18.3, a novel thylakoid lumen protein regulating photosystem II repair cycle. Biochem J 406: 415-425 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Sonnhammer EL, von Heijne G, Krogh A (1998) A hidden Markov model for predicting transmembrane helices in protein sequences. Proc Int Conf Intell Syst Mol Biol 6: 175-182 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Strenkert D, Schmollinger S, Sommer F, Schulz-Raffelt M, Schroda M (2011) Transcription factor dependent chromatin remodeling at heat shock and copper responsive promoters in Chlamydomonas reinhardtii. Plant Cell 23: 2285-2301 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Sugiura M, Inoue Y, Minagawa J (1998) Rapid and discrete isolation of oxygen-evolving His-tagged photosystem II core complex from Chlamydomonas reinhardtii by Ni2+ affinity column chromatography. FEBS Lett 426: 140-144 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Sun X, Fu T, Chen N, Guo J, Ma J, Zou M, Lu C, Zhang L (2010) The stromal chloroplast Deg7 protease participates in the repair of photosystem II after photoinhibition in Arabidopsis. Plant Physiol 152: 1263-1273 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Takahashi S, Murata N (2005) Interruption of the Calvin cycle inhibits the repair of Photosystem II from photodamage. Biochim Biophys Acta 1708: 352-361 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28: 2731-2739 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Tietz S, Puthiyaveetil S, Enlow HM, Yarbrough R, Wood M, Semchonok DA, Lowry T, Li Z, Jahns P, Boekema EJ, Lenhert S, Niyogi KK, Kirchhoff H (2015) Functional implications of Photosystem II crystal formation in photosynthetic membranes. J Biol Chem 290: 14091-14106 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Tikkanen M, Nurmi M, Kangasjarvi S, Aro EM (2008) Core protein phosphorylation facilitates the repair of photodamaged photosystem II at high light. Biochim Biophys Acta 1777: 1432-1437 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Tokutsu R, Kato N, Bui KH, Ishikawa T, Minagawa J (2012) Revisiting the supramolecular organization of photosystem II in Chlamydomonas reinhardtii. J Biol Chem 287: 31574-31581 Pubmed: Author and Title CrossRef: Author and Title Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. Google Scholar: Author Only Title Only Author and Title Tokutsu R, Minagawa J (2013) Energy-dissipative supercomplex of photosystem II associated with LHCSR3 in Chlamydomonas reinhardtii. Proc Natl Acad Sci U S A 110: 10016-10021 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Tomizioli M, Lazar C, Brugiere S, Burger T, Salvi D, Gatto L, Moyet L, Breckels LM, Hesse AM, Lilley KS, Seigneurin-Berny D, Finazzi G, Rolland N, Ferro M (2014) Deciphering thylakoid sub-compartments using a mass spectrometry-based approach. Mol Cell Proteomics 13: 2147-2167 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Tyystjarvi E (2013) Photoinhibition of Photosystem II. Int Rev Cell Mol Biol 300: 243-303 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Tyystjarvi E, Aro EM (1996) The rate constant of photoinhibition, measured in lincomycin-treated leaves, is directly proportional to light intensity. Proc Natl Acad Sci U S A 93: 2213-2218 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Umena Y, Kawakami K, Shen JR, Kamiya N (2011) Crystal structure of oxygen-evolving photosystem II at a resolution of 1.9 A. Nature 473: 55-60 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Uniacke J, Colon-Ramos D, Zerges W (2011) FISH and immunofluorescence staining in Chlamydomonas. Methods Mol Biol 714: 1529 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Uniacke J, Zerges W (2007) Photosystem II assembly and repair are differentially localized in Chlamydomonas. Plant Cell 19: 36403654 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Vallon O, Wollman FA, Olive J (1985) Distribution of intrinsic and extrinsic subunits of the PSII protein complex between appressed and non-appressed regions of the thylakoid membrane: an immunocytochemical study. FEBS Lett 183: 245-250 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Vallon O, Wollman FA, Olive J (1986) Lateral distribution of the main protein complexes of the photosynthetic appartus in Chlamydomonas reinhardttii and in spinach: an immunocytochemical study using inteact thylakoid membranes and PS II enriched membrane preparation. Photobiochem Photobiophys 12: 203-220 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Wang Q, Sullivan RW, Kight A, Henry RL, Huang J, Jones AM, Korth KL (2004) Deletion of the chloroplast-localized Thylakoid formation1 gene product in Arabidopsis leads to deficient thylakoid formation and variegated leaves. Plant Physiol 136: 3594-3604 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Wittenberg G, Levitan A, Klein T, Dangoor I, Keren N, Danon A (2014) Knockdown of the Arabidopsis thaliana chloroplast protein disulfide isomerase 6 results in reduced levels of photoinhibition and increased D1 synthesis in high light. Plant J 78: 1003-1013 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Woessner JP, Gillham NW, Boynton JE (1986) The sequence of the chloroplast atpB gene and its flanking regions in Chlamydomonas reinhardtii. Gene 44: 17-28 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Wu HY, Liu MS, Lin TP, Cheng YS (2011) Structural and functional assays of AtTLP18.3 identify its novel acid phosphatase activity in thylakoid lumen. Plant Physiol 157: 1015-1025 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved. Wunder T, Xu W, Liu Q, Wanner G, Leister D, Pribil M (2013) The major thylakoid protein kinases STN7 and STN8 revisited: effects of altered STN8 levels and regulatory specificities of the STN kinases. Front Plant Sci 4: 417 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Xue H, Tokutsu R, Bergner SV, Scholz M, Minagawa J, Hippler M (2015) Photosystem II subunit R is required for efficient binding of light-harvesting complex stress-related protein 3 to Photosystem II-light-harvesting supercomplexes in Chlamydomonas reinhardtii. Plant Physiol 167: 1566-1578 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Yamatani H, Sato Y, Masuda Y, Kato Y, Morita R, Fukunaga K, Nagamura Y, Nishimura M, Sakamoto W, Tanaka A, Kusaba M (2013) NYC4, the rice ortholog of Arabidopsis THF1, is involved in the degradation of chlorophyll - protein complexes during leaf senescence. Plant J 74: 652-662 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Ye F, Zhang M (2013) Structures and target recognition modes of PDZ domains: recurring themes and emerging pictures. Biochem J 455: 1-14 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Yokthongwattana K, Chrost B, Behrman S, Casper-Lindley C, Melis A (2001) Photosystem II damage and repair cycle in the green alga Dunaliella salina: involvement of a chloroplast-localized HSP70. Plant Cell Physiol 42: 1389-1397 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Zerges W, Rochaix JD (1998) Low density membranes are associated with RNA-binding proteins and thylakoids in the chloroplast of Chlamydomonas reinhardtii. J Cell Biol 140: 101-110 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Zhang LX, Paakkarinen V, van Wijk KJ, Aro EM (1999) Co-translational assembly of the D1 protein into photosystem II. J Biol Chem 274: 16062-16067 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Zhang LX, Paakkarinen V, van Wijk KJ, Aro EM (2000) Biogenesis of the chloroplast-encoded D1 protein: Regulation of translation elongation, insertion, and assembly into photosystem II. Plant Cell 12: 1769-1781 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Zienkiewicz M, Ferenc A, Wasilewska W, Romanowska E (2012) High light stimulates Deg1-dependent cleavage of the minor LHCII antenna proteins CP26 and CP29 and the PsbS protein in Arabidopsis thaliana. Planta 235: 279-288 Pubmed: Author and Title CrossRef: Author and Title Google Scholar: Author Only Title Only Author and Title Downloaded from on June 15, 2017 - Published by www.plantphysiol.org Copyright © 2015 American Society of Plant Biologists. All rights reserved.