* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download The genetic code and tRNA Biochemistry 302 February 15, 2006
Citric acid cycle wikipedia , lookup
Ribosomally synthesized and post-translationally modified peptides wikipedia , lookup
Western blot wikipedia , lookup
Interactome wikipedia , lookup
Gene expression wikipedia , lookup
Messenger RNA wikipedia , lookup
Peptide synthesis wikipedia , lookup
Metalloprotein wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Point mutation wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Proteolysis wikipedia , lookup
Protein structure prediction wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Biochemistry wikipedia , lookup
Epitranscriptome wikipedia , lookup
Biosynthesis wikipedia , lookup
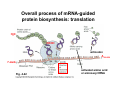
The genetic code and tRNA Biochemistry 302 February 15, 2006 Major advances in defining mechanism of protein biosynthesis • Identification of ribosomes (Paul Zamecnik, 1950s) – Injected radioactive amino acids into rats, fractionated liver homogenates at various time points, then analyzed subcellular distribution of labeled proteins. – At early time points, “hot” proteins only in “small” RNP particles. • Ribosomes attached to outer face of ER Identification of tRNAs – M. Hoagland & Zamecnik – Amino acids become “activated” i.e. attached to heat-stable soluble RNA species if incubated with cytosolic extract. – R. Holley → tRNA • mRNAguided protein synthesis Crick’s adaptor hypothesis Predicted triplet code 43 = 64 possible combinations Overall process of mRNA-guided protein biosynthesis: translation H2N 3′ 5′ COOH 5′ anticodon A200-250 7-meG Fig. 4-22 activated amino acid or aminoacyl tRNA Deciphering the triplet genetic code…. Francis Crick and Sidney Brenner, 1961 • Used growth conditions which generated mutations in T4 bacteriophage DNA. • Deletion or insertion of 1 or 2 nt led to nonsense proteins. • Deletion or insertion of 3 nt led to insertion or deletion of one amino acid residue. • Code must be in triplets and read in units of three. • Eliminated overlapping and punctuated code hypotheses. • First codon read establishes the reading frame. Fig. 27-1 H.G. Khorana experiments confirm the code is triplet and unpunctuated Fig. 27-2 • Developed methods for 1) synthesizing polyribonucleotides and 2) carrying out cell-free ribosome-mediated translation. • (AAG)n template gave rise to three different homopolymers. • Confirmed the importance of the reading frame and revealed that AAG, AGA, and GAA must be codons for K, R, and E. ~1961-1964 Genetic code is broken • Heinrich Matthaei, Philip Leder, and Marshall Nirenberg •1: Used synthetic RNA triplets that would bind ribosomes and specify the binding of only certain aminoacylated-tRNAs • 2: Developed cell-free translation system using random RNA polymers prepared with polynucleotide phosphorylase (NMP)n + NDP ⇌ (NMP)n+1 + Pi • 1+2: Codon assignment 22nd pyrrolysine 21st selenocysteine Fig. 27-3 Genetic code is not quite universal (alternate use generally confined to mitochondrial transcripts in certain organisms) * Table 27-1 Important features of the genetic code • Triplet and non-overlapping • Redundancy of the code • Most amino acids are encoded by several codons, others by only one (e.g. Met, Trp). • Generally, degeneracy is found at the third position. • Special codons for starting and stopping protein synthesis • Start: AUGMet (uncommon: UUGLeu, AUUIle, GUGVal) • Stop: UAA, UGA, UAG (rare alternate use: UGASec, UGATrp, UAGPyl) • Open reading frame • Biochemical basis for complexity of the code • Base-pairing between tRNA anticodon and mRNA codon • Optimize speed and accuracy of protein synthesis Note antiparallel alignment of tRNA and mRNA Wobble hypothesis: attempt to explain redundancy of the genetic code • Rationale for why single a tRNA can recognize several different codons. • Multiple recognition involves the 3′ residue of the codon and the 5′ residue of the anticodon. • In 1966, Crick proposed that the 5′ base of the anticodon could wobble during translation. − Movement permits nonWatson-Crick H-bonding. − Model explains frequent degeneracy observed in the 3′ site of some codons. e.g. AlatRNAAla can base pair with codons GCC and GCU. Crick’s wobble rules (base-pairing possibilities in wobble pairs) • First 2 base pairs in codon-anticodon interaction form strong WC H-bonds and, usually specify the amino acid. • First base (5′) of anticodon determines # of codons read: • C, A → only one codon read by tRNA, binding is specific. • U, G → two codons read by tRNA, binding a little less specific. • I → three codons read by tRNA (tRNAArg), “weak” base-pairing • Codons that specify the same amino acid but differ in either of the first two bases require different tRNAs (e.g. Arg, Ser, Leu) • Rules require a minimum of 32 tRNAs to translate all 61 amino acid codons (∼70 in E. coli). Table 27-2 Wobble bps Wobble hypothesis does not explain all complexities inherent in decoding • • • tRNAs whose anticodons (e.g. CUA, UCA) decode stop codons: UAG (pyrrolysine), UGA (selenocysteine) Selenocysteine found in formate dehydrogenase (bacteria), glutathione peroxidase & thioredoxin reductase (mammals). Alternate use of stop codon dictated by… – Specific 3′ mRNA hairpin element (SECIS) and binding protein (SBP2) – Specialized elongation factor known as (eEFSec or SelB) – Sec-tRNASec (actually special type of serine tRNA that gets converted to Sec) Pyrrolysine L-lysine in amide linkage (4R,5R)-4-substituted pyrroline-5-carboxylate Overview of tRNA structural elements and interaction partners • • 73 to 93 nucleotides long (eukaryotes) and arranged in a canonical 2-D cloverleaf (tRNAAla R. Holley, 1965) Intrastrand H-bonding specifies structure – Four “arms” and a “stub” – pG on 5′ end, CCA acceptor stem on 3′ end (all tRNAs) • Interaction partners – – – – • Processing enzymes Aminoacyl-tRNA synthetases EF-Tu (elongation factor) Ribosome Where does specificity come from in 73 to 93 nucleotides? 76 residue yeast tRNAAla 1st nucleic acid to be completely sequenced. Note the number of modified bases. General secondary and tertiary (twisted L) structure of tRNAs 3-D Structure of yeast tRNAPhe deduced from xray diffraction Invariant residues are boxed and shaded in pink. The D- and TψC-arms are important for tertiary folding & geometric separation of acceptor stem & anticodon loop. The TψC arm interacts with the large ribosomal subunit. Many modified bases are commonly found in tRNA no double bond N and C in altered positions ψ Fig. 27-7 • • • Provide increased stability by facilitating proper 3-D folding. Modify the specificity of anticodon-codon interactions (inosine). Serve as recognition elements for amino acyl-tRNA synthetases. Interactions mediating conserved 3-D tRNA folding (hand drill or twisted L) • • The D loop and TψC loops fold inward to provide maximum Hbonding and base stacking. Fig. 4-20 Yeast tRNAPhe 76 bases • Single-stranded but extensive base pairing (self-complementary) and base stacking Two A-form helices at right angles to each other Non Watson:Crick interactions: – Unusual base pairs – Base triples – Base backbone contacts Conservation of 3-D structure is to ensure correct fit in the ribosome. Aminoacylation of tRNA adaptor, the first step in protein synthesis aaRS Mg++ Amino acid + ATP Amino Acid~AMP + PPi aaRS +tRNA Amino Acid~AMP aaRS: aminoacyl-tRNA synthetase (21 synthetases in E. coli each recognizing one AA and one or more tRNAs. Lys has two aaRSs. aaRSs also have hydrolytic “proofreading” ability.) AA-tRNA + AMP Chemistry of aminoacylation…. Enzyme bound Class II aaRS: AA is transferred to the 3′ OH of invariant adenosine residue on acceptor arm. Class II aaRS Class I aaRS: AA is transferred to the 2′ OH of invariant adenosine then to the 3′ OH via transesterification. Class I aaRS Fig. 27-10 How an aminoacyl-tRNA synthetase identifies the right tRNA….no simple rule, depends on each enzyme, “second genetic code” Recognition elements exist at many different locations but tend to cluster in the anticodon loop and acceptor stem. Fig. 27-11 Structural features of aa-tRNA synthetases Acceptor stem, ATP, & AA in close proximity • • • • Class I: glutaminyl-tRNA synthetase (monomers) Class II: aspartyl-tRNA synthetase (dimers/tetramers) (Met, Ile, Val, Leu, Tyr, Trp, Glu, Arg, Gln, Cys, Lys) (Ser, Pro, His, Thr, Gly, Asp, Asn, Lys, Ala, Phe) Different protein folds • Different consensus sequences for • ATP binding Different mode of tRNA recognition • Different stereochemistry • Subgroups: tRNA binding domains Examples of class switching (e.g. TyrRS binds tRNA in class II mode) Induced fit facilitates catalysis Hydrolytic proofreading