* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Chem*3560 Lecture 27: Membrane transport
Adenosine triphosphate wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
NADH:ubiquinone oxidoreductase (H+-translocating) wikipedia , lookup
Magnesium in biology wikipedia , lookup
Vesicular monoamine transporter wikipedia , lookup
Photosynthetic reaction centre wikipedia , lookup
Electron transport chain wikipedia , lookup
Light-dependent reactions wikipedia , lookup
Fatty acid metabolism wikipedia , lookup
Metalloprotein wikipedia , lookup
Catalytic triad wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Ultrasensitivity wikipedia , lookup
Genetic code wikipedia , lookup
Signal transduction wikipedia , lookup
Proteolysis wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Magnesium transporter wikipedia , lookup
SNARE (protein) wikipedia , lookup
Protein structure prediction wikipedia , lookup
Biosynthesis wikipedia , lookup
Oxidative phosphorylation wikipedia , lookup
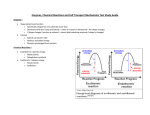
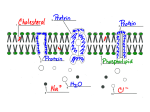
Chem*3560 Lecture 27: Membrane transport Diffusion across permeable membranes If solutions are separated by a permeable membrane, solutes will cross the membrane by random diffusion. Molecules leave the side with higher concentration at a faster rate, causing the concentration to decrease on the source side and increase on the destination side, until equilibrium is reached when both sides are at equal concentration. If the solute is charged, a concentration difference across the membrane may induce a voltage difference or membrane potential Vm (same as ∆ψ used in the context of oxidative phosphorylation). If negative ions are in excess on the left, the membrane potential will be negative on the left. The membrane potential also provides a driving force, because negative ions will tend to move towards the positive side (Lehninger p.408). Ionic substances move across a permeable membrane in response to combination of the membrane potential and the concentration difference, which is referred to as the electrochemical gradient. The net direction of ion movement will decrease both concentration difference and membrane potential, until both are zero and net flux ceases. Lipid membranes are impermeable to most polar molecules Membranes surround cells to enclose their contents - leaving open the question of how necessary nutrients enter cells or pass through internal membranes. The bilayer portion of the membrane is impermeable to most polar molecules, which represent most of the intermediates of metabolism. Membrane proteins play a direct role in allowing transport of substances across membranes. In addition to simple transport, which is governed by the relative concentrations on each side of the membrane, some membrane proteins are also involved in active transport, in which a particular solute can be driven across a membrane from a region of low concentration to a region of higher concentration. This requires an external energy source such as ATP hydrolysis. Facilitated diffusion Many transport proteins simply provide a passage through the membrane for specific solutes, governed solely by the electrochemical gradient, without any external energy being applied. This is facilitated diffusion. The transport protein permits passage of solute, but does not drive solute against an electrochemical gradient. The problem for polar solutes and lipid bilayers is that solutes exist in aqueous solution in a hydrated state. In order to enter the hydrocarbon core of the bilayer, they must lose their hydration layer, and the energy required to dehydrate a solute may be hundreds of kJ/mol. Desolvation contributes to the activation free energy ∆GI for the transport process, hence transport of polar solutes across a lipid bilayer is imperceptibly slow (Lehninger p.409). A transport protein provides a channel through the membrane , and provides binding sites for selected solutes such that the binding energy of solute in the channel is roughly equal to the solvation energy in the surroundings. The result is that solute can leave the aqueous medium, and become dehydrated, but immediately passes to a binding site of equivalent energy. The energy change is thus minimal so that activation energy is low. When the solute reaches the other side, it must be released from the binding site, but immediately becomes hydrated again. If the channel binds a solute too tightly, it will enter the channel easily, but now there's an activation energy barrier for the solute to reenter the aqueous solution on the other side. Many passive substrate transporters share a 12-helix structure These proteins belong to the Major Facilitator Superfamily. A superfamily is a grouping of proteins from many different organisms that share a common structural arrangement. The Facilitator superfamily perform facilitated diffusion of substrates across membranes. There were several structures published during 2002-2003 illustrating how these proteins may function. (The current edition of Lehninger was written 1998-1999 so the description on pp. 411-413 is speculative but not far off the mark). These proteins are arranged in two subgroups of six helices each that surround the central substrate binding site like a pair of cupped hands. The outward faces of the helices are lined with nonpolar amino acids where they contact the hydrocarbon core of the bilayer, but have inward facing polar amino acids facing the substrate binding site. The schematic layout of the twelve helices I-XII is shown above viewed looking down with the bilayer in the plane of the paper. This produces a favourable binding site for a polar substrate in the middle of the protein (S in the schematic). On the right above is a cross section through the actual protein structure of LacY, the lactose sugar transporter in the E. coli cell membrane. Yellow amino acids are nonpolar, and face the hydrocarbon core of the bilayer. Green amino acids are polar, and face the substrate (grey C and red O atoms). The section has been cut in a plane at mid bilayer level. Atoms are shown in their true volume, so there are few gaps. The helical wheel The asymmetric spatial arrangement of polar and non-polar amino acids in the α-helices that make up this structure can be detected in the amino acid sequence by a technique called helical wheel analysis. The method is based on the geometry of the α-helix, which has 3.6 amino acids per turn of helix, or in other words the orientation of each amino acid is 100o away from the previous. Amino acids in a sequence are written out around the perimeter of a circle, one amino acid every 100o . If the amino acids have the correct distribution in the sequence, one side of the helix will be predominantly polar and the side that contacts bilayer will be non polar. The two halves of LacY alternate between two conformations IColour coding in the figure runs from blue at N-terminus to red at the C-terminus. If colours are traced it can be seen that the six helices making the N-terminal half are all on the left of the substrate, and the six making the C-terminal half are all on the right. In the left hand structure, the substrate side is exposed to the exterior ///s \\\ and can bind substrate. The two halves rock about a pivot point roughly at the midline, so that the channel closes on the exterior half and opens on the cytoplasmic side, \\\s ///, and this allows the substrate to be released into the cell. The channel is therefore always closed on one side or the other, so non-substrates can't leak through (see Lehninger p. 412, Fig. 12-26 for a good guess at how the erythrocyte glucose transporter might work). Substrate transport follows Michaelis-Menten kinetics Transport depends on 1) a substrate occupying a binding site, and 2) the conformational changes which accesses the opposite side. This pattern is little different from the standard Michaelis-Menten behaviour of an enzyme. The transport of a single substrate is called uniport Uniport: a single substrate is transported in the direction governed by the electrochemical gradient. After passage of substrate, the transporter randomly flips between conformations. If there is more substrate on the left, the site is more likely to become occupied when open to the left, so net transport is towards the right where the concentration is lower. Many transporters operate by a cotransport mechanism Symport: two substrates are co-transported in the same direction. This could be due to the requirements of the substrate site, so the conformational change only occurs when both components are present. Alternatively, there could be two channels and passage of the second substrate is necessary for the transporter to return to its initial state. In many cases, the second substrate is H+, and the transporter takes advantage of the H+ gradient of oxidative phosphorylation. Net direction of transport is governed by whichever of the two substrates has the steeper electrochemical gradient. Antiport: substrates are exchanged; for each molecule A that enters, a molecule of B must leave. Substrate A flips the conformation in one direction, substrate B brings it back to the original state. Net direction of transport is governed by whichever of the two substrates has the steeper gradient. An example of antiport is the ATP:ADP exchange translocator of the mitochondrial membrane, which exports ATP while bringing in ADP: ATPin + ADPout ‡ ATPout + ADPin The advantage of antiport is that it maintains balance of substrates between compartments. The mitochondrion should never accumulate too much or run out of adenosine compounds.