* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Supplementary Information (doc 290K)
History of genetic engineering wikipedia , lookup
Gene desert wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Metabolic network modelling wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
Minimal genome wikipedia , lookup
Genomic imprinting wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Ridge (biology) wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Genome evolution wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Genome (book) wikipedia , lookup
Mir-92 microRNA precursor family wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Microevolution wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Designer baby wikipedia , lookup
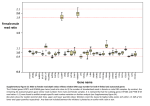
Supporting Information For Data mining reveals a network of early-response genes as a consensus signature of drug-induced in vitro and in vivo toxicity Jitao David Zhanga,1, Nikolaos Berntenisa, Adrian Rotha and Martin Ebelinga,1 a Non-Clinical Safety, F. Hoffmann-La Roche AG, Grenzacherstrasse 124, 4070 Basel, Switzerland 1 Corresponding author: Jitao David Zhang ([email protected]) or Martin Ebeling ([email protected]), F. Hoffmann-La Roche AG, Grenzacherstrasse 124, 4070 Basel, Switzerland. Contents Supplementary Methods ........................................................................................................................ 1 Identify early gene signatures with hierarchical linear models .......................................................... 1 SVM performance measures .............................................................................................................. 2 Temporal expected node occupancy fractions from Boolean network ensembles ........................... 2 Supplementary Tables ............................................................................................................................ 2 Supplementary Figure Legends............................................................................................................... 3 Supplementary Methods Identify early gene signatures with hierarchical linear models As a starting point we built a simple linear model for regression analysis for differential gene expression only with regard to time, written as where denotes differential gene expression, denotes the intercept, and denotes the coefficient of time points t (2, 8, or 24 hours). This model, however, does not include the information of treatments that induced progressive profiles. Therefore we next built a two-level model with varying intercepts and slopes to capture commonly regulated genes in progressive profiles. The model can be represented as implying that intercept and slope of the linear model can vary by treatments. Early gene signatures were defined by the genes of with logFC (log2 fold change) at 2h equal to or larger than 0.5 (or equal to or smaller than -0.5), and with an associated p value (derived from the t-distribution followed by , , and the number of conditions C) smaller than 0.001. SVM performance measures With TP, FP, FN denoting counts of true positive, false positive, and false negative cases, the measures to evaluate performance are defined as: Temporal expected node occupancy fractions from Boolean network ensembles In Boolean networks, activators and inhibitors in a biological system are represented as activating and inhibiting nodes in a network. All nodes can be in only one of two states, ON or OFF, simplifying the typically sigmoidal stimulus-response relationship to a step function. In any specific network state, the values of some nodes may be subject to change, according to well-specified rules capturing the biology of the system, giving rise to a successor state. If it so happens that no node is subject to change, the system is in a fixed-state, i.e., it has reached an equilibrium point. Most studies focus on a single network, looking for possible fixed network states (i.e., possible phenotypes) and then probing fixed state stability under various perturbations. However, Boolean network ensembles can successfully generate semi-quantitative time-response information. We developed an algorithm that generates time-response profiles of expected node occupancy fractions from such an ensemble. In this approach, each ensemble member follows one of the possible time evolution scenarios. Starting from an initial state the ensemble goes through successive iteration steps and at each step all possible successor states of all ensemble members are assigned a weight that is a measure of their likelihood to be occupied. A state-network is thus generated that encompasses the discrete-time evolution of the ensemble. Each path within it corresponds to an ensemble member. The expected occupancy fraction of a node at a given time step is the sum of the weights (at that step) of the ensemble members for which the node is ON. The manuscript describing the algorithm is under review. Supplementary Tables Supplementary Table 1 (Attached XLS file): Compounds and natural molecules that were tested in TG-GATEs: names, ATC codes, and TG-GATEs download links organized by species, experimental systems, and dosing paradigms. Supplementary Table 2: Pathological terms used as prediction targets of Support Vector Machines. TG-GATEs provides pathological records of in vivo experiments, which we used to test the predictive power of the early-response network. If any of the pathological changes listed in the following table was observed in a sample, its pathological state was set as TRUE. Otherwise the state was set as FALSE. Liver Kidney Anisonucleosis Anisonucleosis Cellular infiltration Cellular infiltration Change, basophilic Change, basophilic Degeneration Degeneration Fibrosis Dilatation Increased mitosis Eosinophilic body Proliferation, bile duct Fibrosis Single cell necrosis Hyperplasia Hypertrophy Necrosis Proliferation Supplementary Figure Legends Figure 1: There are fewer early-response genes but they are more generic than late-response genes in rat hepatocytes. The legends follow the definition in the caption of Figure 2 of the manuscript. (A) Number of differentially expressed genes (DEGs) induced by compounds in rat primary hepatocytes, stratified by dose and time. (B) The normalized generality score of DEGs, stratified by dose and time. We observed that the normalized generality scores of 2h DEGs in rat are generally lower than the scores of their counterparts in human. On average, however, they still tend to be higher than for the 24h DEGs. Figure 2: Cytotoxicity matrices of TG-GATEs compounds that were tested in human primary hepatocytes. Figure 3: Expression patterns of EGR1, ATF3, GDF15, and FGF21 in (A) human and (B) rat primary hepatocytes where treatments did not cause any moderate or even strong cytotoxicity. Bold lines indicate average changes of gene expression, and error bars indicate standard errors of the mean. The line representing GDF15 is overwritten by the line representing FGF21 in both panels. Figure 4: Unsupervised clustering of differential expression profiles in rat primary hepatocytes revealed four groups (separated by black lines). Note that the NGTX group (indicated by the black arrow) contrasts strongly with other profiles. Columns of the matrix are frequently induced genes, which were induced in more than five percent of all samples. Figure 5: Cytotoxicity matrices of TG-GATEs compounds that were tested in rat primary hepatocytes. Figure 6: Identifying early-response signatures in rat primary hepatocytes. (A) Amino-acid (AA) sequence identity and similarity between orthologous proteins in human and rat. (B) Cxcl2, the rat ortholog of human CXCL3, which shares 47.8% protein sequence similarity with human IL8, is an early-response gene in rat. (D) Temporal profiles of Tob2, the rat ortholog of human TOB2. Its dynamics are highly conserved in both species; however its induction at 2 hours did not meet the predefined significance level (see supplementary methods). Figure 7: Robustness of the early-response network assessed by edge permutations. In each permutation step, one edge of the network is deleted, and dynamics of the permutated network is simulated with the two initial states defined in the main text. Each panel in the figure represents one permutation with one initial state. For visualization purposes random noise was added to the simulation results.