* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Poster
Protein moonlighting wikipedia , lookup
Signal transduction wikipedia , lookup
Magnesium transporter wikipedia , lookup
Endomembrane system wikipedia , lookup
Cytokinesis wikipedia , lookup
Homology modeling wikipedia , lookup
Intrinsically disordered proteins wikipedia , lookup
Protein structure prediction wikipedia , lookup
List of types of proteins wikipedia , lookup
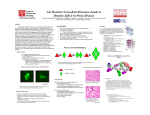
Who’s Afraid of the Big, Bad, Misfolded Protein? Meghan Murphy1, Moses Misplon1, Joel Pollen1, Christine Pollnow1, Maia Stack1, David Stack PhD2 and Anita Manogaran PhD3. 1 Community SMART Team of Southeastern WI. 2 University of Wisconsin–Milwaukee 3 Department of Biological Sciences, University of Illinois at Chicago; Chicago, IL Abstract Fatal prion diseases, such as Creutzfeldt-Jakob disease in humans, are associated with the conversion of the normally folded mammalian PrP protein to a misfolded prion form that aggregates. Understanding how prions behave has been greatly facilitated by the study of prions in Saccharomyces cerevisiae, or baker’s yeast. In yeast, the translation release factor, Sup35, misfolds to form the [PSI+] prion. Although the Sup35 protein has a significantly different primary sequence from PrP, its prion form behaves similarly to human prions. The N-terminus of Sup35 is Q/N-rich and responsible for prion formation. Determining the structure of this region has proven difficult since the N-terminus forms aggregates instead of the ordered crystals required for structural studies. However, a small seven amino acid sequence (GNNQQNY) from Sup35's N-terminus was crystallized by Nelson et al. (2005). We have constructed a 3D physical model of GNNQQNY as part of Prion diseases are fatal, transmissible neurodegenerative Healthy brain CJD infected brain the MSOE SMART Teams program using 3D printing technology. GNNQQNY's structure suggests that multiple prion molecules assemble into strong fibrous aggregates through tightly structured interlocking parallel beta sheets called a cross-β spine. The structure of GNNQQNY suggests a potential model of how human prions assemble, which could potentially help in developing therapies that prevent aggregation.. Summary diseases such as Mad Cow Disease and Creutzfeldt-Jakob Disease (CJD) in humans. CJD often has decadelong incubation periods and cannot be effectively diagnosed while in incubation. Brain sections of postmortem CJD patients (lower) exhibit sponge-like holes compared to healthy brain sections (upper). There are currently no cures for prion diseases and prions are resistant to common methods of sterilization. The [PSI+] yeast prion can be used as a model of mammalian prion behavior. Understanding aggregation of the GNNQQNY sequence may lead to possible therapies for human prion diseases. GNNQQNY Aggregate Structure The Q/N rich N-terminus of Sup35, which is responsible for [PSI+] formation, cannot form crystals. Nelson et al. were able to crystallize 7 amino acids (GNNQQNY) from the prion domain. GNNQQNY binds in interlocking antiparallel units, which form a stack held together by hydrogen bonds (left). Model of Prion Formation Prions convert correctly folded proteins to prion form Misfolded proteins form prion aggregates GNNQQNY Sequence from the N-terminus of the [PSI+] Prion 1YJP.pdb Introduction of Foreign Prion Prion Aggregation Prion oligo formation Prions are misfolded proteins that escape the cell’s mechanisms for destroying them. When a prion (red shape, left) is introduced into a cell, it converts correctly folded proteins (blue ovals, left) into the prion form (center). These misfolded proteins assemble into prion aggregates (right). In mammalian prion diseases, the presence of these aggregates are a hallmark of cell death. Using Yeast as a Model Researchers study prions in yeast rather than mammalian prions because of: ● Similar behavior ● Nominal risk ● Yeast have short life cycle ● Yeast are easy to cultivate Budding Yeast Cells (Saccharomyces cerevisiae) Community SMART team Sup35, [PSI+], and Related Research The translational release factor, Sup35, can misfold to form the [PSI+] prion. Fusing the Green Fluorescent Protein (GFP) to Sup35 allows tracking within the cell. [PSI+] appearance is enhanced by increasing the levels of Sup35-GFP in the cell. Before prion induction, Sup35GFP is spread evenly (1. left panel). During initial stages of prion formation, Sup35 forms intermediate ring and line structures (1. center) that eventually turn into fluorescent dots (1. right) that indicate [PSI+] aggregates. 1. 2. When researchers induce [PSI+] formation in cells lacking a cell polarity gene, BEM1 (2.), the intermediate ring stages (2. center) appear normally, but cells do not become [PSI+]. Therefore, bem1 deletion mutants inhibit [PSI+] prion formation. Structure of GNNQQNY molecule: Nelson, R., Sawaya, MR., Balbirnie, M., Madsen, AO., Riekel, C., Grothe, R. & Eisenberg, D. (2005). Structure of the crossβ-spine of amyloid-like fibrils. Nature. 435, 773-778. Brain images: Aguzzi, A., Glatzel, M., Montrasio, F., Prinz, M & Heppner FL. (2001). Interventional strategies against prion disease. Nature Reviews Neuroscience. 2, 745-749. A SMART Team project supported by the National Institutes of Health (NIH) – National Center for Research Resources Science Education Partnership Award (NCRR-SEPA).