* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download FinalReview
Citric acid cycle wikipedia , lookup
Human digestive system wikipedia , lookup
Signal transduction wikipedia , lookup
Transformation (genetics) wikipedia , lookup
Polyclonal B cell response wikipedia , lookup
Evolution of metal ions in biological systems wikipedia , lookup
Oxidative phosphorylation wikipedia , lookup
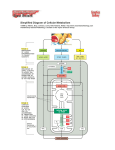
Figure 2.11b Cytoplasmic membrane Endoplasmic reticulum Ribosomes Nucleus Nucleolus Nuclear membrane Golgi complex Cytoplasm Mitochondrion Chloroplast Eukaryote © 2012 Pearson Education, Inc. Figure 2.11a Cytoplasm Nucleoid Ribosomes Plasmid Cell wall Prokaryote © 2012 Pearson Education, Inc. Cytoplasmic membrane Bacterial Structures Microscopic Techniques: Dyes and Staining •Simple stains •Basic dyes with positive charge stick to cells •Acid dyes provide background stain •Differential stains Gram stain - separates bacteria into two categories based on type of cell wall Acid Fast Stain Gram-positive Gram-negative Gram-positive Cell Wall Thick layer of peptidoglycan Teichoic acids Gram-negative Cell Wall Thin layer of peptidoglycan Outer membrane – and cytoplasmic membrane lipopolysaccharide (LPS) Morphology of Prokaryotic Cells: Cell Groupings Flagella - motility E. coli O157:H7 Rotate like a propeller Proton motive force used for energy Presence/arrangement can be used as an identifying marker Cell Wall Provides rigidity to the cell (prevents it from bursting) Cell Wall •Peptidoglycan - rigid molecule; unique to bacteria •Alternating subunits of NAG and NAM form glycan chains •Glycan chains are connected to each other via peptide chains on NAM molecules Cell Wall Cell Wall Gram-negative Thin layer of peptidoglycan Outer membrane - additional membrane barrier; porins permit passage lipopolysaccharide (LPS) - ex. E. coli O157:H7 endotoxin - recognized by innate immune system Cytoplasmic membrane •Defines the boundary of the cell •Semi-permeable; excludes all but water, gases, and some small hydrophobic molecules •Transport proteins function as selective gates (selectively permeable) •Control entrance/expulsion of antimicrobial drugs •Receptors provide a sensor system •Phospholipid bilayer, embedded with proteins Cytoplasmic membrane Electron transport chain - Series of proteins that eject protons from the cell, creating an electrochemical gradient Proton motive force is used to fuel: •Synthesis of ATP (the cell’s energy currency) •Rotation of flagella (motility) •One form of transport If a function of the cell membrane is transport….. • How is material transported in/out of the cell? – Passive transport • No ATP • Along concentration gradient – Active transport • Requires ATP • Against concentration gradient Active Transport Internal Structures: Endospores Enzymes bind substrate and generate a product, enzyme is unchanged Some enzymes require a cofactor to bind substrate Competitive Inhibition Non-competitive Inhibition 6.1. Principles of Metabolism Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display. • Can separate metabolism into two parts CATABOLISM ANABOLISM Energy source (glucose) – Catabolism Cell structures (cell wall, membrane, ribosomes, surface structures) • Processes that degrade compounds to release energy • Cells capture to make ATP Energy Macromolecules (proteins, nucleic acids, polysaccharides, lipids) Energy Subunits (amino acids, nucleotides, sugars, fatty acids) – Anabolism • Biosynthetic processes • Assemble subunits of macromolecules • Use ATP to drive reactions – Processes intimately linked Energy Precursor metabolites Waste products Nutrients (acids, carbon dioxide) (source of nitrogen, sulfur, etc.) Catabolic processes harvest the energy released during the breakdown of compounds and use it to make ATP. The processes also produce precursor metabolites used in biosynthesis. Anabolic processes (biosynthesis) synthesize and assemble subunits of macromolecules that make up the cell structures. The processes use the ATP and precursor metabolites produced in catabolism. Components of Metabolic Pathways • Role of the Chemical Energy Source and the Terminal Electron Acceptor • Some atoms, molecules more electronegative than others Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display. Terminal electron acceptors – Greater affinity for electrons – Energy released when electrons move from low affinity molecule to high affinity molecule H2 • (E.g., glucose to O2) Relative tendency to give up electrons H2S S0 Organic carbon compounds Energy release Organic carbon compounds CO2 SO4 FeOOH Fe2+ NH4+ NO2– ( to form NH4+) NO3– ( to form NH4+) Mn2+ MnO2 Relative tendency to give up electrons Energy sources NO3– ( to form NH2) O2 (a) Energy is released when electrons are moved from an energy source with a low affinity for electrons to a terminal electron acceptor with a higher affinity. ATP is made in catabolic reactions and used in anabolic reactions • 6.3. The Central Metabolic Pathways Transition Step Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display. GLUCOSE – CO2 is removed from pyruvate – Electrons reduce NAD+ to NADH + H+ – 2-carbon acetyl group joined to coenzyme A to form acetyl-CoA – Takes place in mitochondria in eukaryotes 2 Pentose phosphate pathway Starts the oxidation of glucose Yields Glycolysis Oxidizes glucose to pyruvate 1 ~ ~ + Reducing power ATP by substrate-level phosphorylation Pyruvate Yields CO2 Reducing power Biosynthesis Pyruvate 3a Pyruvate NAD+ Acids, alcohols, and gases CoA Transition step CO2 Yields Transition step: CO2 is removed, a redox reaction generates NADH, and coenzyme A is added. Fermentation Reduces pyruvate or a derivative 5 CO2 Reducing power AcetylCoA AcetylCoA NADH + H+ x2 CO2 CoA CO2 3b TCA cycle Incorporates an acetyl group and releases CO2 (TCA cycles twice) Acetyl-CoA 1 The acetyl group is transferred to oxaloacetate to start a new round of the cycle. Yields ~ ~ + Reducing power ATP by substrate-level phosphorylation CoA Respiration Uses the electron transport chain to convert reducing power to proton motive force 4 Yields ~ ~ ATP by oxidative phosphorylation NADH + H+ 2 A chemical rearrangement occurs. Oxaloacetate Citrate A redox reaction generates NADH. 8 NAD+ Isocitrate NAD+ 3 Malate Water is added. 7 A redox reaction generates NADH and CO2 is removed. NADH + H+ H 2O CO2 Fumarate -ketoglutarate NAD+ 4 FADH2 6 CoA A redox reaction generates FADH2- NADH + H+ FAD 5 The energy released during CoA removal is harvested to produce ATP. CoA Succinyl-CoA Succinate CoA ~ ~ ATP ~ + Pi ADP CO2 A redox reaction generates NADH, CO2 is removed, and coenzyme A is added. The Electron Transport Chain—Generating Proton Motive Force • Calculating theoretical maximum yields – In prokaryotes: • • • • Glycolysis: 2 NADH 6 ATP Transition step: 2 NADH 6 ATP TCA Cycle: 6 NADH 18 ATP; 2 FADH2 4 ATP Total maximum oxidative phosphorylation yield = 34 ATP – Slightly less in eukaryotic cells • NADH from glycolysis in cytoplasm transported across mitochondrial membrane to enter electron transport chain – Requires ~1 ATP per NADH generated Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display. GLUCOSE 2 Pentose phosphate pathway Starts the oxidation of glucose Glycolysis Oxidizes glucose to pyruvate 1 Yields ~ ~ + Reducing power ATP by substrate-level phosphorylation Yields Reducing power Biosynthesis 5 Acids, alcohols, and gases Pyruvate Pyruvate 3a Fermentation Reduces pyruvate or a derivative Transition step CO2 CO2 Yields Reducing power AcetylCoA AcetylCoA X2 CO2 CO2 3b TCA cycle Incorporates an acetyl group and releases CO2 (TCA cycles twice) Yields ~ ATP by substrate-level phosphorylation ~ + Reducing power 4 Respiration Uses the electron transport chain to convert reducing power to proton motive force Yields ~ ATP by oxidative phosphorylation ~ Fermentation • The incomplete breakdown of glucose with an organic compound serving as the final electron acceptor • Only pathway operating is glycolysis 6.5. Fermentation • Fermentation end products varied; helpful in identification, commercially useful – Ethanol – Butyric acid – Propionic acid • 2,3-Butanediol • Mixed acids Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display. Pyruvate Fermentation pathway Microorganisms End products Lactic acid Ethanol Butyric acid Propionic acid Mixed acids 2,3-Butanediol Streptococcus Lactobacillus Saccharomyces Clostridium Propionibacterium E. coli Enterobacter Lactic acid Ethanol CO2 Butyric acid Butanol Acetone Isopropanol CO2 H2 Propionic acid Acetic acid CO2 Acetic acid Lactic acid Succinic acid Ethanol CO2 H2 CO2 H2 (yogurt, dairy, pickle), b (wine, beer), (acetone): © Brian Moeskau/McGraw- Hill; (cheese): © Photodisc/McGraw-Hill; (Voges-Proskauer Test), (Methyl-Red Test): © The McGraw-Hill Companies, Inc./Auburn University Photographic Services The Electron Transport Chain—Generating Proton Motive Force • Components of an Electron Transport Chain – Most carriers grouped into large protein complexes • Serve as proton pumps • Three general groups are notable – Quinones • Lipid-soluble molecules • Move freely, can transfer electrons between complexes – Cytochromes • Contain heme, molecule with iron atom at center • Several types – Flavoproteins • Proteins to which a flavin is attached • FAD, other flavins synthesized from riboflavin Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display. Fig. 6.20 Prokaryotic cell Cytoplasmic membrane Electron Transport Chain NADH dehydrogenase Uses of Proton Motive Force Ubiquinol veoxidase force rive: H+ (2 or 4) H+ (0 or 4) Ubiquinone Path of electrons ATP synthase (ATP synthesis) 10 Active transport (one mechanism) H+ Rotation of a flagella H+ H+ Proton motive force is used to drive: Transported molecule Outside of cytoplasmic membrane 2 e– – Cytoplasm Succinate dehydrogenase NADH + NAD+ 2 H+ 1/ H2O 2 O2 Terminal electron acceptor H+ 3 ATP + 3 Pi 3 ADP Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display. Glycolysis Pentose phosphate pathway Glucose 6-phosphate Fructose 6-phosphate Lipopolysaccharide (polysaccharide) Ribose 5-phosphate Erythrose 5-phosphate Nucleotides amino acids (histidine) Amino acids (phenylalanine, tryptophan, tyrosine) Peptidoglycan Dihydroxyacetone phosphate Lipids (glycerol component) 3-phosphoglycerate Amino acids (cysteine, glycine, serine) Anabolic Pathways— Synthesizing Subunits from Precursor Molecules Phosphoenolpyruvate Amino acids (phenylalanine, tryptophan, tyrosine) Pyruvate Pyruvate Acetyl-CoA Acetyl-CoA Amino acids (alanine, leucine, valine) Lipids (fatty acids) Oxaloacetate Amino acids (aspartate, asparagine, isoleucine, lysine, methionine, threonine) X2 - ketoglutarate TCA cycle Amino acids (arginine, glutamate, glutamine, proline) • Role of Electron Carriers – Energy harvested in stepwise process • Electrons transferred to electron carriers, which represent reducing power (easily transfer electrons to molecules) – Raise energy level of recipient molecule • NAD+/NADH, NADP+/NADPH, and FAD/FADH2 Microbial Growth • Growth= an increase in the number of cells, not an increase in size • Generation=growth by binary fission • Generation time=time it takes for a cell to divide and the population to double 4.2. Prokaryotic Growth in Nature • Microorganisms historically studied in laboratory • But dynamic, complex conditions in nature have effects on microbial growth, behavior – Cells sense changes, adjust to surroundings – Synthesize compounds useful for growth – Can live singly • Most live in polysaccharideencased communities • Termed biofilms • Cause slipperiness of rocks in stream bed, slimy “gunk” in sink drains, scum in toilet bowls, dental plaque Biofilms • Formation of biofilm Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display. Planktonic bacteria move to the surface and adhere. Bacteria multiply and produce extracellular polymeric substances (EPS). Other bacteria may attach to the EPS and grow. Cells communicate and create channels in the EPS that allow nutrients and waste products to pass. Some cells detach and then move to other surfaces to create additional biofilms. Bacteria divide by binary fission Calculating cell number over time t=time; 0=cell number at start; n= number of divisions based on generation time Nt=N0 x 2n The Growth Curve • Growth curve characterized by five stages Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display. Stationary phase Number of cells (logrithmic scale) 1010 Death phase 108 106 Log or exponential phase Phase of prolonged decline 104 102 Lag phase 100 Time (hr) (days/months/years) Primary and Secondary metabolites Some factors that influence growth in foods…temperature • Remember that some microbes grow well at cooler temperature, others more slowly Some of the factors that influence growth in foods… Water Availability (aw) Food (aw) Microbe Minumum (aw) Fresh meat 0.99 Spoilage microbes 0.91 Hot dog 0.92 Pseudomonas 0.97 Ham 0.91 Staphylococcus aureus 0.86 Dried fruit 0.72-0.8 Yeasts 0.81 Molds 0.80 Some factors that influence growth in foods….pH Foods pH of food Microbe Minimum pH of microbe beef 5.5 Most spoilage microbes 4.0 milk 6.3 molds 1.5 spinach 5.5 yeast 2.5 apples 3.0 E. coli 4.0 Microbes in food production • Lactic Acid bacteria • Yeasts…Saccharomyces cerevisiae • Molds Overview of Digestion Fig. 24.1 Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display. Oral cavity containing tongue and teeth Dental caries Periodontal disease Parotid salivary gland Mumps Stomach Gastritis Gastric ulcer Gallbladder Function Oral cavity Obtains and processes food Salivary glands Secrete saliva Esophagus Transports food to stomach Stomach Stores food; mechanical digestion; breaks down some proteins Pancreas Secretes digestive enzymes Liver Produces bile to assist in fat digestion Salivary glands Esophagus Esophagitis Liver Hepatitis Organ Pancreas Pancreatitis Small intestine Enteritis Duodenal ulcer Appendix Appendicitis Large intestine Dysentery Colitis Rectum Anus Food molecules Gallbladder Stores bile until needed Small intestine Site of most digestion and absorption of nutrients Large intestine Absorbs some water and minerals; prepares waste Villus Epithelial cells Microvilli Capillaries Lymphatic vessel Smooth muscle Nerve fibers Upper digestive tract Lower digestive tract How do organisms cause food poisoning? • Food borne intoxication: bacteria grow within the food and produce toxins, the toxins are what lead to food poisoning symptoms • Examples: Clostridium botulinum Staphylococcus aureus Mechanisms of pathogenesis • Attachment: pili or adhesins • Toxin production: two kinds of toxins 1)increase secretion of water and electrolytes 2)cause cell death • Alterations in host cells • Cell invasion Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display. Features of intestinal infections 1. Attachment—pili and adhesins 2. Toxin production 1. Enterotoxins cause release of electrolytes 2. Cytotoxins cause cell death 3. Alteration of epithelial cells 1. Attaching and effacing 2. Type III secretion systems 4. Cell invasion Toxin production Cl– Na+ H2O Cytotoxins cause cell death. Toxins absorbed into the bloodstream result in systemic effects. Enterotoxins increase secretion of water and electrolytes. Alterations in the host cells Pedestal Inject effector proteins Attachment and effacing (A/E) lesions formed after bacterium injects various effector proteins. One protein functions as a receptor for the bacterium. Another induces rearrangement of actin filaments, resulting in the formation of a pedestal under the bacterium. Cell invasion Bacterium is engulfed and multiplies within host cell. An effector protein injected by the bacterium induces the engulfment by causing rearrangement of host cell actin. Type III Secretion System Membrane ruffling Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display. Shigella enters via M cells A-B toxin on chromosome. bad. Causes HUS, hemolytic uremic syndrome. 1 Epithelial cell M cell Intestinal lumen Macrophage Shigella cells 2 Dead macrophage Bacteria make it into blood vessels, and 3 kill the endothelial cells. A-B toxin: B-binds cell, A goes into cell—causes cell death. 4 M cells take up Shigella cells and transport them across the epithelium. They multiply in the macrophages that ingest them, leading to death of that host cell. Shigella cells attach to the base of the epithelial cells and induce those cells to take them in. From there, they escape the endosome and multiply in the cytoplasm. Shigella cells cause the host cell actin to polymerize. This forms an “actin tail” that propels a bacterium within the host cell, sometimes with enough force to move it into a neighboring cell. Neutrophils Infected epithelial cells die and slough off. An intense inflammatory response leads to bleeding and abscess formation. Courtesy of Philippe J. Sansonette, M.D., Professeur Institut Pasteur Shiga-toxin E. coli (STEC) • Obtain from the consumption of animal products • Attacks the colon, produce A/E lesions • Produces Shiga toxins • O157:H7 causes bloody diarrhea which may lead to hemolytic uremic syndrome Diarrhea causing E. coli • Classified according to virulence – Entertoxigenic E. coli (ETEC) – Enterpathogenic E. coli (EPEC) – Shiga toxin-producing E. coli (STEC) – Enteroinvasive E. coli (EIEC) – Enteroaggregative E. coli (EAEC) – Diffusely adhering E. coli (DAEC) Vibrio cholerae • Causative agent of cholera • General Characteristics: Curved gram negative rod, facultative anaerobe, single polar flagella, pili • Can exist in saltwater for extended periods of time, halotolerant • Different serotypes based on O antigen, O1 is current serotype circulating, O antigen is part of the lipopolysaccharide. Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display. Cholera Fig. 24.1 Vibrio cholerae . a marine bacterium, halotolerant, and tolerant to alkaline conditions. Gr- curved rod. Oral cavity containing tongue and teeth Dental caries Periodontal disease Parotid salivary gland Mumps NOT acid tolerant—high infective dose Stomach Gastritis Gastric ulcer Gallbladder Function Oral cavity Obtains and processes food Salivary glands Secrete saliva Esophagus Transports food to stomach Stomach Stores food; mechanical digestion; breaks down some proteins Pancreas Secretes digestive enzymes Liver Produces bile to assist in fat digestion Salivary glands Esophagus Esophagitis Liver Hepatitis Organ Pancreas Pancreatitis Small intestine Enteritis Duodenal ulcer There have been 7 pandemics of cholera Appendix Appendicitis Large intestine Dysentery Colitis Rectum Anus Food molecules Colonizes the small intestine, binds to cells via pili and an A-B toxin causes electrolytes to leave the cells. Gallbladder Stores bile until needed Small intestine Site of most digestion and absorption of nutrients Large intestine Absorbs some water and minerals; prepares waste Villus Epithelial cells Microvilli Capillaries Lymphatic vessel Smooth muscle Nerve fibers Upper digestive tract Lower digestive tract A-B toxin Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display. B binds to cell A entersV. cholerae bacterium 1 The A-B toxin’s B subunit attaches to receptors on cell membrane; the A subunit enters the cell. B 2 The A subunit locks a G protein in the “active” mode, turning on adenylate cyclase. 4 A OFF 10 µm Plasma membrane of intestinal cell A G protein ON cAMP activates ion transport channels in the membrane causing Cl– and other electrolytes to pour out of the cell. ATP Cl– 3 Adenylate cyclase causes the conversion of ATP to cAMP. Adenylate cyclase K+ Na+ ● HCO3– H2O cAMP 5 Water follows electrolytes out of the cell by osmosis. © VeronikaBurmeister/Visuals Unlimited Infections of the GI tract that are not from food spoilage • Helicobacter pylori can colonize the stomach. • Gr-, curved rod with polar, sheathed flagella • Many people have H. pylori with no symptoms (1 in 5). However, some strains can cause stomach cancer and 90% of stomach cancer patients have H. pylori in their stomach. How does H. pylori colonize the stomach? It’s sterile, right? • High Acid – H. pylori survives acids by using urease to transform urea into ammonia, creating an alkaline pocket • Thick Mucus layer protects stomach surface cells (epithelium) – H. pylori cells have sheathed flagella that allow them to swim down into the thick mucus (less acidic under the mucus). Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display. • Clostridium difficile Fig. 24.1 Gr+, rod, spore forming, obligate Parotid anaerobe. salivary gland Mumps Oral cavity Produces containingcytotoxins tongue and teeth •DentalMember of gut microbiome, in low numbers, caries Salivary glands Organ Function Oral cavity Obtains and processes food Salivary glands Secrete saliva Esophagus Transports food to stomach Stomach Stores food; mechanical digestion; breaks down some proteins Pancreas Secretes digestive enzymes Liver Produces bile to assist in fat digestion Periodontal disease --It most commonly occurs in patients in Esophagus hospitals on antibiotic therapy. Esophagitis • Can also be acquired Difficult to kill with disinfectants (spores) Liver Hepatitis Stomach Gastritis Gastric ulcer Gallbladder Pancreas • Mild to severe symptoms includingPancreatitis colitis Small(inflammation intestine of the colon). Enteritis Large intestine ulcer •Duodenal Treatment: Often stopping the antibiotics, if Dysentery Appendix possible, alleviates the problem. Colitis Appendicitis Rectum Gallbladder Stores bile until needed Small intestine Site of most digestion and absorption of nutrients Large intestine Absorbs some water and minerals; prepares waste Anus Food molecules Villus Epithelial cells Microvilli Capillaries Lymphatic vessel Smooth muscle Nerve fibers Upper digestive tract Lower digestive tract Control of Microbial Growth A few terms • Bacteriostatic: inhibits bacterial growth • Bactericidal: something capable of killing bacteria • Antiseptic: an agent that is used to inhibit/kill bacterial growth on skin and mucus membranes • Disinfectant: an agent that is used to inhibit/kill bacterial growth on inanimate objects What parts of a bacterial cell are sensitive to physical treatments and chemicals? • Plasma membrane • DNA and proteins Are all microbes equally sensitive? What level of microbial control or elimination is needed ? Appropriate procedures depend on 4 main features: 1. Type and number of microbes (are there spores? What kind? Are there pathogens? ) Bacterial endospores Mycobacterium, Pseudomonas sp. Protozoan cysts and oocysts (Giardia lambia and Cryptosporidium parvum). Naked viruses lack a lipid envelope, this makes them more resistant. HIV—with lipid envelop—is very sensitive. 2. Environmental conditions 3. Risk of infection 4. Composition of the item to be treated. Physical Methods Moist Heat Dry Heat Filtration Radiation electromagnetic ionizing High Pressure Chemical Control Alcohol Aldehydes Biguanides Ethyline Oxide Gas Metals Ozone Peroxygens Phenolics Quaternarey Ammonium Compounds Gene transfer in bacteria • There are three types of gene transfer 1. Transformation—DNA enters and is incorporated (cell imports it) 2. Conjugation—DNA moves from one bacterial cell to another during cell to cell contact 3. Transduction—a virus injects DNA into the bacterial cell All types of gene transfer • Involve unidirectional transfer of information (donor-->recipient) • Require the integration of newly acquired DNA “homologous recombination” • Increases genetic diversity Terms to remember • Replicon – DNA containing an origin of replication (O.R.) This allows the DNA to be copied. If no O.R. then it must be incorporated into the chromosome for replication. • Homologous recombination – DNA has parts that match the DNA in the chromosome, and bind to it to be incorporated into the chromosome. • Competent– A bacterial cell that can take up DNA from the environment is termed ‘competent’ • Naked DNA—outside of the cell, or virus Conjugation • Transfer of genes between 2 bacterial cells • Gram negative cells use a sex pilus • F(+) cells have F plasmid, F(-) lack F plasmid Conjugation between (F+) and F() cells First, the F pilus binds to specific receptor on the F- cell. Pilus retracts (gets shorter) and brings the cells closer. The F plasmid requires an Origin of Transfer—we will refer to this as the OoT!! Without the OoT, DNA can not be moved using the F pilus. The OoT is nicked to open the DNA and one single strand of the plasmid moves through the pilus to the other cell. Then the complement of each of the ssDNA plasmids is made, and voila! 2 F+ cells. Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display. Transformation There are structures bacterial cells use to bind and import DNA. Gene conferring StrS 1 Recipient chromosome Gene Conferring StrR Double-stranded DNA binds to the surface of a competent cell. 2 Single strand enters the cell; the other strand is degraded. These include the type 4 pili aparatus, which are also used for making pili. Some bacteria take in any DNA, while others stick to specific DNA sequences and take in only DNA they want. 3 The strand integrates into the recipient cell’s genome by homologous recombination. 4 Streptomycin-sensitive daughter cell Streptomycin-resistant daughter cell After replicating the DNA, the cell divides. 5 Non-transformed cells (StrS) die on streptomycin-containing medium, whereas transformed cells (StrR) can multiply. Transduction • Transfer of genes from a phage to bacterial cell • Generalized transduction: occurs with lytic or lysogenic phage (section 8.7) • Specialized transduction: occurs with lysogenic phage (section 13.3) Plasmids-types • Can be broad host range • Or specific to particular species • Some can be maintained within the same cell as others (Arranged in groups by compatibility) • Some can not, and are not compatible • High copy number vs. low copy number • Conjugative plasmids, carry genes needed for conjugation • Mobilizable plasmids have an OoT, but not the conjugation genes. If conjugative and mobilizable are together, they can both be moved to a new cell. Resistance Plasmids (R plasmids) Transposons…way to move genes between organisms Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display. Insertion sequence Mobile element Transposase gene Inverted repeat 5′ 3′ Inverted repeat 3′ T C G A T G… A G C T A C… 5′ 5′ 3′ …C A T C G A ....G T A G C T 3′ 5′ Composite transposon Mobile element Insertion sequence Antibiotic-resistance gene Insertion sequence Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display. Vancomycinresistance gene (encoded on a transposon on a plasmid) • How did this S. aureus become Vancomycin resistant S. aureus (VRSA)? Plasmid Staphylococcus aureus sensitive to vancomycin Enterococcus faecalis resistant to vancomycin Enterococcus faecalis plasmid transferred by conjugation Transposon jumps from one plasmid to another. Plasmid from Enterococcus faecalis is destroyed. Vancomycin-resistant Staphylococcus aureus