* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Lecture 21 : Introduction to Neutral Theory
Public health genomics wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Genome (book) wikipedia , lookup
Gene therapy wikipedia , lookup
Gene desert wikipedia , lookup
Genetic drift wikipedia , lookup
Gene expression profiling wikipedia , lookup
Viral phylodynamics wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Genome evolution wikipedia , lookup
Koinophilia wikipedia , lookup
Gene nomenclature wikipedia , lookup
Polymorphism (biology) wikipedia , lookup
The Selfish Gene wikipedia , lookup
Genetics and archaeogenetics of South Asia wikipedia , lookup
Human genetic variation wikipedia , lookup
Helitron (biology) wikipedia , lookup
Computational phylogenetics wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Gene expression programming wikipedia , lookup
Designer baby wikipedia , lookup
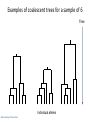
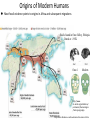
Lecture 23 : Introduction to Coalescence April 7, 2014 Last Time Introduction to phylogenetics Phylogeography Limitations of phylogenetic analysis Today Gene trees versus species trees Coalescence Influence of demographic factors on coalescence times Coalescence and human origins Gene Trees vs Species Trees Genes (or loci) evolve at different rates Why? Topology derived by a single gene may not match topology based on whole genome, or morphological traits Gene Tree B C A Coalescence Retrospective tracing of existing alleles to a common ancestral allele A reverse reconstruction of the evolution of modern variation Allows explicit simulation of sequence evolution Incorporation of factors that cause deviation from neutrality: selection, drift, and gene flow 9 generations in the history of a population of 14 gene copies Time present Slide courtesy of Yoav Gilad Individual alleles Gene Trees vs Species Trees Failure to coalesce within species lineages drives divergence of relationships between gene and species trees Divergent Gene Tree: Concordant Gene Tree b is closer to a than to c a b c b is closer to c than to a a b c How to model this process? Modeling from Theoretical Ancestors: Forward Evolution Can model populations in a forward direction, starting with theoretical past Fisher-Wright model of neutral evolution Very computationally intensive for large populations Alternative: Start at the end and work your way back Most recent common ancestor (MRCA) Time present Slide courtesy of Yoav Gilad Individual alleles The genealogy of a sample of 5 gene copies Most recent common ancestor (MRCA) Time present individuals Slide courtesy of Yoav Gilad The genealogy of a sample of 5 gene copies Most recent common ancestor (MRCA) Individual alleles Slide courtesy of Yoav Gilad Time present Examples of coalescent trees for a sample of 6 Time Individual alleles Slide courtesy of Yoav Gilad Coalescence Advantages Don’t have to model dead ends Only consider lineages that survive to modern day: computationally efficient Based on actual observations Can simulate different evolutionary scenarios to see what best fits the observed data Coalescent Tree Example Coalescence: Merging of two lineages in the Most Recent Common Ancestor (MRCA) Waiting Time: time to coalescence for two lineages Increases with each coalescent event Probability of Coalescence For any two lineages, function of population size Pcoalescence 1 2Ne Also a function of number of lineages Pcoalescence k (k 1) 1 2 2Ne where k is number of lineages Probability of Coalescence Probability declines over time Lineages decrease in number Can be estimated based on negative exponential Pcoalescence e k ( k 1) 1 t 2 2 Ne where k is number of lineages Time to Coalescence Affected by Population History Bottleneck Time to Coalescence Affected by Population History Population Growth How will population structure affect coalescence times? Time to Coalescence Affected by Population Structure Applications of the Coalescent Approach Framework for efficiently testing alternative models for evolution Inferences about effective population size Detection of population structure Signatures of selection (coming attraction) Reconstructing history of populations Origins of Modern Humans Most fossil evidence points to origins in Africa and subsequent migrations Skulls found in Omo Valley, Ethiopia Dated at ~195K Omo 1 Modern http://wwwv1.amnh.org/exhibitions/ permanent/humanorigins /history/origin.php http://www.dhushara.com/book/unraveltree/unravel.htm Human Phylogeography: mtDNA Most ancient and diverse haplotypes in Africa (dots) Migration and admixture is evident from presence of African haplotypes in other clades Complexities to Human Phylogeography Some genes show evidence of Asian origin Sequence of X-linked ribonucleotide reductase M2 pseudogene 4 (RRM2P4) Garrigan 2007 Nature Reviews Genetics 7:669 Why might some X-linked genes show a human origin in Africa (e.g., PDHA1), while others suggest an Asian origin e.g., (RRM2P4)? Evidence of Population Structure in Ancient Humans Garrigan and Hammer 2006 Nature Reviews Genetics 7:669 Time to Coalescence Affected by Population Structure Evidence for Ancient Population Structure in Nuclear but not Mitochondrial Trees Garrigan and Hammer 2006 Nature Reviews Genetics 7:669 Why does mitochondrion show shorter coalescence times than nuclear loci? Why does rate vary much more for nuclear loci?