* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download File
Gene expression profiling wikipedia , lookup
Adeno-associated virus wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Metagenomics wikipedia , lookup
Deoxyribozyme wikipedia , lookup
DNA supercoil wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Primary transcript wikipedia , lookup
Cancer epigenetics wikipedia , lookup
Epigenomics wikipedia , lookup
Minimal genome wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Point mutation wikipedia , lookup
Non-coding DNA wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Genome (book) wikipedia , lookup
Neocentromere wikipedia , lookup
DNA vaccination wikipedia , lookup
Genetic engineering wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Genome evolution wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Helitron (biology) wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Microevolution wikipedia , lookup
Molecular cloning wikipedia , lookup
Designer baby wikipedia , lookup
History of genetic engineering wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
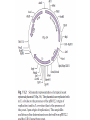
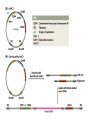
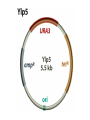
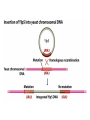
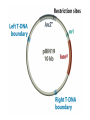
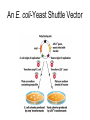
Unit I Lecture 4 B. Tech. (Biotechnology) III Year V th Semester EBT-501, Genetic Engineering EBT 501, Genetic Engineering Unit I Gene cloning -concept and basic steps; application of bacteria and viruses in genetic engineering; Molecular biology of E. coli and bacteriophages in the context of their use in genetic engineering, Cloning vectors: Plasmid cloning vector PBR322, Vectors for cloning large piece of DNA; –Bacteriophage-l and other phage vectors; Cosmids, Phagemids; YAC and BAC vectors, Model vectors for eukaryotes – Viruses, Unit II Restriction modification, enzymes used in recombinant DNA technology endonucleases, ligases and other enzymes useful in gene cloning, PCR technology for gene/DNA detection, cDNA, Use of Agrobacterium for genetic engineering in plants; Gene libraries; Use of marker genes. Cloning of foreign genes: DNA delivery methods -physical methods and biological methods, Genetic transformation of prokaryotes: Transferring DNA into E. coli –Chemical induction and Electroporation, Unit III Gene library: Construction cDNA library and genomic library, Screening of gene libraries – screening by DNA hybridization, immunological assay and protein activity, Marker genes: Selectable markers and Screenable markers, nonantibiotic markers, Gene expression in prokaryotes: Tissue specific promoter, wound inducible promoters, Strong and regulatable promoters; increasing protein production; Fusion proteins; Translation expression vectors; DNA integration into bacterial genome; Increasing secretions; Metabolic load, Recombinant protein production in yeast: Saccharomyces cerevisiae expression systems; Mammalian cell expression vectors: Selectable markers; Unit IV Origins of organismal cloning in developmental biology research on frogs; nuclear transfer procedures and the cloning of sheep (Dolly) & other mammals; applications in conservation; therapeutic vs. reproductive cloning; ethical issues and the prospects for human cloning; Two-vector expression system; two-gene expression vector, Directed mutagenesis; transposon mutagenesis, Gene targeting, Site specific recombination Unit V General principles of cell signaling, Extracellular signal molecule and their receptors, Operation of signaling molecules over various distances, Sharing of signal information, Cellular response to specific combinations of extracellular signal molecules; Different response by different cells to same extracellular signal molecule, NO signaling by binding to an enzyme inside target cell, Nuclear receptor; Ion channel linked, G-protein- linked and enzyme-linked receptors, Relay of signal by activated cell surface receptors via intracellular signaling proteins, Intracellular signaling proteins as molecular switches, Interaction between modular binding domain and signaling proteins, Remembering the effect of some signal by cells. Unit I • Gene cloning -concept and basic steps • Application of bacteria and viruses in genetic engineering • Molecular biology of E. coli and bacteriophages in the context of their use in genetic engineering • Cloning vectors: Plasmid cloning vector pBR322, • Vectors for cloning large piece of DNA – Bacteriophage-l and other phage vectors – Cosmids – phasmids – Phagemids – BAC vectors – PAC vectors – YAC vectors • Model vectors for eukaryotes - Viruses, Table 2.4 Genomes 3 (© Garland Science 2007) Cloning in Saccharomyces cerevisiae • In 1996 the sequencing of the entire 12 Mb genome of S. cerevisiae was completed and most, if not all, of the genes have been identified. • fungi are not naturally transformable and artificial means have to be used for introducing foreign DNA. • One method involves the use of spheroplasts (i.e. wallless cells) and was first developed for S. cerevisiae (Hinnen et al. 1978). • In this method, the cell wall is removed enzymically and the resulting spheroplasts are fused with ethylene glycol in the presence of DNA and CaCl2 Reasons for cloning DNA in S. cerevisiae • At that time the primary purpose of cloning was to understand what particular genes do in vivo • to understand those cellular functions unique to eukaryotes such as mitosis, meiosis, signal transduction, obligate cellular differentiation • offer a number of advantages, such as the ability to glycosylate protein, the absence of pyrogenic toxins, and in the case of the methylotrophic yeast Pichia pastoris, the ability to get very high yields of recombinant proteins. • ability to clone very large pieces of DNA • Certain yeast vectors can accept inserts greater than 1 Mb, much greater than those found in BACs and PACs • All contain unique target sites for a number of restriction endonucleases. • Secondly, they can all replicate in E. coli, often at high copy number. • all employ markers that can be selected readily in yeast and which will often complement the corresponding mutations in E. coli as well. • The four most widely used markers are His3, Leu2, Trp1, and Ura3. Kinds of vectors have been developed for use in S. cerevisiae Four types of plasmid vector have been developed: • yeast episomal plasmids (YEps) • yeast replicating plasmids (YRps) • Yeast centromere plasmids (Ycps) • yeast artificial chromosomes (YACs). Yeast episomal plasmids • YEps were first constructed by Beggs (1978) by recombining an E. coli cloning vector with the naturally occurring yeast 2 μm plasmid. • This plasmid is 6.3 kb in size, has a copy number of 50–100 per haploid cell and has no known function. A representative YEp is shown in Fig. 11.2. Yeast replicating plasmids • YRps were initially constructed by Struhl et al. (1979). • carry sequences that enable E. coli vectors to replicate in yeast cells. • sequences are known as ars autonomously replicating sequences). • An ars is quite different from a centromere: • The ars acts as an origin of replication • Centromer is involved in chromosome segregation Yeast centromere plasmids • plasmids carrying an ars, most of the recombinants were unstable in yeast • plasmid-borne centromere sequences have the same distinctive chromatin structure that occurs in the centromere region of yeast chromosomes (Bloom & Carbon 1982) • YCps exhibit three characteristics of chromosomes in yeast cells. • First, they are mitotically stable in the absence of selective pressure. • Secondly, they segregate during meiosis in a Mendelian manner. • Finally, found at low copy number in the host cell. Yeast artificial chromosomes • All three autonomous plasmid vectors described above are maintained in yeast as circular DNA molecules, even the YCp vectors, which possess yeast centromeres. • Thus, none of these vectors resembles the normal yeast chromosomes, which have a linear structure. • The ends of all yeast chromosomes, like those of all other linear eukaryotic chromosomes, have unique structures that are called telomeres. • Telomere structure has evolved as a device to preserve the integrity of the ends of DNA molecules, which often cannot be finished by the conventional mechanisms of DNA replication. • In 1982 Szostak & Blackburn developed the first vector which could be maintained as a linear molecule, thereby mimicking a chromosome, by cloning yeast telomeres into a YRp. • Such vectors are known as yeast artificial chromosomes (YACs). • The method for cloning large DNA sequences in YACs developed by Burke et al. (1987) is shown in next slide Fig 11.3. Yeast Artificial Chromosomes (YACs) • Genetically engineered yeast minichromosomes. • Accept foreign DNA inserts of 200-500 kb. • Contain a yeast origin of replication, yeast centromere, two yeast telomeres, a selectable marker, and a polycloning site. • Recombinogenic engineering can be used to move genes from one vector to another Promoter systems have been developed to facilitate overexpression of recombinant proteins in yeast • The first overexpression systems developed were for S. cerevisiae and used promoters from genes encoding abundant glycolytic enzymes, e.g. alcohol dehydrogenase (ADH1), PGK or glyceraldehyde-3- phosphate dehydrogenase (GAP). • These are strong promoters and mRNA transcribed from them can accumulate up to 5% of total. • They were at first thought to be constitutive but later were shown to be induced by glucose • The ideal promoter is one that is tightly regulated so that the growth phase can be separated from the induction phase. • This minimizes the selection of non-expressing cells and can permit the expression of proteins normally toxic to the cell. • ideal promoter will also have a high induction ratio. • One promoter which has these characteristics and which is now the most widely used is that from the GAL1 gene. • Galactose regulation in yeast is now extremely well studied and has become a model system for eukaryotic transcriptional regulation specialist multi-purpose vectors have been developed for use in yeast • incorporate the useful features found in the corresponding E. coli vectors e.g. an f1 origin to permit sequencing of inserts, production of the cloned gene product as a purification fusion Heterologous proteins can be synthesized as fusions for display on the cell surface of yeast • used to detect protein–ligand interactions • to select mutant proteins with altered binding capacity Figure 2.25 Genomes 3 (© Garland Science 2007) Figure 2.25a Genomes 3 (© Garland Science 2007) Figure 2.25b Genomes 3 (© Garland Science 2007) Figure 2.26a Genomes 3 (© Garland Science 2007) Figure 2.26b Genomes 3 (© Garland Science 2007) Figure 2.27 Genomes 3 (© Garland Science 2007) An E. coli-Yeast Shuttle Vector BACs and PACs are vectors that can carry much larger fragments of DNA than cosmids because they do not have packaging constraints Figure 2.24 Genomes 3 (© Garland Science 2007) The PAC Mammalian Shuttle Vector pJCPAC-Mam1 Episomes • An episome is a genetic element that is not essential to the host and that can either replicate autonomously or be integrated into the bacterial chromosome. • Integration depends on the presence of IS elements.