* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download proteinCompression
Community fingerprinting wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Genetic code wikipedia , lookup
Western blot wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Non-coding DNA wikipedia , lookup
Protein (nutrient) wikipedia , lookup
Gene expression wikipedia , lookup
Protein adsorption wikipedia , lookup
Proteolysis wikipedia , lookup
Ancestral sequence reconstruction wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Molecular evolution wikipedia , lookup
Protein moonlighting wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
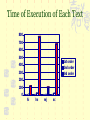
Compression of Protein Sequences EE-591 Information Theory FEI NAN, SUMIT SHARMA May 3, 2003 GENERAL OVER VIEW OF THE PROJECT DNA and Protein form the basic structure of the life and they can be represented by any normal text file. When it comes to compression, we see that they don’t show the good result from the normal compression techniques. In this project we have used a new technique called CP (Compression Scheme) for their compression and analysis it how far it is good in Protein Compression. THE NEED OF THE COMPRESSION There are two different motivation for the compression Compression enables efficient use of the resource such as storage and bandwidth From scientific perspective it provides a way of capturing and quantifying structure in the sequence We here have put stress on the second one as we are here dealing with the biological sequence A good model for compression will contain a few symbols with high probability (and preferably one dominant symbol), thus allowing very compact coding of those probable symbols. We see that DNA is the genetic material of the life it transmits information from one generation to another We can represent DNA as sequence of the symbol represented by four symbol alphabet of nucleotides A adenine C cytosine G guanine T thymine A will only bond with T and G will only bond with C DNA STRUCTURE DNA STRUCTURE Protein Structure Since we know that Protein sequence is a very large one as it is composed of 20 amino acid so they have a high level of redundancy since we can represent the sequence in a normal text file. But when we talk abut the compression we have to take care of the fact they are biological sequence and protein are subjected to mutation that destroy repetition. 3-D Protein Structure PROTEIN STRCTURE The redundancy in protein majorly comes through two sources New genes arise through duplication Mistake made while copying DNA and other cellular process In compression of the protein we take into account a distance metric this distance reflect their mutation probabilities that is symbol that are close together are derived from the same symbol by mutation and if far apart other wise . In our scheme we have taken distance to combine the prediction made by different context ,we sum up over all possible context up to a certain length weighted by their similarity to the current context. In our project we have make use of the following concepts PPM Adaptive coding PROBABILITIC PREDICTION METHOD(PPM) The basic idea of the PPM is to used last few character to predict the probabilities of the coming one . Example –we have a sequence abcd that end with aba so PPM will calculate the previous occurrence of the aba and tallies with the symbol that will occur next CP COMPRESSION SCHEME In this scheme we take into consideration that biological sequence constantly undergo into mutation and as long as the new sequence has similar properties the mutation will be accepted ,thus exact repletion is overlaid with mutation which is modeled by the distance function ,thus it is desirable to take into account in compression scheme In general this is given by the formula We see in the equation that weight frequent context more highly this has some merits as it has less variance in distribution thus to improve this more we take the weight context equally ,we do that by converting f Explanation of the program Calculation of the first order Calculation of the second order Calculation of the third order Major Function There are major functions in the code. 1. 2. 3. 4. 5. Function to read the mutation matrix Function to read the target sequence Function to compute the n-order value Function to compute the dynamic probability based on the CP algorithm (a major part distinct from some other existing algorithm of the 1987 CACM article by Witten, Neal, and Cleary. ) Main function Modules in the Code There is a function to calculate the dynamic probability of each symbol. i.e. When we read a new symbol, we increase the occurrence of that symbol by 1 and increase the total number of symbols we have read by 1 and use this function module to compute the probability dynamically. ADAPTIVE CODING Adaptive coding doesn’t require the probabilities to be transmitted with the encoded data Require only one pass through the data coding Doesn’t use the fixed symbol probabilities Time of Execution of Each Text 800 700 600 500 1st order 2nd order 3rd order 400 300 200 100 0 hi hs mj sc Time of Execution of Each Text As we can see from the diagram in the last slide, the time for the successful execution of n-order CP is increased with the n by a factor of 20*. So we cannot make n too large although we might receive a good compression ratio. *Craig G, Nevill-Manning, Ian H. Witten Protein is incompressible Improvements Improvement of the execution time. Improvement of the float precision. Solution By addition of high-performance hardware By Optimization of software algorithm Thanks ?