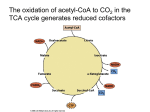
* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Bio 226: Cell and Molecular Biology
Survey
Document related concepts
Artificial gene synthesis wikipedia , lookup
Biochemistry wikipedia , lookup
Mitochondrion wikipedia , lookup
Magnesium in biology wikipedia , lookup
Metalloprotein wikipedia , lookup
Photosynthesis wikipedia , lookup
NADH:ubiquinone oxidoreductase (H+-translocating) wikipedia , lookup
Microbial metabolism wikipedia , lookup
Citric acid cycle wikipedia , lookup
Evolution of metal ions in biological systems wikipedia , lookup
Electron transport chain wikipedia , lookup
Photosynthetic reaction centre wikipedia , lookup
Adenosine triphosphate wikipedia , lookup
Transcript
Next Assignment? In lab time Friday March 27 or in class starting March 27 or March 30? •GMO plants? • Herbicide resistance • Pathogen/herbivore resistance • Improving nutrition • Making vaccines, other useful biochems •Plant/Algal biofuels? PSI and PSII work together in the “Z-scheme” Light absorbed by PS II makes ATP Light absorbed by PS I makes reducing power Ultimate e- source O2 released? Terminal e- acceptor Form in which energy is temporarily captured Photosystems required cyclic None No None ATP PSI non-cyclic water yes NADP+ ATP & NADPH PSI & PSII Z-scheme energetics PSII Photochemistry 1) LHCII absorbs a photon 2) energy is transferred to P680 PSII Photochemistry 3) P680* reduces pheophytin ( chl a with 2 H+ instead of Mg2+) = primary electron acceptor PSII Photochemistry 3) P680* reduces pheophytin ( chl a with 2 H+ instead of Mg2+) = primary electron acceptor charge separation traps the electron PSII Photochemistry 4) pheophytin reduces PQA (plastoquinone bound to D2) moves electron away from P680+ & closer to stroma PSII Photochemistry 5) PQA reduces PQB (forms PQB- ) PSII Photochemistry 6) P680+ acquires another electron , and steps 1-4 are repeated PSII Photochemistry 7) PQA reduces PQB - -> forms PQB2- PSII Photochemistry 8) PQB2- acquires 2 H+ from stroma forms PQH2 (and adds to ∆pH) PSII Photochemistry 9) PQH2 diffuses within bilayer to cyt b6/f - is replaced within D1 by an oxidized PQ Photolysis: Making Oxygen 1) P680+ oxidizes tyrZ ( an amino acid of protein D1) Photolysis: Making Oxygen 2) tyrZ + oxidizes one of the Mn atoms in the OEC Mn cluster is an e- reservoir Photolysis: Making Oxygen 2) tyrZ + oxidizes one of the Mn atoms in the OEC Mn cluster is an e- reservoir Once 4 Mn are oxidized replace e- by stealing them from 2 H2O Shown experimentally that need 4 flashes/O2 Shown experimentally that need 4 flashes/O2 Mn cluster cycles S0 -> S4 Reset to S0 by taking 4 efrom 2 H2O Electron transport from PSII to PSI 1) PQH2 diffuses to cyt b6/f 2) PQH2 reduces cyt b6 and Fe/S, releases H+ in lumen since H+ came from stroma, transports 2 H+ across membrane (Q cycle) Electron transport from PSII to PSI 3) Fe/S reduces plastocyanin via cyt f cyt b6 reduces PQ to form PQ- Electron transport from PSII to PSI 4) repeat process, Fe/S reduces plastocyanin via cyt f cyt b6 reduces PQ- to form PQH2 Electron transport from PSII to PSI 4) PC (Cu+) diffuses to PSI, where it reduces an oxidized P700 Electron transport from PSI to Ferredoxin 1) LHCI absorbs a photon 2) P700* reduces A0 3) e- transport to ferredoxin via A1 & 3 Fe/S proteins Electron transport from Ferredoxin to NADP+ 2 Ferredoxin reduce NADP reductase Electron transport from Ferredoxin to NADP+ 2 Ferredoxin reduce NADP reductase NADP reductase reduces NADP+ Electron transport from Ferredoxin to NADP+ 2 Ferredoxin reduce NADP reductase NADP reductase reduces NADP+ this also contributes to ∆pH Overall reaction for the Z-scheme 8 photons + 2 H2O + 10 H+stroma + 2 NADP+ = 12 H+lumen + 2 NADPH + O2 Chemiosmotic ATP synthesis PMF mainly due to ∆pH is used to make ATP Chemiosmotic ATP synthesis PMF mainly due to ∆pH is used to make ATP -> very little membrane potential, due to transport of other ions Chemiosmotic ATP synthesis PMF mainly due to ∆pH is used to make ATP -> very little membrane potential, due to transport of other ions thylakoid lumen pH is < 5 cf stroma pH is 8 Chemiosmotic ATP synthesis PMF mainly due to ∆pH is used to make ATP -> very little membrane potential, due to transport of other ions thylakoid lumen pH is < 5 cf stroma pH is 8 pH is made by ETS, cyclic photophosphorylation,water splitting & NADPH synth Chemiosmotic ATP synthesis Structure of ATP synthase CF1 head: exposed to stroma CF0 base: Integral membrane protein a & b2 subunits form stator that immobilizes a & b F1 subunits a is also an H+ channel c subunits rotate as H+ pass through g & e also rotate c, g & e form a rotor Binding Change mechanism of ATP synthesis H+ translocation through ATP synthase alters affinity of active site for ATP Binding Change mechanism of ATP synthesis H+ translocation through ATP synthase alters affinity of active site for ATP ADP + Pi bind to subunit then spontaneously form ATP Binding Change mechanism of ATP synthesis H+ translocation through ATP synthase alters affinity of active site for ATP ADP + Pi bind to subunit then spontaneously form ATP ∆G for ADP + Pi = ATP is ~0 role of H+ translocation is to force enzyme to release ATP! Binding Change mechanism of ATP synthesis 1) H+ translocation alters affinity of active site for ATP 2) Each active site ratchets through 3 conformations that have different affinities for ATP, ADP & Pi due to interaction with the subunit Binding Change mechanism of ATP synthesis 1) H+ translocation alters affinity of active site for ATP 2) Each active site ratchets through 3 conformations that have different affinities for ATP, ADP & Pi 3) ATP is synthesized by rotational catalysis g subunit rotates as H+ pass through Fo, forces each active site to sequentially adopt the 3 conformations Evidence supporting chemiosmosis 1) ionophores (uncouplers) 2) can synthesize ATP if create ∆pH a) Jagendorf expt: soak cp in pH 4 in dark, make ATP when transfer to pH 8