* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Thermodynamics of Protein Folding
Protein moonlighting wikipedia , lookup
List of types of proteins wikipedia , lookup
Self-assembling peptide wikipedia , lookup
Genetic code wikipedia , lookup
Homology modeling wikipedia , lookup
Expanded genetic code wikipedia , lookup
Ribosomally synthesized and post-translationally modified peptides wikipedia , lookup
Western blot wikipedia , lookup
Peptide synthesis wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Protein (nutrient) wikipedia , lookup
Folding@home wikipedia , lookup
Cell-penetrating peptide wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Intrinsically disordered proteins wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
Bottromycin wikipedia , lookup
Protein domain wikipedia , lookup
Metalloprotein wikipedia , lookup
Protein mass spectrometry wikipedia , lookup
Biochemistry wikipedia , lookup
Protein adsorption wikipedia , lookup
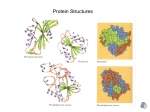
Thermodynamics of Protein Folding Introduction and Literature Review Overview • Applications of what we have learned – Intermolecular forces – Effect of acid/base chemistry – Calorimetry – Free energy of folding – Equilibrium and stability of solvation – Entropy: The hydrophobic effect Protein Folding • Activity of proteins depends on 3-D shape • Primary structure • Secondary and Tertiary structure Amino Acids • Nonpolar: vDW forces Amino Acids • Polar: Hydrogen bonding Amino Acids Acid/base: Ion/ion pH and Amino Acids Primary Structure Polar Peptide bonds Secondary Structure: H-bonds Secondary Structure: H-bonds Tertiary Structure Thermodynamics of Taq • Work from LiCata, et al. • Polymerase – E. coli – Thermus aquaticaus (Taq) • Active fragments – Klenow – Klentaq Calorimetry of Taq • Differential Scanning Calorimetry measures difference in energy needed to keep sample and reference increasing in temperature • Marks energy input into non-kinetic mode (degree of freedom) • DH = CDT Free Energy of Folding Free Energy of Folding for Taq • Experiment – pH 9.5 – Guanidinium chloride – To compare, need same conditions for both without aggregation of proteins • Taq DGunfold = 27 kcal/mol • Klenow DGunfold = 4.5 kcal/mol Structural Basis of Taq Stability • Steitz et al. suggest Taq has 4 additional internal H-bonds and 2 additional ion/ion interactions compared to Klenow • Waksman et al. suggest fewer unfavorable electrostatic charges lead to global rearrangement of electrostatic distribution and more buried nonpolar space • LiCata suggests that unfolded Taq has more surface area, leading to greater relative destabilization of unfolded relative to folded Thermodynamic Principles of Protein Folding • Very difficult to determine how all factors blend together to give overall DGfolding – Use of averages contributions, but – Each protein is unique – Large stabilization factors, large destabilization factors, but small difference between them – Use RNase T1 as a model for study (because structure is well known and many mutants have been studied) • Based on work of Pace, et al. Factors in Folding/Unfolding • Stabilizing effects – Ionization/disulfide bonds – Specific hydrogen bonding – Hydrophobic effect • Destabilizing effects – Conformational entropy – Buried polar groups Specific Hydrogen Bonding • Folding not only forms H-bonds—it also destroys them! • But which are stronger? – Transient solvent H-bonds – Specific H-bonds • Mutants show that formation of specific H-bonds stabilize protein by average of 1.6 kcal – Replacing asparagine H-bond with alanine (no Hbond) leads to destabilization of mutant enzyme – Assumptions about changed hydrophobicity, etc Specific H-Bonding Data • Quite a range of H-bond energies—valid approximation? Hydrophobic Effect • Free energy of burying nonpolar groups not primarily vDW—it is an entropic effect • Water “freezes” around nonpolar surface— clatherate shell • vDW important— cavities are destabilizing • Traditionally, thought to be actual driving force of protein folding Hydrophobic Effect: Quantitative • Free energy of transfer between water and octanol— transfer of side chain from water to model of non-polar protein core • Data suggest about 0.8 kcal stabilization for each –CH2 group buried • Mutant models show energy difference of 1.1 kcal/methylene • Suggests that burial of hydrophobic group has van der Waals contribution Conformational Entropy • Spolar and Record used calorimetry to predict an average entropy of folding of -5.6 e.u. • What does this translate to for the free energy change for freezing conformational entropy in RNase T1 (104 residues) at 25 oC? Burying Polar Groups • Water dielectric constant vs protein dielectric constant • Even if H-bonding is maintained, it is unfavorable to put polar group in nonpolar environment • Model: Partitioning of amino acid sidechains and peptide bonds between water and octanol – Determine K – Calculate DG Burying Polar Groups DG of transfer between water and octanol is thought to be best model (Transfer between water and cyclohexane also includes loss of H-bond) Summary: Contributions to RNase • Conformational entropy: calculated • Peptide buried = 73.4 peptides (1.1 kcal/peptide) • Polar buried based on previous table Summary: Contributions to RNase • Ionization and disulfide: experimental • Hydrophobic groups: from DGtr • H-bonding = 1.6 kcal (104 H-bonds) Summary: Contributions to RNase How valid are these approximations? Conclusions: Hydrophobic Effect or H-Bonding? • Pace is making the case for the importance of H-bonds vs hydrophobic effect in protein folding. How did he do? Bibliography LiCata, V.K. et al. Proteins: Struct., Funct., Bioinf. 2004, 54, 616-621. LiCata, V.K. et al. Biochem. J. 2003, 374, 785-792. Pace, C.N., et al. FASEB J. 1996, 10, 75-83. Pace, C.N. Meth. Enz. 1995, 259, 538-554.