* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Prof. Kamakaka`s Lecture 3 Notes
Survey
Document related concepts
List of types of proteins wikipedia , lookup
Protein purification wikipedia , lookup
Structural alignment wikipedia , lookup
Western blot wikipedia , lookup
Homology modeling wikipedia , lookup
Circular dichroism wikipedia , lookup
Protein domain wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Protein folding wikipedia , lookup
Intrinsically disordered proteins wikipedia , lookup
Metalloprotein wikipedia , lookup
Ribosomally synthesized and post-translationally modified peptides wikipedia , lookup
Protein mass spectrometry wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
Transcript
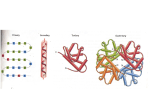
Proteins Proteins? What is its What is its How does it How does it How is its How does it How is it Where is it What are its R2 H O H2N C N H R1 O H H O H C C C OH H O R2 H H O H2N H C R1 C N C OH C H O Condensation reaction forms a peptide bond. a a Peptide bond formation The peptide bond Peptide The planar peptide bond Three bonds separate sequential a carbons in a polypeptide chain. The N—Ca and Ca—C bonds can rotate, described by dihedral angles designated f and y, respectively. The C—N peptide bond is not free to rotate. • Rotation around the peptide bond is not permitted • Rotation around bonds connected to the alpha carbon is permitted • f (phi): angle around the a-carbon—amide nitrogen bond • y (psi): angle around the a-carbon—carbonyl carbon bond • In a fully extended polypeptide, both f and y are 180° QuickTime™ and a TIFF (Uncompressed) decompressor are needed to see this picture. Steric Hindrance QuickTime™ and a TIFF (Uncompressed) decompressor are needed to see this picture. While many angles of rotation are possible, only a few are energetically favorable Ramchandran plot • Some f and y combinations are very unfavorable because of steric crowding of backbone atoms with other atoms in the backbone or side-chains • Some f and y combinations are more favorable because of chance to form favorable H-bonding interactions along the backbone • Ramachandran plot shows the distribution of f and y dihedral angles that are found in a protein • shows the common secondary structure elements • reveals regions with unusual backbone structure While many angles of rotation are possible only a few are energetically favorable QuickTime™ and a TIFF (Uncompressed) decompressor are needed to see this picture. Rotation Alpha helix • The backbone is more compact with the y dihedral (N–Ca—C–N) in the range ( 0° <y < -70°) • Helical backbone is held together by hydrogen bonds between the nearby backbone amides • Right-handed helix with 3.6 residues (5.4 Å) per turn • Peptide bonds are aligned roughly parallel with the helical axis • Side chains point out and are roughlyperpendicular with the helical axis Left and right handedness QuickTime™ and a TIFF (Uncompressed) decompressor are needed to see this picture. • Not all polypeptide sequences adopt a helical structures • Small hydrophobic residues such as Ala and Leu are strong helix formers • Pro acts as a helix breaker because the rotation around the N-Ca bond is impossible • Gly acts as a helix breaker because the tiny R group supports other conformations Peptide dipole Beta Sheet • The backbone is more extended with the y dihedral (N–Ca—C–N) in the range ( 90° < y < 180°) • The planarity of the peptide bond and tetrahedral geometry of the a-carbon create a pleated sheetlike structure • Sheet-like arrangement of backbone is held together by hydrogen bonds between the more distal backbone amides • Side chains protrude from the sheet alternating in up and down direction • Parallel or antiparallel orientation of two chains within a sheet are possible • In parallel b sheets the H-bonded strands run in the same direction • In antiparallel b sheets the H-bonded strands run in opposite directions Beta strand is an extended structure… 3.5 A between R groups in sheet compared to 1.5 in alpha helix QuickTime™ and a TIFF (Uncompressed) decompressor are needed to see this picture. Anti‐parallel B sheet R‐groups spaced at 3.5 A Distance R groups alternate above and below plane of sheet Parallel B sheet R‐groups spaced at 3.25 A distance R groups alternate above and below plane of sheet Parallel and antiparallel QuickTime™ and a TIFF (Uncompressed) decompressor are needed to see this picture. The Beta turn • b-turns occur frequently whenever strands in b sheets change the direction • The 180° turn is accomplished over four amino acids • The turn is stabilized by a hydrogen bond from a carbonyl oxygen to amide proton three residues down the sequence • Proline in position 2 or glycine in position 3 are common in b-turns The Beta turn Cis and Trans proline Tertiary Structures • Tertiary structure refers to the overall spatial arrangement of atoms in a polypeptide chain or in a protein • One can distinguish two major classes – fibrous proteins typically insoluble; made from a single secondary structure – globular proteins water-soluble globular proteins lipid-soluble membrane proteins Fibrous Proteins Keratin Hair Collagen Collagen Silk Silk Globular Proteins Myoglobin Tertiary A simple motif An elaborate motif X-ray diffraction NMR (1D) NMR (2D) Constructing large motifs Quaternary structure • Quaternary structure is formed by spontaneous assembly of individual polypeptides into a larger functional cluster • Oligomeric Subunits are arranged in Symmetric Patterns Hemoglobin Rotational symmetry Dihedral symmetry Protein Denaturation Protein Denaturation Protein Renaturation Protein folding QuickTime™ and a TIFF (Uncompressed) decompressor are needed to see this picture. Folding pathway Molten globules Chaperones