* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Document
Interactome wikipedia , lookup
Enzyme inhibitor wikipedia , lookup
Biochemical cascade wikipedia , lookup
Evolution of metal ions in biological systems wikipedia , lookup
Multi-state modeling of biomolecules wikipedia , lookup
Paracrine signalling wikipedia , lookup
Proteolysis wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Biochemistry wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Homology modeling wikipedia , lookup
Ligand binding assay wikipedia , lookup
Protein structure prediction wikipedia , lookup
Metalloprotein wikipedia , lookup
Western blot wikipedia , lookup
Signal transduction wikipedia , lookup
Clinical neurochemistry wikipedia , lookup
Drug discovery wikipedia , lookup
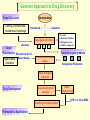
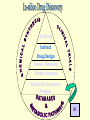
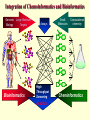
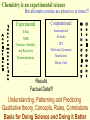
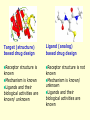
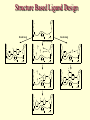
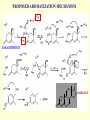
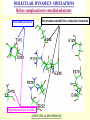
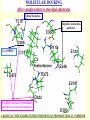
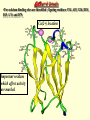
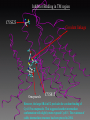
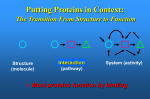
G. Narahari Sastry Molecular Modelling Group Organic Chemical Sciences Indian Institute of Chemical Technology Hyderabad – 500 007 [email protected]; [email protected] http://203.199.182.73/gnsmmg National Seminar on BioInformatics - Pondicherry Drug Discovery & Development It starts with disease identification Isolate protein involved in disease (2-5 years) Find a drug effective against disease protein (2-5 years) Scale-up Human clinical trials (2-10 years) Preclinical testing (1-3 years) Formulation FDA approval (2-3 years) Discovery and Development of Drugs Discover mechanism of action of disease Identify target protein Screen known compounds against target or Chemically develop promising leads Find 1-2 potential drugs Toxicity, pharmacology Clinical Trials Genomic Approach to Drug Discovery Target Discovery Genome data Existing Chemical and biochemical knowledge Functional & comparative Genomics Target gene annotation Literature Target Prioritization Biochemical & Cell Experimental Based Assays Validation A B C GO terms 1. Molecular Function 2. Biological process 3. Cellular component Translated gene products Functionally validated target A B C Comparative Proteomics Role of targets in disease Drug Development Sequence-structure analysis Small molecule lead HTS+/- in silico SBDD Screening and improvement Therapeutic Application Screening and Optimization Cycle with in-silico components Database clustering Target Selected Similarity analysis QSAR pharmaco Structure phore based design Assay HTS Chemistry Target developed begins structure obtained Candidate taken forward Docking Indirect Drug Design Protein Modeling Protein Analysis Nucleotide Sequence Analysis BI Virtual Screening 106 small-molecule compounds vHTS: MM + scoring functions N x 102 leads Filters: ADMET / QSAR M x 101 leads Filters: synthesis / manufacturing / IP / patent / biological assays 1 - 5 leads Integration of Chemoinformatics and Bioinformatics Genomic Biology Large Molecule Targets Bioinformatics Assays High Throughput Screening In silico Small Molecules Computational chemistry Cheminformatics Much About Structure • Structure Function • Structure Mechanism • Structure Origins/Evolution • Structure Anything!!! Quantum Mechanics “The underlying physical laws necessary for the mathematical theory of…the whole of chemistry are thus completely known, and the difficulty is only that the exact application of these laws leads to equations much too complicated to be soluble.” -P. A. M. Dirac • Exact solutions are available only for Hydrogen atom. • Modeling any realistic system needs approximations (mathematically not solvable) • Plenty of approximations were put forward to tackle mathematic complexity Chemistry is an experimental science But alternative routes are attractive at times!!! Experimental Computational X-Ray NMR Structure, Stability and Reactivity Thermochemistry … … Semiempirical Ab Initio DFT Molecular Dynamics Simulations Monte Carlo … Results Factual Data!!! Understanding, Patterning and Predicting Qualitative theory, Concepts, Rules, Correlations Basis for Doing Science and Doing it Better The Jargon of nomenclature • • • • • • • • Molecular Modeling Computational Chemistry Theoretical Chemistry Simulations Quantum Chemistry Computational Biology Molecular Dynamics Mathematical Chemistry Central Paradigm: Deriving information on molecular systems without really synthesizing them. Computational Chemistry Quantum Mechanics (QM) Hybrid QM / MM Molecular Mechanics (MM) Semi-empirical (SE) The current scenario in chemistry • Computation has become an effective alternative to explore the structural, energetic, mechanistic and other properties of small molecules (say less than 8-10 atoms). SOMETIMES THE COMPUTATIONAL ACCURACY SUPERCEDES THE EXPERIMNTAL ACCURACY Every Computational Experiment at Any Level of Theory Yields an Answer… Usually Answers for Many Questions Judging the Reliability is the Crucial Task Just Like Experiments Fail, Computations Fail The paradigm shift … However, the challenges are of different kind in modeling chemistry and biology!! It is not only the size but the philosophy!!!..!!! Biological Structure Sequence 3D structure MESDAMESETMESSRSMYN AMEISWALTERYALLKINCAL LMEWALLYIPREFERDREVIL MYSELFIMACENTERDIRATV ANDYINTENNESSEEILIKENM RANDDYNAMICSRPADNAPRI MASERADCALCYCLINNDRKI NASEMRPCALTRACTINKAR KICIPCDPKIQDENVSDETAVS WILLWINITALL Structural Scales polymerase SSBs Complexes helicase primase Organism Assemblies Cell Structures System Dynamics Cell Bottlenecks in developing Structure – Function Relationships Structures determined by NMR, computation, or X-ray crystallography are static snapshots of highly dynamic molecular systems Biological process (recognition, interaction, chemistry) require molecular motions and time dependent. To comprehend and facilitate thinking about the dynamic structure of molecules is crucial. Relevant timescales Bond vibration 10-15 femto MD step Isomeris- Water ation dynamics 10-12 pico Helix Fastest forms folders 10-9 nano long MD run Conformati onal transitions 10-6 typical folders 10-3 100 micro milli where we need to be Enzyme catalysis slow folders seconds where we’d love to be Protein folding Ligand binding How does the drug differ from an inhibitor? Selectivity Less toxicity Bioavailability Reach the target Ease of synthesis Low price Slow (or) no development of resistance Stability upon storage as tablet or solution Pharmacokinetic parameters No allergies Bioavailability (ADMET) • ADMET • Adsorption • Distribution • Metabolism • Excretion • Toxicity • Model and Predict: • Biotransformations & metabolites • Catalytic reactions • Drug-receptor interactions • GI physiology • Transepithelial transport • Epithelial permeability • Solubility • Toxicity Which Strategy? • Do you have a validated target? • Do you have active ligands? • Do you have both? Computer Aided Drug Design Science Support Drug Design Structure based Ligand based Target (structure) based drug design Ligand (analog) based drug design Receptor Receptor structure is known Mechanism is known Ligands and their biological activities are known/ unknown structure is not known Mechanism is known/ unknown Ligands and their biological activities are known Various Steps Involved • • • • • • Get the structure of the receptor Identify the active site Build a library of possible ligands Docking & Scoring Understand receptor-ligand interactions Design new ligands Structure Based Ligand Design H N O Docking Linking Building H N O O H ? ? ? H N H N ? O H O O H O O O ? H N S O ? O N H N O O H H O O H N S O H O CADD Success Stories • FKBP Ligand • docking and scoring • P. Burkhard et al., J. Mol. Biol. 287, 853-858, 1999 + • K ion channel blocker • fragment-based evolutionary design • G. Schneider et al., J. Computer-Aided Mol. Design 14, 487-494, 2000 2+ • Ca antagonist / T-channel blocker • pharmacophore similarity search • G. Schneider et al., Angew. Chem. Int. Ed. Engl. 39, 4130-4133, 2000 • Glyceraldehyde-phosphate DH inhibitors • combinatorial docking • J.C. Bressi et al., J. Med. Chem. 44, 2080-2093, 2001 • Thrombin inhibitor • docking, de-novo design • H.J. Bohm et al., J. Computer-Aided Mol. Design 13, 51-56, 1999 • HIV-1 RNA TAR inhibitor • docking, database search • A.V. Filikov et al., J. Computer-Aided Mol. Design 14, 593-610, 2000 • Aldose reductase inhibitors • 3-D database searching • Y. Iwata et al., J. Med. Chem. 44, 1718-1728, 2001 • DNA gyrase inhibitor • structure-based virtual screening • H.J. Boehm et al., J. Med. Chem. 43, 2664-2674, 2000 • Let us look at some of recent interests Broad Objectives: Aiding the experimentalists in Drug/Molecule/Reaction design We strongly believe that while chemistry and biology are experimental sciences THEORY-EXPERIMENT INTERPLAY IS INDISPENSABLE • Theoretical/computational approaches to bring insights which might trigger interest of the prospective experimental groups (Usually with no collaboration with experimentalists) • Rationalizing the experimental finding with computations and participate in the designing of experiments (In collaboration with experimentalists or groups of experimentalists) Non-availability of the receptor structure is a bottleneck… In our pursuit to engage with experimentalists for lead discovery or optimization, our efforts become restricted in the absence of an experimental structure of the receptor protein/enzyme. When we analyze, it occurred to us that most of these ‘important target receptors’ whose structures are not available belong to the class of ‘membrane proteins’. MEMBRANE PROTEINS – What are they • Membrane proteins are those that exist in cell membranes. • They can serve as structural supports, as both passive and active channels for ions and chemicals, or serve more specialized functions such as light reception. • Membrane proteins form about 25% of all protein sequences. (They constitute close to 70% of drug targets) • Only 2% of PDB structures belong to membrane proteins! Sastry et al, Computational Biology and Chemistry, 2006, in press Membrane proteins form about 25% of all protein sequences. Only 2% of PDB structures belong to this class! Membrane Proteins: Classification… • Receptors for extracellular ligands Ex :- G-Protein coupled receptors Tyrosine kinase receptors • Transport proteins Ex :- Molecular translocators Ion channels • Membrane-bound enzymes Ex :- Lipid synthases Cytochrome P-450 enzymes • Proteins associated with cytoskeletal network Ex :- Cytoskeletal attachments • Proteins associated with energy production Ex :- Photosynthetic complexes Respiratory chain complexes Challenges in computer simulations of membrane proteins. • Heavy molecular weight and size. • Their association with lipid bilayer. • Technical limitations related to the accuracy of the empirical potential function. • Difficulties with accurately incorporating important variables such as pH, transmembrane potential. • Starting configuration of a simulation may also bias the results in undesirable ways. • Comparative protein modelling approaches are very essential Sastry et al, Computational Biology and Chemistry, 2006, in press HUMAN AROMATASE: A PERIPHERAL MP PLAY A MAJOR ROLE IN STEROID AND INHIBITOR BINDING. ACIDIC RESIDUES HEME HOMOLOGY MODEL •Membrane bound microsomal cytochrome P450 enzyme. •Converts androgens to estrogens by aromatisation of A-ring of steroids. •Estrogens and their carcinogenic metabolites are responsible for progression of breast cancer WHAT IS THE ROLE OF THESE ACIDIC RESIDUES IN THE AROMATIZATION MECHANISM? Sastry et al, J. Com. Aided Mol. Design, 2006, in press Our Attempts of Modeling Aromatase • A protein model is constructed (based on CYP 2C5 (pdb code: 1NR6, sequence identity is found to be 28%) • The role of acidic residues in controlling the function(substrate binding with androstenedione, testosterone and norandrogens) is studied. • Studies help in designing putative inhibitors to control the aromatase activity. Sastry et al, J. Com. Aided Mol. Design, 2006, in press PROPOSED AROMATIZATION MECHANISM A-ring of ANDROGENS O ANDROGEN A O MOLECULAR DYNAMICS SIMULATIONS Before complexation to steroidal substrates No H-bond interaction Environment suitable for carboxylate formation High conformational flexibility ACTIVE SITE ACIDIC RESIDUES MOLECULAR DOCKING After complexation to steroidal substrates H-Bond formation Repulsive interaction predicted. CLAMPED ! Flexibility decreases. Environment suitable for carboxylate formation. A MOLECULE WHICH ARRESTS THESE PROPERTIES IS PROPOSED TO BE AN INHIBITOR Inhibition of aromatase activity by 4-hydroxy androstenedione (formestane) Critical H-bond between inhibitor and T310 hampering its’ role in the mechanism. ONE COULD DESIGN A MOLECULE BY ADDING OR DELETING A GROUP FROM ANDROGEN SKELETON TO ARREST THE PROPERTIES OBSERVED FOLLOWING COMPLEXATION. O ANDROSTENEDIONE (Substrate) A O O FORMESTANE (Inhibitor) A O OH ACTIVE SITE Human 5-lipoxygenase (5-LO)-Peripheral MP Catalytic domain Non-heme iron MODEL Ca(2+) binding Mg(2+) binding Tryptophan residues β-barrel domain •5-LO catalyses the rate limiting steps in leukotriene synthesis. •Calcium binds reversibly to 5-LO, triggering its translocation from the cytoplasm to the nuclear membrane. Sastry et al, Biophys. Biochem. Res. Comm, 2004, 320, 461-467 -barrel domain •Two calcium binding sites are identified ; ligating residues: F14, A15, G16, D18, D19, L76 and D79. Ca(2+) location Important residues which affect activity are marked. Gastric Proton Pump H(+)K(+)-ATPase – Integral MP ANTI-ULCER TARGET ATP binds here Cytoplasmic Phosphorylation. E1 E2 Inhibitor binding sites. Cation binding sites Transmembrane Lumenal •Expose ion binding sites sequentially to each side of the membrane. Sastry et al, Biophys. Biochem. Res. Comm, 2004, 319, 312-320; Biophys. Biochem. Res. Comm, 2005, 336, 961-966 Inhibitor Binding in TM region Inhibitor binding sites CYS323 Covalent linkage Omeprazole CYS815 However, the large SBA in E2 precludes the covalent binding of Cys815 to omeprazole. This suggested another intermediate conformation with slightly more exposed Cys815. The existence of stable intermediate structures has been proved in 2004. Cation binding in E1 conformation T825 Q941 D826 E822 H3O+ N794 E345 H3O+ V343 A341 V340 E797 Proposed hydronium binding. Cα – carbons of arenes in the pump. Regular disposition aids hydronium transport. Amino acid ligands (D,E,N,Q) that bind to metal ions in proteins # of Binding structures for metals PDB (June 2004) Ca2+ : 2020; Cu(II) : 298 Ni(II) : 118 Na+ : 678; Mn (II) : 454 Co(II) :101 K+ Fe (II) : 100 Fe(III) :269 : 258; Typical non-covalent binding to cations (from Mg2+ : 1167; PDB). The distances between the ligating atoms and ion vary for different cations. Asp Glu Asn Zn (II) : 1545 Gln In general, the acidic amino acid or their amides (ASP, GLU, ASN, GLN) are present in the ligating sphere of the cations (Ca, Na, K, Mg, etc.) . Additional ligating amino acid residues: Ala, Val, Thr, Leu, Phe etc. An investment in knowledge pays the best interest. Benjamin Franklin CAUTION…. •Don't be a naive user!?! •When computers are applied to biology, it is vital to understand the difference between mathematical & biological significance •computers don’t do biology, they do sums quickly macromolecular structure methods protocols Structure determinations methods Traditional Approach Rational Approach It’s like a game of LUDO Done 99 98 97 96 95 94 93 92 91 81 82 83 84 85 86 87 88 89 90 80 79 78 77 76 75 74 73 72 71 61 62 63 64 65 66 67 68 69 70 60 59 58 57 56 55 54 53 52 51 41 42 43 44 45 46 47 48 49 50 40 39 38 37 36 35 34 33 32 31 21 22 23 24 25 26 27 28 29 30 20 19 18 17 16 15 14 13 12 11 1 2 3 4 5 6 7 8 9 10 Drug Discovery “This isn’t rocket science. This is much harder.” -- Alan Holmer -- President, PhRMA GNS, Dr. G. Madhavi Sastry, Dr. Y. Soujanya, Srinivas Reddy, Punnagai, Gayatri, Srivani, Sateesh, Nagaraju, Dolly, Srinivasa Rao, Prasad, Mukesh, Murty, Usha Rani, Srinivas, Janardhan, Bharat, Upendra. Past Ph.D. students: Dr. U. Deva Priyakumar, Mr. T.C. Dinadayalane