* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Document
Biosynthesis wikipedia , lookup
Gel electrophoresis wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Evolution of metal ions in biological systems wikipedia , lookup
Oxidative phosphorylation wikipedia , lookup
Interactome wikipedia , lookup
Signal transduction wikipedia , lookup
Basal metabolic rate wikipedia , lookup
Multi-state modeling of biomolecules wikipedia , lookup
Protein purification wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Size-exclusion chromatography wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Protein structure prediction wikipedia , lookup
Western blot wikipedia , lookup
Metalloprotein wikipedia , lookup
Photosynthetic reaction centre wikipedia , lookup
1
Protein folding
Primary structure itself results in some folding constraints:
See bottom of handout 3-3
2
3
And these 4 atoms are in
one plane (N central)
These 4 redatoms
are ininone
so 6 atoms
oneplane
plane
(C of C=O central)
4
5
6
7
8
There’s still plenty of flexibility
Secondary structure: the alpha helix
9
Amino acids shown
simplified, without
side chains and H’s.
Almost every N-H and C=O
group can participate
10
Alpha helix depictions
C = grays
N = blue
O = red
Poly alanine
Side chains = -CH3 (lighter
gray)
H’s not shown
11
Linus Pauling and a model of the alpha helix.1963
Secondary structure:
H-bond
AA residue
beta pleated sheet
12
13
Beta sheet (i.e., beta pleated sheet)
antiparallel
antiparallel
parallel
14
Beta-sheets
Anti-parallel
Parallel
15
secondary structure (my definition):
structure produced by regular
repeated interactions between
atoms of the backbone.
16
Tertiary structure: The overall 3-D structure of a polypeptide.
Neither
This is a popular “ribbon” model
of protein structure. Get familiar
with it. The ribbons are stretches
of single polypeptide chains. A
single ribbon is NOT a sheet.
3 alpha helices
These “ribbon” depictions do not show the side chains, only the backbone
Tertiary structure
(overall 3-D)
17
ionic
hydrophobic
H-bond
cys
Ion - dipole
interaction
covalent
Van der Waals
Examples of bonds
determining 3D structure
Exist in loop regions and in regions of secondary structure
18
Disulfide bond formation
Disulfide bond
(covalent, strong)
Sulfhydryl group
R-CH2-SH
cysteine
+
HS-CH2-R
cysteine
½ O2
R-CH2-S-S-CH2-R + HOH
cystine
Two sulfhydryls have been oxidized (lost H’s)
Oxygen has been reduced (gained H’s).
Oxygen was the oxidizing agent (acceptor of the H’s).
An oxidation-reduction reaction: Cysteines are getting oxidized
(losing H atoms, with electron; NOT losing a proton, not like acids.)
Oxygen is getting reduced, gaining H-atoms and electrons
Actually it’s the loss and gain of the electrons that constitutes oxidation and
reduction, respectively.
No catalyst is usually needed here.
19
Overall 3-D structure of a polypeptide is tertiary structure
Stays intact in the jacuzzi at 37 deg C
Usually does not require the strong covalent disulfide bond
to maintain its 3-D structure
[Tuber mode]l
Protein structures are depicted in a variety of ways
Backbone only
Ribbon
Small molecule
bound
Drawing attention
to a few side groups
Continuous lines, ribbons=
backbone (not sheets)
Space-filing,
with surface charge
blue = +
red =
-
Space-filling
20
21
Most proteins are organized into
22
Handout 4-2
23
Two different
proteins with
almost the
same 3-D
structure !
Handout 4-2
4o,
24
QUATERNARY STRUCTURE
Monomeric protein (no quaternary structure)
Dimeric protein (a homodimer)
The usual
weak
bonds
Dimeric protein (a heterodimer)
Also called:
multimeric proteins
A heterotetramer
A heteropolymeric protein (large one)
25
Hemoglobin
$
One protein
$
Four polypeptide chains,
2 identical alphas
and 2 identical betas
Four “subunits”
Molecular weight
$
16,000
Subunit molecular weight
16,000
Subunit molecular weight
64,000
Protein molecular weight
$
$
64,000, even though the 4 chains are
not covalently bonded to each other
26
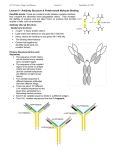
Tetramer
Two heavy chains (H),
Two light chains (L)
Interchain disulfide bonds
The 4 weak bond types
27
Sickle cell disease
Normal
glu
glu
Sickle cell
glu
glu
val
val val
val
Some small molecules can be bound tightly to a protein.
Such associated small molecule are called “prosthetic groups”.
Some are even covalently bound to the protein.
Pyridoxal
phosphate
AA
side chain
= Vitamin B6
Enzyme
28
29
Most prosthetic groups are bound tightly via weak bonds.
Tetrahydrofolic acid
~ vitamin B9
Riboflavin
~ vitamin B2
Heme
Membrane proteins
30
Hydrophobic side
chains on the protein
exterior for the
portion in contact
with the interior of the
phospholipid bilayer.
Anions are
negatively
charged.
Cations are
positively
charged
Small molecules bind with great specificity to pockets on protein surfaces
31
Too far
32
Ligand
Protein
Ligand binding can be equisitely specific:
the estrogen reeptor binds estrogen but not testosterone.
Testosterone
Estrogen
33
34
Protein separation methods
Ultracentrifugation
Mixture of proteins
Ultracentrifuge
35
Causing sedimentation:
centrifugal force =
m(omega)2r
m = mass
omega = angular velocity
r = distance from the center of rotation
Opposing sedimentation = friction = foV.
fo = frictional coefficient (shape)
V = velocity
Constant velocity is soon reached; then, no tnet force
So: centrifugal force = frictional force (balanced each other out)
And so:
m(omega)2r = foV
And: V = m(omega)2r/fo,
Or: V = [(omega)2r] x [m / fo]
V proportional to mass (MW)
V inversely proportional to fo
V inversely proportional to “non-sphericity”
(spherical shape moves fastest)
36
(“native”)37
Glass
plates
Sample loaded here
+
+
polyacrylamide
fibers
+++
+++
+
+++
+
+
+++
Winner:
Small, +++
High positive charge
+++
+
+++
Loser:
Large, +
low positive charge
Intermediate:
Large, +++
high positive charge
Intermediate:
Small, +
Low positive charge
Molecules shown after several
hours of electrophoresis
38
Upper resevoir
Cut out for contact
of buffer with gel
39
Cut out of glass plate
for contact of buffer with gel
Electrode connection
(~ 150 V)
Power supply
Tracking dyes
40
41
42
SDS PAGE = SDS polyacrylamide gel
electrophoresis
• sodium dodecyl sulfate, SDS (or SLS): CH3-(CH2)11- SO4-• CH3-CH2-CH2-CH2-CH2-CH2-CH2-CH2-CH2-CH2-CH2-CH2-SO4--
SDS
All the polypeptides are denatured and behave as random coils
All the polypeptides have the same charge per unit length
All are subject to the same electromotive force in the electric field
Separation based on the sieving effect of the polyacrylamide gel
Separation is by molecular weight only
SDS does not break covalent bonds (i.e., disulfides) (but can treat with
mercaptoethanol for that) (and perhaps boil for a bit for good measure)
43
Disulfides between 2 cysteines can be cleaved in the laboratory by reduction, i.e.,
adding 2 Hs (with their electrons) back across the disulfide bond.
One adds a reducing agent:
mercaptoethanol (HO-CH2-CH2-SH).
In the presence of this reagent, one gets exchange among the disulfides and the
sulfhydryls:
Protein-CH2-S-S-CH2-Protein + 2 HO-CH2CH2-SH --->
Protein-CH2-SH + HS-CH2-Protein + HO-CH2CH2-S-S-CH2CH2-OH
The protein's disulfide gets reduced (and the S-S bond cleaved), while the
mercaptoethanol gets oxidized, losing electrons and protons and itself forming a
disulfide bond.
44
P.A.G.E.
e.g., “p53”
Molecular weight
markers
(proteins of known
molecular weight)
45
Molecular sieve chromatography
(= gel filtration, Sephadex chromatography)
Sephadex bead
46
Molecular sieve chromatography
Sephadex bead
47
Molecular sieve chromatography
Sephadex bead
48
Molecular sieve chromatography
Sephadex bead
49
Molecular sieve chromatography
Sephadex bead
50
Fancy
Plain
4oC (cold room)
Larger molecules get to the bottom faster, and ….
Non-spherical molecules get to the bottom faster
~infrequent
orientation
Non-spherical
molecules get to
the bottom faster
51
52
Handout 4-3: protein separations
53
Winners: Largest and
most spherical
Lowest MW
Largest and
least spherical
Similar to handout 4-3,
but Winners &
native PAGE added
Winners:
Most charged
and smallest
54
Enzymes =
protein catalysts
55
Flow of glucose in E. coli
Macromolecules
Polysaccharides
Lipids
Nucleic Acids
Proteins
yn
th
e
tic
pa
t
hw
ay
monomers
bi
os
intermediates
glucose
Each arrow = an ENZYME
Each arrow = a specific chemical reaction
56
Chemical reaction between 2 reactants
H 2 + I2
2 HI
H 2 + I2
2 HI + energy
“Spontaneous” reaction:
Energy released
Goes to the right
H-I is more stable than H-H or I-I here
i.e., the H-I bond is stronger, takes more energy to break it
That’s why it “goes” to the right,
i.e., it will end up with more products than reactants
i.e., less tendency to go to the left, since the products are more stable
57
say, 100
kcal/mole
say, 103
kcal/mole
H2 + I2
2 HI
{
Change in Energy (Free Energy)
2H + 2I
Atom pulled completely apart
(a “thought” experiment)
-3 kcal/mole
Reaction goes spontaneously to the right
If energy change is negative: spontaneously to the right = exergonic: energy-releasing
If energy change is positive: spontaneously to the left = endergonic: energy-requiring
58
Different ways of writing chemical reactions
H 2 + I2
2 HI
H 2 + I2
2 HI
H 2 + I2
2 HI
H 2 + I2
2 HI
H 2 + I2
2 HI
59
say, 100
kcal/mole
But: it is not necessary to break
molecule down to its atoms in order
to rearrange them
say, 103
kcal/mole
H2 + I2
2 HI
{
Change in Energy (Free Energy)
2H + 2I
-3 kcal/mole
60
Reactions proceed through a transition state
I
I
+
H H
I
I
+
H H
I
I
H H
I
I
H
H
Transition state
(TS)
(H2 + I2)
I
H
+
I
H
(2 HI)
Products
61
Change in Energy
2H + 2I
~100 kcal/mole
H-H
| |
I-I
(TS)
Say,
~20 kcal/mole
2 HI
{
H 2 + I2
-3 kcal/mole
Activation
energy
Allows it to happen
Energy needed
to bring molecules
together to form
a TS complex
determines speed =
VELOCITY =
rate of a reaction
H 2 + I2
2 HI
{
Change in Energy (new scale)
62
HHII
(TS)
Activation
energy
3 kcal/mole
Net energy change:
Which way it will end up.
the DIRECTION
of the reaction, independent of the rate
2 separate
concepts
63
Concerns about the cell’s chemical reactions
• Direction
– We need it to go in the direction we want
• Speed
– We need it to go fast enough to have the
cell double in one generation
64
Example
Biosynthesis of a fatty acid
3 glucose’s
18-carbon fatty acid
Free energy change: ~ 300 kcal per mole of glucose used is REQUIRED
So: 3 glucose
18-carbon fatty acid
So getting a reaction to go in the direction you want is a major problem
(to be discussed next time)
65
Concerns about the cell’s chemical reactions
• Direction
– We need it to go in the direction we want
• Speed
– We need it to go fast enough to have the
cell double in one generation
– Catalysts deal with this second problem, which we will now
consider
66
The velocity problem is solved by catalysts
The catalyzed reaction
The catalyst takes part in the reaction,
but it itself emerges unchanged
67
Change in Energy
HHII
(TS)
Activation
energy
without
catalyst
TS
complex
with
catalyst
H 2 + I2
2 HI
Activation
energy
WITH the
catalyst
68
Reactants in an enzyme-catalyzed reaction = “substrates”
69
Reactants (substrates)
Active site
or
Not a substrate
substrate binding site
(not exactly synonymous,
could be just part of the active site)
70
Unlike inorganic catalysts,
enzymes are specific
Substrate Binding
71
Small molecules bind with great specificity to pockets on ENZYME surfaces
Too far
72
Unlike inorganic catalysts,
enzymes are specific
succinic dehydrogenase
HOOC-HC=CH-COOH <-------------------------------> HOOC-CH2-CH2-COOH
+2H
fumaric acid
succinic acid
NOT a substrate for the enzyme:
1-hydroxy-butenoate:
HO-CH=CH-COOH
(simple OH instead of one of the carboxyl's)
Maleic acid
73
+
Enzymes work as catalysts for two reasons:
1. They bind the substrates putting them in close proximity.
2. They participate in the reaction, weakening the covalent bonds
of a substrate by its interaction with their amino acid residue side
groups (e.g., by stretching).