* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Alignment of pairs of sequences
Polycomb Group Proteins and Cancer wikipedia , lookup
Transposable element wikipedia , lookup
Pathogenomics wikipedia , lookup
Human genome wikipedia , lookup
Metagenomics wikipedia , lookup
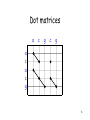
Genome evolution wikipedia , lookup
Gene expression profiling wikipedia , lookup
Copy-number variation wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Genome editing wikipedia , lookup
Protein moonlighting wikipedia , lookup
Helitron (biology) wikipedia , lookup
Multiple sequence alignment wikipedia , lookup
Point mutation wikipedia , lookup
Smith–Waterman algorithm wikipedia , lookup
Sequence Alignment I Dot Matrices Reading • Mount, Chapters 1, 2, and 3 (up to page 94) 2 Why compare sequences? • To find whether two (or more) genes or proteins are evolutionarily related to each other • To find structurally or functionally similar regions within proteins 3 Similar genes arise by gene duplication • Copy of a gene inserted next to the original • Two copies mutate independently • Each can take on separate functions • All or part can be transferred from one part of genome to another 4 Sequence Comparison Methods • Dot matrix analysis • Dynamic Programming • Word or k-tuple methods (FASTA and BLAST) 5 Dot matrices a c g c g a c a c g 6 Dot matrix comparison 7 Interpretation • Regions of similarity appear as diagonal runs of dots • Reverse diagonals (perpendicular to diagonal) indicate inversions • Reverse diagonals crossing diagonals (Xs) indicate palindromes 8 Interpretation • Can link separate diagonals to form alignment with gaps – Each a.a. or base can only be used once • Can't double back – A gap is introduced by each vertical or horizontal skip 9 Filtering • Dot matrices for long sequences can be noisy due to insignificant matches • Solution: use a window and a threshold – compare character by character within a window (have to choose window size) – require certain fraction of matches within window in order to display it with a dot 10 Dot plot comparison using windows Window size = 11 Stringency = 7 (Put a dot only if 7 out of next 11 positions are identical.) 11 Uses for dot matrices • Aligning two proteins or two nucleic acid sequences • Finding amino acid repeats within a protein by comparing a protein sequence to itself – Repeats appear as a set of diagonal runs stacked vertically and/or horizontally 12 Repeats 100 200 300 400 500 600 700 800 100 100 200 200 300 300 400 400 500 500 600 600 700 700 800 800 100 200 300 400 500 600 700 800 Human LDL receptor protein sequence (Genbank P01130) W=1 S=1 (Mount, Fig. 3.6) 13 Repeats 100 200 300 400 500 600 700 800 100 100 200 200 300 300 400 400 500 500 600 600 700 700 800 800 W = 23 S=7 (Mount, Fig. 3.6) 100 200 300 400 500 600 700 800 14 Using substitution matrices • Dots can have weights • Some matches are rewarded more than others, depending on likelihood – Use PAM or BLOSUM matrix (more on these later) • Put a dot only if a minimum total or average weight is achieved – See Mount, Fig. 3.5 15