* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download allosteric inhibition
Proteolysis wikipedia , lookup
Biochemistry wikipedia , lookup
Endocannabinoid system wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Nicotinamide adenine dinucleotide wikipedia , lookup
Oxidative phosphorylation wikipedia , lookup
Development of analogs of thalidomide wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Restriction enzyme wikipedia , lookup
Ultrasensitivity wikipedia , lookup
Metabolic network modelling wikipedia , lookup
Lipid signaling wikipedia , lookup
Metalloprotein wikipedia , lookup
Evolution of metal ions in biological systems wikipedia , lookup
Biosynthesis wikipedia , lookup
Catalytic triad wikipedia , lookup
NADH:ubiquinone oxidoreductase (H+-translocating) wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Discovery and development of neuraminidase inhibitors wikipedia , lookup
ENZYME INHIBITION
• Studies on Inhibitors are useful for:
• Mechanistic studies to learn about how enzymes interact
with their substrates.
• Role of inhibitors in enzyme regulation.
• Drugs if they inhibit abberrant biochemical reactions:
– penicillin, ampicillin, et al.: interfere with the synthesis
of bacterial cell walls.
– methotrexate: anti-cancer drug that affects DNA
metabolism in actively growing cells
• Understanding the role of biological toxins.
– Arsenate: mimics phosphate esters in enzyme reactions,
but are easily hydrolysed.
– Amino acid analogs: useful herbicides (i.e. roundup)
– Insecticides: chemicals targeted for the insect nervous
system.
TYPES OF INHIBITION
• REVERSIBLE INHIBITION
They are of three types
1. Competitive Inhibition
2. Non Competitive Inhibition and
3. Un Competitive Inhibition
• IRREVERSIBLE INHIBITION
• ALLOSTERIC INHIBITION
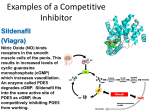
1.COMPETITIVE INHIBITION
•
Inhibitor binds to the same site on the enzyme as the
substrate, the active site.
Inhibitor only binds to the free enzyme.
Inhibitor is usually structurally very similar to the
substrate. Examples:
•
•
–
–
•
Succinate / Malonate
ATP/AMP
The reaction scheme that corresponds to competitive
inhibition is:
• The inhibitor reduces the amount of E available for
productive catalysis by the formation of the EI
complex. The inhibitor does not affect the ES complex
after it has formed. The dissociation constant for the
inhibitor is KI = [E][I]/[EI].
• There are two anticipated consequences of this
additional competitive equilibrium:
• Vmax is unchanged: At high levels of substrate all of
the inhibitor is displaced by substrate.
• KM is increased: Higher substrate concentrations are
required to reach the maximal velocity.
2.NON COMPETITIVE INHIBITION
• Noncompetitive Inhibition:
• In this case the inhibitor binds to both E and ES. Both
the slope (KM/Vmax), and the Y-intercept (1/Vmax)
of the Lineweaver-Burk plot increase (see figure 5.11).
The KI ('s) are determined as above by replotting the
slope and intercept values vs. [I].
• Vmax is decreased: At high levels of substrate the
inhibitor is still bound.
• KM is increased: Higher [S] is required to reach the
lower maximal velocity.
3.UN COMPETITIVE INHIBITION
•
•
•
•
•
The inhibitor binds directly to the ES complex.
The inhibitor does not have to bind at the active site.
The inhibitor does not have to resemble the substrate
(e.g. an allosteric inhibitor).
Vmax is reduced
The slopes of the Lineweaver-Burk plot (KM/Vmax)
are unchanged, but the Y-intercept increases by a
factor of (1 + [I]/KI). The X-intercept shifts to the
left by a factor (1 + [I]/KI).
ALLOSTERIC INHIBITION
• This type of inhibition is exclusive for allosteric
enzymes.
• Allosteric enzymes differs in their structure from
other enzymes by possesing another site apart
from active site. This site is called allosteric site
where modulators can bind
• Modulators may be of two types,
• Positive Modulator and
• Negative Modulator
Contd..
• The rate determining pathways in most of the
metabolic reactions are catalysed by Allosteric
enzymes
• The significance of this specialty is to regulate
the metabolic pathway.
• Allosteric enzymes do not follow MM-Kinetics