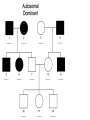
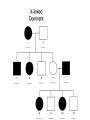
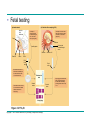
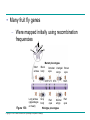
* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download c - Hialeah Senior High School
Survey
Document related concepts
Adenosine triphosphate wikipedia , lookup
Biosynthesis wikipedia , lookup
Fatty acid metabolism wikipedia , lookup
Biochemical cascade wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Photosynthetic reaction centre wikipedia , lookup
Photosynthesis wikipedia , lookup
Polyclonal B cell response wikipedia , lookup
Oxidative phosphorylation wikipedia , lookup
Signal transduction wikipedia , lookup
Citric acid cycle wikipedia , lookup
Evolution of metal ions in biological systems wikipedia , lookup
Transcript
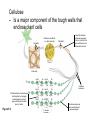
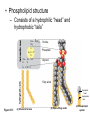
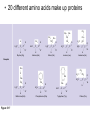
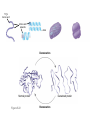
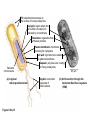
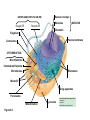
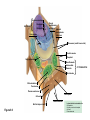
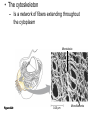
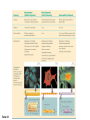
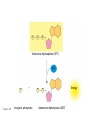
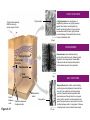
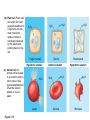
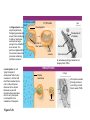
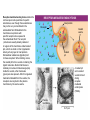
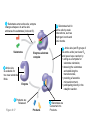
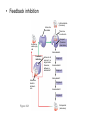
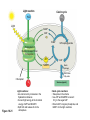
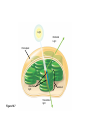
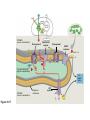
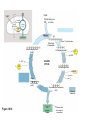
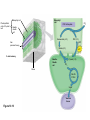
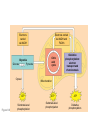
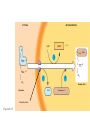
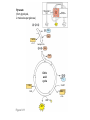
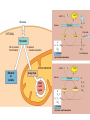
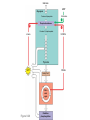
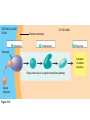
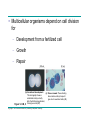
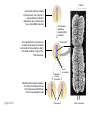
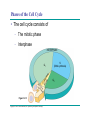
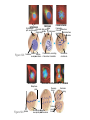
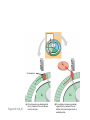
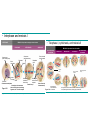
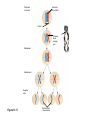
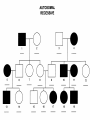
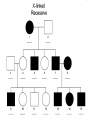
AP REVIEW 1 Hydrogen 1H Atomic mass First shell 2 He 4.00 Atomic number Helium 2He Element symbol Electron-shell diagram Lithium 3Li Beryllium 4Be Boron 3B Carbon 6C Nitrogen 7N Oxygen Fluorine 8O 9F Neon 10Ne Second shell Sodium Magnesium Aluminum Silicon Phosphorus Sulfur 13Al 16S 11Na 12Mg 14Si 15P Third shell Figure 2.8 Chlorine 17Cl Argon 18Ar Because oxygen (O) is more electronegative than hydrogen (H), shared electrons are pulled more toward oxygen. d– This results in a partial negative charge on the oxygen and a partial positive charge on the hydrogens. O d+ H H H2O d+ Carbon Nitrogen Hydrogen Sulfur Oxygen Natural endorphin Morphine (a) Structures of endorphin and morphine. The boxed portion of the endorphin molecule (left) binds to receptor molecules on target cells in the brain. The boxed portion of the morphine molecule is a close match. Natural endorphin Brain cell Morphine Endorphin receptors (b) Binding to endorphin receptors. Endorphin receptors on the surface of a brain cell recognize and can bind to both endorphin and morphine. Properties of water What is electronegativity and how does it affect interactions between water molecules? Cohesion Adhesion Surface Tension Describe how properties of water contribute to the upward movement of water in a tree. Turnover in lakes? • Lakes – Are sensitive to seasonal temperature change – Experience seasonal turnover Lake depth (m) In winter, the coldest water in the lake (0°C) lies just below the surface ice; water is progressively warmer at deeper levels of the lake, typically 4–5°C at the bottom. O2 (mg/L) 0 4 Spring Winter 8 12 8 16 2 4 4 4 4C 24 O2 concentration 0 Lake depth (m) 1 2 In spring, as the sun melts the ice, the surface water warms to 4°C and sinks below the cooler layers immediately below, eliminating the thermal stratification. Spring winds mix the water to great depth, bringing oxygen (O2) to the bottom waters (see graphs) and nutrients to the surface. O2 (mg/L) 0 4 8 12 8 16 4 4 4 4 4 4C 24 High Medium O2 (mg/L) 0 4 8 12 8 16 24 Figure 50.13 4 Autumn 4 4 4 4C 4 In autumn, as surface water cools rapidly, it sinks below the underlying layers, remixing the water until the surface begins to freeze and the winter temperature profile is reestablished. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings 4 Thermocline 3 22 20 18 8 6 5 4C Summer Lake depth (m) Lake depth (m) Low O2 (mg/L) 0 4 8 12 8 16 24 In summer, the lake regains a distinctive thermal profile, with warm surface water separated from cold bottom water by a narrow vertical zone of rapid temperature change, called a thermocline. The four emergent properties of water that are important for life are: a) Cohesion, expansion upon freezing, high heat of evaporation, and capillarity b) Cohesion, moderation of temperature, expansion upon freezing, and solvent properties c) Moderation of temperature, solvent properties, high surface tension, and capillarity d) Heat of vaporization, high specific heat, high surface tension, and capillarity e) Polarity, hydrogen bonding, high specific heat, and high surface tension Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • Ascent of xylem sap Xylem sap Outside air Y = –100.0 MPa Leaf Y (air spaces) = –7.0 MPa Transpiration Leaf Y (cell walls) = –1.0 MPa Trunk xylem Y = – 0.8 MPa Atmosphere Xylem cells Water potential gradient Y Mesophyll cells Stoma Water molecule Cohesion and adhesion in the xylem Adhesion Cell wall Cohesion, by hydrogen bonding Water molecule Root xylem Y = – 0.6 MPa Root hair Soil Y = – 0.3 MPa Soil particle Figure 36.13 Water uptake from soil Water pH Scale 0 Increasingly Acidic [H+] > [OH–] 1 2 Digestive (stomach) juice, lemon juice 3 Vinegar, beer, wine, cola 4 Tomato juice 5 Black coffee Rainwater Urine 6 Neutral [H+] = [OH–] Battery acid 7 Pure water Human blood 8 Increasingly Basic [H+] < [OH–] Seawater 9 10 11 Milk of magnesia Household ammonia 12 Household bleach 13 Oven cleaner Figure 3.8 14 Name and Comments Molecular Structural Formula Formula H (a) Methane CH4 H C H H (b) Ethane H H C2H H C C H 6 (c) Ethene Figure 4.3 A-C (ethylene) H H H C2H4 H C C H H Ball-andStick Model SpaceFilling Model H (a) Structural isomers H H H H H H C C C C C H H H H H X H (c) Enantiomers C C C H H H X C C C X H H CO2H CO2H C C H H NH2 NH2 CH3 Figure 4.7 A-C H C H H H C H X C (b) Geometric isomers H H H H CH3 H FUNCTIONAL GROUP HYDROXYL CARBONYL CARBOXYL O OH (may be written HO C C OH ) STRUCTURE In a hydroxyl group (—OH), a hydrogen atom is bonded to an oxygen atom, which in turn is bonded to the carbon skeleton of the organic molecule. (Do not confuse this functional group with the hydroxide ion, OH–.) Figure 4.10 O The carbonyl group ( CO) consists of a carbon atom joined to an oxygen atom by a double bond. When an oxygen atom is doublebonded to a carbon atom that is also bonded to a hydroxyl group, the entire assembly of atoms is called a carboxyl group (— COOH). AMINO SULFHYDRYL H N H Figure 4.10 O SH (may be written HS The amino group (—NH2) consists of a nitrogen atom bonded to two hydrogen atoms and to the carbon skeleton. PHOSPHATE ) O P OH OH The sulfhydryl group consists of a sulfur atom bonded to an atom of hydrogen; resembles a hydroxyl group in shape. In a phosphate group, a phosphorus atom is bonded to four oxygen atoms; one oxygen is bonded to the carbon skeleton; two oxygens carry negative charges; abbreviated P . The phosphate group (—OPO32–) is an ionized form of a phosphoric acid group (— OPO3H2; note the two hydrogens). The Synthesis and Breakdown of Polymers HO 1 3 2 H Unlinked monomer Short polymer Dehydration removes a water molecule, forming a new bond HO Figure 5.2A 1 2 H HO 3 H2O 4 H Longer polymer (a) Dehydration reaction in the synthesis of a polymer • Polymers can disassemble by – Hydrolysis HO 1 2 3 4 Hydrolysis adds a water molecule, breaking a bond HO 1 2 3 H Figure 5.2B (b) Hydrolysis of a polymer H H2O HO H Macromolecules • • • • Carbohydrates Proteins Lipids Nucleic acids Starch – Is the major storage form of glucose in plants Chloroplast Starch 1 m Amylose Amylopectin Figure 5.6 (a) Starch: a plant polysaccharide • Glycogen – Consists of glucose monomers – Is the major storage form of glucose in animals Mitochondria Giycogen granules 0.5 m Glycogen Figure 5.6 (b) Glycogen: an animal polysaccharide Cellulose – Is a major component of the tough walls that enclose plant cells Cell walls Cellulose microfibrils in a plant cell wall Microfibril About 80 cellulose molecules associate to form a microfibril, the main architectural unit of the plant cell wall. 0.5 m Plant cells Parallel cellulose molecules are held together by hydrogen bonds between hydroxyl groups attached to carbon atoms 3 and 6. Figure 5.8 OH CH2OH OH CH2OH O O O O OH OH OH OH O O O O O O CH OH OH CH2OH 2 H CH2OH OH CH2OH OH O O O O OH OH OH OH O O O O O O CH OH OH CH2OH 2 H CH2OH OH OH CH2OH O O O O OH OH OH O O OH O O O O CH OH OH CH2OH 2 H Glucose monomer Cellulose molecules A cellulose molecule is an unbranched glucose polymer. Fats • Fats – Are constructed from two types of smaller molecules, a single glycerol and usually three fatty acids H H C O C OH HO H C OH H C OH H C H H C H H C H H C H H C H H C H H C H H C H H C H H C H H C H H C H H C H H C H Fatty acid (palmitic acid) H Glycerol (a) Dehydration reaction in the synthesis of a fat Ester linkage O H H C O C H C H O H C O C O H C H Figure 5.11 O C H C H H C H C H H H C H C H H H C H H C H H C H H C H C H H H C H H C H H C H H C H H C H H C H H C H H C H H C H H C H (b) Fat molecule (triacylglycerol) H C H H C H H C H H C H H C H H C H H C H H C H H C H H C H H C H H C H H C H H H C C H H H C H H C H H C H H C H H C H H C H H C H H H C H H H C H H H C H H • Phospholipid structure – Consists of a hydrophilic “head” and hydrophobic “tails” CH2 + N(CH ) 3 3 Choline CH2 O O P O– Phosphate O CH2 CH O O C O C CH2 Glycerol O Fatty acids Hydrophilic head Hydrophobic tails Figure 5.13 (a) Structural formula (b) Space-filling model (c) Phospholipid symbol • One steroid, cholesterol – Is found in cell membranes – Is a precursor for some hormones H3C CH3 CH3 Figure 5.15 HO CH3 CH3 Table 5.1 • 20 different amino acids make up proteins CH3 CH3 H H3N+ C CH3 O H3N+ C H Glycine (Gly) O– C H3N+ C H Alanine (Ala) O– CH CH3 CH3 O C CH2 CH2 O H3N+ C H Valine (Val) CH3 CH3 O– C O H3N+ C H Leucine (Leu) H3C O– CH C O C O– H Isoleucine (Ile) Nonpolar CH3 CH2 S NH CH2 CH2 H3N+ C H H3N+ C O– Methionine (Met) Figure 5.17 CH2 O C H CH2 O H3N+ C C O– Phenylalanine (Phe) H O H2C CH2 H2N C O C O– H C O– Tryptophan (Trp) Proline (Pro) Amino Acid Polymers • Amino acids – Are linked by peptide bonds OH Peptide bond OH CH2 CH2 H N H SH CH2 H C C H N C C OH H N C H O H O H (a) C OH O DESMOSOMES H2O OH DESMOSOMES DESMOSOMES SH OH Peptide CH2 bond CH2 CH2 H H N C C H O Figure 5.18 (b) Amino end (N-terminus) H H N C C H O N C C OH H O Carboxyl end (C-terminus) Side chains Backbone +H 3N Amino end Amino acid subunits helix Denaturation Normal protein Figure 5.22 Denatured protein Renaturation The Structure of Nucleic Acids 5’ end 5’C O 3’C O O 5’C O 3’C OH 3’ end Figure 5.26 (a) Polynucleotide, or nucleic acid Pili: attachment structures on the surface of some prokaryotes Nucleoid: region where the cell’s DNA is located (not enclosed by a membrane) Ribosomes: organelles that synthesize proteins Bacterial chromosome (a) A typical rod-shaped bacterium Figure 6.6 A, B Plasma membrane: membrane enclosing the cytoplasm Cell wall: rigid structure outside the plasma membrane Capsule: jelly-like outer coating of many prokaryotes 0.5 µm Flagella: locomotion organelles of some bacteria (b) A thin section through the bacterium Bacillus coagulans (TEM) ENDOPLASMIC RETICULUM (ER) Rough ER Smooth ER Nuclear envelope Nucleolus NUCLEUS Chromatin Flagelium Plasma membrane Centrosome CYTOSKELETON Microfilaments Intermediate filaments Ribosomes Microtubules Microvilli Golgi apparatus Peroxisome Mitochondrion Figure 6.9 Lysosome In animal cells but not plant cells: Lysosomes Centrioles Flagella (in some plant sperm) Nuclear envelope Nucleolus Chromatin NUCLEUS Centrosome Rough endoplasmic reticulum Smooth endoplasmic reticulum Ribosomes (small brwon dots) Central vacuole Tonoplast Golgi apparatus Microfilaments Intermediate filaments CYTOSKELETON Microtubules Mitochondrion Peroxisome Plasma membrane Chloroplast Cell wall Plasmodesmata Wall of adjacent cell Figure 6.9 In plant cells but not animal cells: Chloroplasts Central vacuole and tonoplast Cell wall Plasmodesmata • The cytoskeleton – Is a network of fibers extending throughout the cytoplasm Microtubule Figure 6.20 0.25 µm Microfilaments Table 6.1 P P P Adenosine triphosphate (ATP) H2O P Figure 8.9 i + Inorganic phosphate P P Adenosine diphosphate (ADP) Energy Free energy Products Amount of energy released (∆G>0) Energy Reactants Progress of the reaction Figure 8.6 (b) Endergonic reaction: energy required Free energy Reactants Amount of energy released (∆G <0) Energy Products Progress of the reaction Figure 8.6 (a) Exergonic reaction: energy released TIGHT JUNCTIONS Tight junction Tight junctions prevent fluid from moving across a layer of cells At tight junctions, the membranes of neighboring cells are very tightly pressed against each other, bound together by specific proteins (purple). Forming continuous seals around the cells, tight junctions prevent leakage of extracellular fluid across A layer of epithelial cells. 0.5 µm DESMOSOMES Desmosomes (also called anchoring junctions) function like rivets, fastening cells Together into strong sheets. Intermediate Filaments made of sturdy keratin proteins Anchor desmosomes in the cytoplasm. Tight junctions Intermediate filaments Desmosome Gap junctions Space between cells Figure 6.31 1 µm GAP JUNCTIONS Gap junctions (also called communicating junctions) provide cytoplasmic channels from one cell to an adjacent cell. Gap junctions consist of special membrane proteins that surround a pore through which ions, sugars, amino acids, and other small molecules may pass. Gap junctions are necessary for communication between cells in many types of tissues, including heart muscle and animal embryos. Extracellular matrix Gap junction Plasma membranes of adjacent cells 0.1 µm Glycoprotein Carbohydrate Glycolipid EXTRACELLULAR SIDE OF MEMBRANE Microfilaments of cytoskeleton Figure 7.7 Cholesterol Peripheral protein Integral protein CYTOPLASMIC SIDE OF MEMBRANE (a) Diffusion of one solute. The membrane has pores large enough for molecules of dye to pass through. Random movement of dye molecules will cause some to pass through the pores; this will happen more often on the side with more molecules. The dye diffuses from where it is more concentrated to where it is less concentrated (called diffusing down a concentration gradient). This leads to a dynamic equilibrium: The solute molecules continue to cross the membrane, but at equal rates in both directions. Figure 7.11 A Molecules of dye Membrane (cross section) Net diffusion Net diffusion Equilibrium (b) Plant cell. Plant cells are turgid (firm) and generally healthiest in a hypotonic environment, where the uptake of water is eventually balanced by the elastic wall pushing back on the cell. Turgid (normal) Hypotonic solution (a) Animal cell. An animal cell fares best in an isotonic environment unless it has special adaptations to offset the osmotic uptake or loss of water. H2O Lysed Figure 7.13 H2O H2O H2O Flaccid H2O Plasmolyzed Isotonic solution Hypertonic solution H2O H2O Normal H2O Shriveled Passive transport. Substances diffuse spontaneously down their concentration gradients, crossing a membrane with no expenditure of energy by the cell. The rate of diffusion can be greatly increased by transport proteins in the membrane. Active transport. Some transport proteins act as pumps, moving substances across a membrane against their concentration gradients. Energy for this work is usually supplied by ATP. ATP Diffusion. Hydrophobic Facilitated diffusion. Many hydrophilic molecules and (at a slow substances diffuse through membranes rate) very small uncharged with the assistance of transport proteins, polar molecules can diffuse through either channel or carrier proteins. the lipid bilayer. EXTRACELLULAR 1 µm CYTOPLASM FLUID In phagocytosis, a cell Pseudopodium PHAGOCYTOSIS engulfs a particle by Wrapping pseudopodia Pseudopodium around it and packaging of amoeba it within a membraneenclosed sac large “Food” or enough to be classified other particle Bacterium as a vacuole. The particle is digested after Food the vacuole fuses with a Food vacuole vacuole lysosome containing hydrolytic enzymes. An amoeba engulfing a bacterium via phagocytosis (TEM). In pinocytosis, the cell “gulps” droplets of extracellular fluid into tiny vesicles. It is not the fluid itself that is needed by the cell, but the molecules dissolved in the droplet. Because any and all included solutes are taken into the cell, pinocytosis is nonspecific in the substances it transports. Figure 7.20 PINOCYTOSIS 0.5 µm Plasma membrane Pinocytosis vesicles forming (arrows) in a cell lining a small blood vessel (TEM). Vesicle Receptor-mediated endocytosis enables the cell to acquire bulk quantities of specific substances, even though those substances may not be very concentrated in the extracellular fluid. Embedded in the membrane are proteins with specific receptor sites exposed to the extracellular fluid. The receptor proteins are usually already clustered in regions of the membrane called coated pits, which are lined on their cytoplasmic side by a fuzzy layer of coat proteins. Extracellular substances (ligands) bind to these receptors. When binding occurs, the coated pit forms a vesicle containing the ligand molecules. Notice that there are relatively more bound molecules (purple) inside the vesicle, other molecules (green) are also present. After this ingested material is liberated from the vesicle, the receptors are recycled to the plasma membrane by the same vesicle. RECEPTOR-MEDIATED ENDOCYTOSIS Coat protein Receptor Coated vesicle Ligand Coated pit A coated pit and a coated vesicle formed during receptormediated endocytosis (TEMs). Coat protein Plasma membrane 0.25 µm • Enzymes – Are a type of protein that acts as a catalyst, speeding up chemical reactions 1 Active site is available for a molecule of substrate, the reactant on which the enzyme acts. Substrate (sucrose) 2 Substrate binds to enzyme. Glucose OH Enzyme (sucrase) H2O Fructose H O 4 Products are released. Figure 5.16 3 Substrate is converted to products. 1 Substrates enter active site; enzyme changes shape so its active site embraces the substrates (induced fit). Substrates Enzyme-substrate complex 6 Active site Is available for two new substrate Mole. Enzyme 5 Products are Released. Figure 8.17 Products 2 Substrates held in active site by weak interactions, such as hydrogen bonds and ionic bonds. 3 Active site (and R groups of its amino acids) can lower EA and speed up a reaction by • acting as a template for substrate orientation, • stressing the substrates and stabilizing the transition state, • providing a favorable microenvironment, • participating directly in the catalytic reaction. 4 Substrates are Converted into Products. Course of reaction without enzyme EA without enzyme EA with enzyme is lower Free energy Reactants ∆G is unaffected by enzyme Course of reaction with enzyme Products Progress of the reaction Figure 8.15 Effects of Temperature and pH • Each enzyme – Has an optimal temperature in which it can function Optimal temperature for typical human enzyme Optimal temperature for enzyme of thermophilic Rate of reaction (heat-tolerant) bacteria 0 20 40 Temperature (Cº) (a) Optimal temperature for two enzymes Figure 8.18 80 100 A substrate can bind normally to the active site of an enzyme. Substrate Active site Enzyme (a) Normal binding A competitive inhibitor mimics the substrate, competing for the active site. Figure 8.19 (b) Competitive inhibition Competitive inhibitor • Feedback inhibition Active site available Initial substrate (threonine) Threonine in active site Enzyme 1 (threonine deaminase) Isoleucine used up by cell Intermediate A Feedback inhibition Active site of enzyme 1 no longer binds threonine; pathway is switched off Enzyme 2 Intermediate B Enzyme 3 Intermediate C Isoleucine binds to allosteric site Enzyme 4 Intermediate D Enzyme 5 Figure 8.21 End product (isoleucine) Light reaction Calvin cycle H2O CO2 Light NADP+ ADP +P1 RuBP 3-Phosphoglycerate Photosystem II Electron transport chain Photosystem I ATP NADPH G3P Starch (storage) Amino acids Fatty acids Chloroplast O2 Figure 10.21 Light reactions: • Are carried out by molecules in the thylakoid membranes • Convert light energy to the chemical energy of ATP and NADPH • Split H2O and release O2 to the atmosphere Sucrose (export) Calvin cycle reactions: • Take place in the stroma • Use ATP and NADPH to convert CO2 to the sugar G3P • Return ADP, inorganic phosphate, and NADP+ to the light reactions Light Reflected Light Chloroplast Absorbed light Granum Transmitted light Figure 10.7 H2O CO2 LIGHT NADP+ ADP CALVIN CYCLE LIGHT REACTOR ATP NADPH STROMA (Low H+ concentration) O2 [CH2O] (sugar) Photosystem II Cytochrome complex Photosystem I NADP+ reductase Light 2 H+ Fd 3 NADP+ + 2H+ NADPH + H+ Pq Pc 2 H2O THYLAKOID SPACE (High H+ concentration) 1⁄ 2 1 O2 +2 H+ 2 H+ To Calvin cycle STROMA (Low H+ concentration) Thylakoid membrane ATP synthase ADP ATP P Figure 10.17 H+ H2 O CO2 Input Light (Entering one 3 CO2 at a time) NADP+ ADP CALVIN CYCLE LIGHT REACTION ATP NADPH Rubisco O2 [CH2O] (sugar) 3 P P Phase 1: Carbon fixation Short-lived intermediate 3 P P 6 P Ribulose bisphosphate (RuBP) 3-Phosphoglycerate 6 ATP 6 ADP CALVIN CYCLE 3 ADP 3 6 P ATP P 1,3-Bisphoglycerate 6 NADPH 6 NADPH+ 6 P Phase 3: Regeneration of P 5 the CO(G3P) 2 acceptor (RuBP) 6 P Glyceraldehyde-3-phosphate (G3P) P 1 Figure 10.18 G3P (a sugar) Output Glucose and other organic compounds Phase 2: Reduction Mesophyll cell Mesophyll cell Photosynthetic cells of C4 plant leaf CO CO 2 2 PEP carboxylase Bundlesheath cell PEP (3 C) ADP Oxaloacetate (4 C) Vein (vascular tissue) Malate (4 C) ATP C4 leaf anatomy BundleSheath cell Pyruate (3 C) CO2 Stoma CALVIN CYCLE Sugar Vascular tissue Figure 10.19 Pineapple Sugarcane C4 Mesophyll Cell Organic acid Bundlesheath cell (a) Spatial separation of steps. In C4 plants, carbon fixation and the Calvin cycle occur in different types of cells. CALVIN CYCLE Sugar CAM CO2 1 CO2 incorporated into four-carbon organic acids (carbon fixation) 2 Organic acids release CO2 to Calvin cycle CO2 Organic acid Night Day CALVIN CYCLE Sugar (b) Temporal separation of steps. In CAM plants, carbon fixation and the Calvin cycle occur in the same cells at different times. Electrons carried via NADH and FADH2 Electrons carried via NADH Glycolsis Pyruvate Glucose Cytosol ATP Figure 9.6 Substrate-level phosphorylation Citric acid cycle Oxidative phosphorylation: electron transport and chemiosmosis Mitochondrion ATP Substrate-level phosphorylation ATP Oxidative phosphorylation CYTOSOL MITOCHONDRION NAD+ NADH + H+ O– S CoA C O 2 C C O O 1 3 CH3 Pyruvate Transport protein Figure 9.10 CH3 Acetyle CoA CO2 Coenzyme A Pyruvate (from glycolysis, 2 molecules per glucose) Glycolysis Citric acid cycle ATP ATP Oxidative phosphorylation ATP CO2 CoA NADH + 3 H+ Acetyle CoA CoA CoA Citric acid cycle 2 CO2 3 NAD+ FADH2 FAD 3 NADH + 3 H+ ADP + P i ATP Figure 9.11 P1 2 ADP + 2 2 ATP Glucose Glucose Glycolysis O– C O C O CH3 2 Pyruvate CYTOSOL Pyruvate No O2 present Fermentation 2 NADH 2 NAD+ H O2 present Cellular respiration H 2 CO2 H C C OH CH3 O CH3 2 Ethanol 2 Acetaldehyde (a) Alcohol fermentation MITOCHONDRION Ethanol or lactate 2 ADP + 2 P1 2 ATP Acetyl CoA Glucose Citric acid cycle 2 NAD+ O H O– Glycolysis C O C OH CH3 2 Lactate (b) Lactic acid fermentation 2 NADH C O C O CH3 Glucose AMP Glycolysis Fructose-6-phosphate Stimulates + – Phosphofructokinase – Fructose-1,6-bisphosphate Inhibits Inhibits Pyruvate Citrate ATP Acetyl CoA Citric acid cycle Figure 9.20 Oxidative phosphorylation EXTRACELLULAR FLUID 1 Reception CYTOPLASM Plasma membrane 2 Transduction 3 Response Receptor Activation of cellular response Relay molecules in a signal transduction pathway Signal molecule Figure 11.5 Signal-binding site Segment that interacts with G proteins G-protein-linked Receptor Plasma Membrane Activated Receptor Inctivate enzyme Signal molecule GDP CYTOPLASM G-protein (inactive) Enzyme GDP GTP Activated enzyme GTP GDP Pi Cellular response • Multicellular organisms depend on cell division for – Development from a fertilized cell – Growth – Repair 200 µm 20 µm (b) Growth and development. (c) Tissue renewal. These dividing This micrograph shows a bone marrow cells (arrow) will sand dollar embryo shortly give rise to new blood cells (LM). after the fertilized egg divided, Figure 12.2 B, C forming two cells (LM). Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings 0.5 µm A eukaryotic cell has multiple chromosomes, one of which is represented here. Before duplication, each chromosome has a single DNA molecule. Chromosome duplication (including DNA synthesis) Once duplicated, a chromosome consists of two sister chromatids connected at the centromere. Each chromatid contains a copy of the DNA molecule. Centromere Separation of sister chromatids Sister chromatids Mechanical processes separate the sister chromatids into two chromosomes and distribute them to two daughter cells. Figure 12.4 Centromeres Sister chromatids Phases of the Cell Cycle • The cell cycle consists of – The mitotic phase – Interphase INTERPHASE C M yto ito ki si ne s si s G1 MI (M TOT ) P IC HA SE Figure 12.5 Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings S (DNA synthesis) G2 G2 OF PROPHASE INTERPHASE Centrosomes Aster Early mitotic Chromatin (with centriole pairs) Centromere spindle (duplicated) Figure 12.6 Nucleolus Nuclear Plasma envelope membrane METAPHASE Metaphase plate Figure 12.6 Spindle PROMETAPHASE Fragments Kinetochore of nuclear envelope Nonkinetochore microtubules Chromosome, consisting of two sister chromatids Kinetochore microtubule ANAPHASETELOPHASE AND CYTOKINESIS Daughter Centrosome at chromosomes one spindle pole Cleavage furrow Nuclear envelope forming Nucleolus forming G0 G1 checkpoint G1 Figure 12.15 A, B (a) If a cell receives a go-ahead signal at the G1 checkpoint, the cell continues on in the cell cycle. G1 (b) If a cell does not receive a go-ahead signal at the G1checkpoint, the cell exits the cell cycle and goes into G0, a nondividing state. Pair of homologous chromosomes karyotype Centromere Sister chromatids Figure 13.3 5 µm • Interphase and meiosis I • Telophase I, cytokinesis, and meiosis II MEIOSIS I: Separates homologous chromosomes INTERPHASE PROPHASE I METAPHASE I ANAPHASE I MEIOSIS II: Separates sister chromatids TELOPHASE I AND CYTOKINESIS Sister chromatids Chiasmata Spindle METAPHASE II Homologous Microtubule chromosomes attached to separate kinetochore Pairs of homologous Chromosomes duplicate Tertads line up chromosomes split up Homologous chromosomes (red and blue) pair and exchange Figure 13.8 Figure 13.8 segments; 2n = 6 in this example Chromatin TELOPHASE II AND CYTOKINESIS Metaphase plate Cleavage furrow Nuclear envelope ANAPHASE II Sister chromatids remain attached Centromere (with kinetochore) Centrosomes (with centriole pairs) PROPHASE II Sister chromatids separate Haploid daughter cells forming Tetrad Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Two haploid cells form; chromosomes are still double During another round of cell division, the sister chromatids finally separate; four haploid daughter cells result, containing single chromosomes Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Prophase I of meiosis Nonsister chromatids Tetrad Chiasma, site of crossing over Metaphase I Metaphase II Daughter cells Figure 13.11 Recombinant chromosomes • Mendel’s law of segregation, probability and the Punnett square Each true-breeding plant of the parental generation has identical alleles, PP or pp. Gametes (circles) each contain only one allele for the flower-color gene. In this case, every gamete produced by one parent has the same allele. P Generation Gametes: p P Union of the parental gametes produces F1 hybrids having a Pp combination. Because the purpleflower allele is dominant, all these hybrids have purple flowers. F1 Generation When the hybrid plants produce gametes, the two alleles segregate, half the gametes receiving the P allele and the other half the p allele. Gametes: This box, a Punnett square, shows all possible combinations of alleles in offspring that result from an F1 F1 (Pp Pp) cross. Each square represents an equally probable product of fertilization. For example, the bottom left box shows the genetic combination resulting from a p egg fertilized by a P sperm. Appearance: Purple flowers White flowers Genetic makeup: PP pp Appearance: Genetic makeup: Purple flowers Pp 1/ 1/ 2 P F1 sperm P p PP Pp F2 Generation P F1 eggs p pp Pp Figure 14.5 Random combination of the gametes results in the 3:1 ratio that Mendel observed in the F2 generation. Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings 2 p 3 :1 • A dihybrid cross – Illustrates the inheritance of two characters • Produces four phenotypes in the F2 generation EXPERIMENT Two true-breeding pea plants— one with yellow-round seeds and the other with green-wrinkled seeds—were crossed, producing dihybrid F1 plants. Self-pollination of the F1 dihybrids, which are heterozygous for both characters, produced the F2 generation. The two hypotheses predict different phenotypic ratios. Note that yellow color (Y) and round shape (R) are dominant. P Generation YYRR yyrr Gametes F1 Generation YR Hypothesis of dependent assortment yr YyRr Hypothesis of independent assortment Sperm RESULTS 1? YR 2 CONCLUSION The results support the hypothesis of independent assortment. The alleles for seed color and seed shape sort into gametes independently of each other. Sperm yr 1? 2 Eggs 1 F2 Generation ?2 YR YYRR YyRr (predicted offspring) 1 ? yr 2 YyRr yyrr 3? 4 1? 4 YR 1? 4 Yr 1? 4 yR 1? 4 yr Eggs 1 ? YR 4 1? 4 Yr 1? 4 yR 1? 4 yr YYRR YYRr YyRR YyRr YYrr YYrr YyRr Yyrr YyRR YyRr yyRR yyRr 1? 4 Phenotypic ratio 3:1 9? 16 YyRr 3? 16 Yyrr yyRr 3? 16 yyrr 1? 16 Phenotypic ratio 9:3:3:1 Figure 14.8 Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings 315 108 101 32 Phenotypic ratio approximately 9:3:3:1 • In incomplete dominance – The phenotype of F1 hybrids is somewhere between the phenotypes of the two parental varieties P Generation Red CRCR White CWCW Gametes CR CW Pink CRCW F1 Generation Gametes Eggs F2 Generation Figure 14.10 Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings 1? 2 CR 1? 2 Cw 1? 2 1? 2 CR CR 1? 2 CR 1? CR 2 CR CR CR CW CR CW CW CW Sperm • The ABO blood group in humans – Is determined by multiple alleles Table 14.2 Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • Inheritance patterns of particular traits – Can be traced and described using pedigrees Ww ww Ww ww ww Ww WW or Ww ww Ww Ww ww First generation (grandparents) Second generation (parents plus aunts and uncles) FF or Ff Ff Ff Third generation (two sisters) ww Widow’s peak Ff No Widow’s peak (a) Dominant trait (widow’s peak) Figure 14.14 A, B Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Attached earlobe ff ff Ff Ff Ff ff ff FF or Ff Free earlobe (b) Recessive trait (attached earlobe) • Fetal testing (b) Chorionic villus sampling (CVS) (a) Amniocentesis Amniotic fluid withdrawn A sample of chorionic villus tissue can be taken as early as the 8th to 10th week of pregnancy. A sample of amniotic fluid can be taken starting at the 14th to 16th week of pregnancy. Fetus Fetus Suction tube Inserted through cervix Centrifugation Placenta Placenta Uterus Chorionic viIIi Cervix Fluid Fetal cells Fetal cells Biochemical tests can be Performed immediately on the amniotic fluid or later on the cultured cells. Fetal cells must be cultured for several weeks to obtain sufficient numbers for karyotyping. Biochemical tests Several weeks Several hours Karyotyping Figure 14.17 A, B Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Karyotyping and biochemical tests can be performed on the fetal cells immediately, providing results within a day or so. CONCLUSION Since all F offspring had red eyes, the mutant 1 white-eye trait (w) must be recessive to the wild-type red-eye trait (w+). Since the recessive trait—white eyes—was expressed only in males in the F2 generation, Morgan hypothesized that the eye-color gene is located on the X chromosome and that there is no corresponding locus on the Y chromosome, as diagrammed here. P Generation W+ X X X X Y W W+ W Ova (eggs) F1 Generation Sperm W+ W+ W+ W W+ Ova (eggs) F2 Generation Sperm W+ W W+ W+ W+ W W W+ Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • Morgan crossed flies – That differed in traits of two different characters P Generation (homozygous) EXPERIMENT Morgan first mated true-breeding Wild type wild-type flies with black, vestigial-winged flies to produce (gray body, heterozygous F1 dihybrids, all of which are wild-type in normal wings) appearance. He then mated wild-type F1 dihybrid females with + + b b vg+ vg+ black, vestigial-winged males, producing 2,300 F2 offspring, which he “scored” (classified according to F1 dihybrid phenotype). (wild type) (gray body, normal wings) Double mutant (black body, vestigial wings) Double mutant (black body, vestigial wings) x b b vg vg Double mutant (black body, vestigial wings) b b vg vg Double mutant TESTCROSS (black body, x vestigial wings) CONCLUSION If these two genes were on different chromosomes, the alleles from the F1 dihybrid would sort into gametes independently, and we would expect to see equal numbers of the four types of offspring. If these two genes were on the same chromosome, we would expect each allele combination, B+ vg+ and b vg, to stay together as gametes formed. In this case, only offspring with parental phenotypes would be produced. Since most offspring had a parental phenotype, Morgan concluded that the genes for body color and wing size are located on the same chromosome. However, the production of a small number of offspring with nonparental phenotypes indicated that some mechanism occasionally breaks the linkage between genes on the same chromosome. Figure 15.5 Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings b+ b vg+ vg RESULTS b vg b+vg+ b vg 965 944 Wild type Black(gray-normal) vestigial b+ vg b vg+ 206 Grayvestigial 185 Blacknormal Sperm b+ b vg+ vg b b vg vg b+ b vg vgb b vg+ vg Parental-type offspring Recombinant (nonparental-type) offspring • Linked genes – Exhibit recombination frequencies less than 50% Testcross parents b+ vg+ Gray body, normal wings b vg (F1 dihybrid) Replication of chromosomes b+ vg Meiosis I: Crossing over between b and vg loci produces new allele combinations. b vg b+ vg+ vg b b vg vg b b vg b vg Meiosis II: Segregation of chromatids produces recombinant gametes with the new allele combinations. Gametes Black body, vestigial wings b vg (double mutant) Replication of chromosomes b vg Meiosis I and II: Even if crossing over occurs, no new allele combinations are produced. Recombinant chromosome Ova Sperm b+vg+ b vg b+ vg b vg+ b vg b+ vg+ Testcross offspring Sperm b vg Figure 15.6 b+ vg b vg 965 944 BlackWild type (gray-normal) vestigial b+ vg+ b vg+ b vg b vg 206 Grayvestigial b+ vg+ b vg b vg+ Ova 185 BlackRecombination normal b vg+ frequency b vg Parental-type offspring Recombinant offspring Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings 391 recombinants =2,300 total offspring 100 = 17% • Many fruit fly genes – Were mapped initially using recombination frequencies I Y II X IV III Mutant phenotypes Short aristae Black body 0 Figure 15.8 Long aristae (appendages on head) Cinnabar Vestigial Brown eyes wings eyes 48.5 57.5 67.0 Gray body Red eyes Normal wings Wild-type phenotypes Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings 104.5 Red eyes • Alterations of chromosome structure (a) A deletion removes a chromosomal segment. (b) A duplication repeats a segment. (c) An inversion reverses a segment within a chromosome. (d) A translocation moves a segment from one chromosome to another, nonhomologous one. In a reciprocal translocation, the most common type, nonhomologous chromosomes exchange fragments. Nonreciprocal translocations also occur, in which a chromosome transfers a fragment without receiving a fragment in return. A B C D E F G H A B C D E F G H A B C D E F G H A B C D E F G H Deletion Duplication Inversion A B C E F G H A B C B C D E A D C B E F G H M N O C D E Reciprocal translocation M N O P Q Figure 15.14a–d Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings R A B P Q F G H R F G H