* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download OUTLINE
Behavioural genetics wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
Genetic testing wikipedia , lookup
Maximum parsimony (phylogenetics) wikipedia , lookup
Genetic engineering wikipedia , lookup
Heritability of IQ wikipedia , lookup
History of genetic engineering wikipedia , lookup
Gene expression profiling wikipedia , lookup
Designer baby wikipedia , lookup
Human genetic variation wikipedia , lookup
Pharmacogenomics wikipedia , lookup
Population genetics wikipedia , lookup
Quantitative comparative linguistics wikipedia , lookup
Public health genomics wikipedia , lookup
Quantitative trait locus wikipedia , lookup
Microevolution wikipedia , lookup
Gene expression programming wikipedia , lookup
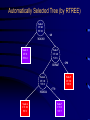
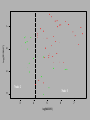
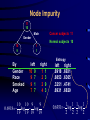
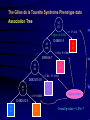
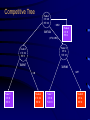
Recursive Partitioning And Its Applications in Genetic Studies Chin-Pei Tsai Assistant Professor Department of Applied Mathematics Providence University OUTLINE Genetic data Example Basic ideas of recursive partitioning Applications in genetic studies linkage analysis association analysis Recursive-partitioning based tools for data analyses Tree-based Analyses in Data Genetic Genetic Studies Nuclear Family Father Mother 1 1 00 1 2 1 2 00 2 1 1 3 12 1 2 1 4 12 2 1 Affected 1 1 5 12 1 1 2 1 6 12 2 2 2 1 00 1 2 2 2 00 2 1 2 3 12 1 1 3 4 5 6 2 4 12 2 2 Genetic Data Genotype 1 2 7 2 1 1 00 1 2 17 22 1 2 00 2 1 26 33 1 3 12 1 2 72 23 1 4 12 2 1 16 23 1 5 12 1 1 12 23 1 6 12 2 2 72 23 2 1 00 1 2 34 25 2 2 00 2 1 32 44 2 3 12 1 1 33 24 2 4 12 2 2 32 54 Application of Recursive Partitioning in Microarray Data (Zhang et al.,PNAS, 2001) Gene expression profiles of 2,000 genes in 22 normal and 40 colon cancer tissues Purpose: to predict new tissue Automatically Selected Tree (by RTREE) Node 1 CT:40 NT:22 >60 M26383 Node 2 CT: 0 NT:14 Node 3 CT: 40 NT: 8 R15447 Node 4 CT: 10 NT: 8 M28214 Node 6 CT: 10 NT: 1 >290 Node 5 CT: 30 NT: 0 >770 Node 7 CT: 0 NT: 7 7 Node 3 4 log(R15447) 6 5 Node 2 3 4 5 log(M26383) 6 7 7.5 Node 5 6.5 6.0 5.5 log(M28214) 7.0 Node 7 Node 6 4 5 6 log(R15447) 7 3-D Representation of Tree Concluding Remarks The three genes, IL-8 (M26383), CANX (R15447) and RAB3B (M28214), were chosen from 2,000 genes. Using three genes can achieve high classification accuracy. These three genes are related to tumors. Tree Growing Basic Ideas in Splitting criterion Classification Trees Goodness of Split = weighted sum of node impurities Impurity functions: entropy For binary outcome, y=0, 1, let p = proportion of (y=1). Entropy: -p log(p) - (1-p) log(1-p) where 0log(0) = 0 1/2 0 1/2 1 p Node Impurity 11 10 Cancer subjects 11 Male Gender Normal subjects 10 10 9 1 1 Entropy By Gender Race Smoked Age left 10 9 9 7 9 1 7 7 right 1 1 2 3 2 9 4 3 10 10 9 9 0.6918 log log 19 19 19 19 left .6918 .6853 .3251 .6931 right .6931 .6365 .4741 .6829 1 1 1 1 0.6931 log log 2 2 2 2 Goodness of Split Goodness of split s = p(L)i(L) + p(R)i(R) By Gender Race Smoked Age Entropy (i(t)) Weight (p(t)) left .6918 .6853 .3251 .6931 left right 19/21 2/21 16/21 5/21 10/21 11/21 14/21 7/21 No split: right .6931 .6365 .4741 .6829 s .6919 .6737 .4031 .6897 .6920 Tree Pruning Fisher Exact Test Misclassification cost and rate Cost-complexity and complexity parameter Optimal sub-trees Genetic Data Genotype 1 2 7 2 1 1 00 1 2 17 22 1 2 00 2 1 26 33 1 3 12 1 2 72 23 1 4 12 2 1 16 23 1 5 12 1 1 12 23 1 6 12 2 2 72 23 2 1 00 1 2 34 25 2 2 00 2 1 32 44 2 3 12 1 1 33 24 2 4 12 2 2 32 54 Key Idea in Tree-based Analysis If a marker locus is close to a disease locus, then individuals from a given family who are phenotypically similar are expected to be genotypically more similar than expected by chance. Sib pair 1 2 3 4 Tree-based Linkage Analysis Unit of observation: sib pair The response variable y takes three possible values depending on whether none, one, or both sibs are affected, which we arbitrarily coded as 0, 1, and 2. Covariate: the expected IBD (identity by descent) sharing at each marker locus Identity by Descent (IBD) 3 2 1 1 2 3 4 IBD=0 4 Sib 1 Father’s genotype Sib 2 Mother’s genotype 3 2 1 3 IBD=1 Genes (or alleles) inherited by relatives from the same ancestor. For two sibs, they can share at most one IBD gene from the father, and at most one from the mother. Thus, 0, 1, or 2 genes can be shared by two siblings. Sib 1 1 Sib 2 3 1 3 IBD=2 Sib 1 Sib 2 The Gilles de la Tourette Syndrome (GTS) Phenotype data (Joint work with Zhang et al., 2002) Genome scan of the hoarding phenotype collected by the Tourette Syndrome Association International Consortium for Genetics (TSAICG) Hoarding is a component of obsessivecompulsive disorder. We used data from 223 individuals in 51 families with 77 sib pairs. Genotypes are allele sizes from 370 markers on 22 chromosomes. The Gilles de la Tourette Syndrome Phenotype data Linkage Tree 23 28 26 > 1.9 P=0.0011 IBD Sharing at D5SMfd154 7 0 8 16 28 18 D5S408 0 8 0 > 0 P=0.0034 Split p-values 16 20 18 D4S1652 6 17 14 > 1.16 P=0.0078 10 3 4 Overall p-value = 2.63e-6 Tree-based Association Study The response variable is affection status. If a marker has n distinct alleles, then n covariates, each taking a value of 0, 1 or 2, are then constructed for this marker. For example, if n=7, then the 7 covariates take values (0,0,0,1,0,1,0) for a genotype of 4/6 and (0,0,0,0,0,0,2) for a genotype of 7/7. The covariates include gender, the parental phenotypes, race and the variables constructed using the marker information. The Gilles de la Tourette Syndrome Phenotype data Association Tree 85 135 >0 P=2e-4 Copies of Allele D4S403-5 46 106 39 29 > 0,NA P= 0.0017 D5S816-7 0 18 46 88 D4S2431-10 > 1,NA P= 0.016 0 11 46 77 > 0 P=0.0023 Split p-values D4S2632-5 19 54 27 23 Overall p-value = 1.03e-7 Why Recursive Partitioning? Attempt to discover possibly very complex structure in huge databases - genotypes for hundreds of markers - expression profiles for thousands of gene - all possibly predictors (continuous, categorical) No need to do transformation Impervious to outliers Easy to use Easy to interpret Recursive partitioning based tools for data analysis Classification and regression RTREE (http://peace.med.yale.edu) CART Multivariate Adaptive Regression Splines MASAL (http://peace.med.yale.edu) MARS Longitudinal data analysis MASAL (http://peace.med.yale.edu) Survival Analysis STREE (http://peace.med.yale.edu) References Books L. Breiman, J. H. Friedman, R. A. Olshen and C. J. Stone, 1984, Classification and Regression Trees, Wadsworth, California. H. Zhang and B. Singer, 1999, Recursive Partitioning in the Health Sciences, Springer, New York. T. Hastie, R. Tibshirani and J. Friedman, 2001, The Elements of Statistical Learning, Springer, New York. References Papers Zhang, Tsai, Yu, and Bonney, 2001, Genetic Epidemiology, 21, Supplement 1, S317-S322. Zhang, Leckman, Pauls, Tsai, Kidd, Campos and The TSAICG, 2002, American Journal of Human Genetic, 70, 896-904. Zhang, Yu, Singer and Xiong, 2001, Proc Natl Acad Sci U S A, 98, 6730-6735. Tsai, Acharyya, Yu and Zhang, 2002, In Recent Research Developments in Human Genetic. Recent Development Instability of Trees (high variance) Bagging – averages many trees to reduce variance (Breiman, 1996) Boosting (Breiman, 1998, Mason et al. 2000, Friedman el al. 1998) Random forest (Breiman, 1999) Lack of Smoothness MARS procedure (Zhang & Singer, 1999, Hastie et al. 2001) Difficulty in Capturing Additive Structure MARS procedure Competitive Tree for Colon Data 1.0 0.6 0.4 M28214 0.2 R15447 R15447 M26383 M28214 M26383 0.0 correlation 0.8 M26383 R15447 M28214 0 500 1000 1500 2000 Competitive Tree Node 1 CT: 40 NT: 22 >1052 R87126 Node 4 CT: 0 NT:6 (372, 1052] Node 3: CT: 6 NT: 13 Node 2 CT: 34 NT: 3 T62947 X15183 >457 >28 Node 5 CT: 0 NT: 3 Node 6 CT: 34 NT: 0 Node 7 CT: 0 NT: 13 Node 8 CT: 6 NT: 0 3-D Representation of Tree