* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download PDH02 - OSU Biochemistry and Molecular Biology
Protein–protein interaction wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Lipid signaling wikipedia , lookup
Catalytic triad wikipedia , lookup
Fatty acid metabolism wikipedia , lookup
Evolution of metal ions in biological systems wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Lactate dehydrogenase wikipedia , lookup
Proteolysis wikipedia , lookup
Oxidative phosphorylation wikipedia , lookup
Genetic code wikipedia , lookup
Point mutation wikipedia , lookup
Fatty acid synthesis wikipedia , lookup
Metalloprotein wikipedia , lookup
15-Hydroxyeicosatetraenoic acid wikipedia , lookup
Butyric acid wikipedia , lookup
Biosynthesis wikipedia , lookup
Specialized pro-resolving mediators wikipedia , lookup
NADH:ubiquinone oxidoreductase (H+-translocating) wikipedia , lookup
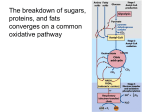
Citric acid cycle wikipedia , lookup
Amino acid synthesis wikipedia , lookup
The-Keto Acid Dehydrogenase Complexes Prepared by Franklin R. Leach Department of Biochemistry and Molecular Biology Oklahoma State University Stillwater, OK 9/15/02 The-Keto Acid Dehydrogenase Complexes A Brief Review 2002 Contents Introduction Background Complex Structure Regulation Whole Genomes Components Parts E1 E2 X E3 Relation of Lipoic Acid to Oxidative Damage Relation to Medicine Therapeutic potential Ischemic heart disease Lactic acidosis and related diseases Maple syrup urine disease and branched chain-related diseases Leigh's necrotizing encephalomyelopathy Alzheimer's disease Primary biliary cirrhosis Systemic sclerosis Lipoic Acid Activation and the Lipoamidase Reaction Recent Results on the Regulatory Enzymes Web Connections Literature Cited Appendix Symposium Honoring Lester Reed - A Tribute Recollection - From lipoic acid to multienzyme complexes Introduction Remarkable progress in understanding the function and mechanism of action of the keto acid dehydrogenases has been made in the last 50 years. These complexes are the classical example of a multienzyme complexes. A conference on "-Keto Acid Dehydrogenase Complexes: Organization, Regulation, and Biomedical Aspects" (1), held November 16-18, 1988 in Austin, Texas to honor Professor Lester J. Reed, summarized much of the early progress. This conference celebrated Dr. Reed's 65th birthday. My interest in this topic is because I did my dissertation research with Lester Reed (1953-57) and worked on the lipoic acid activating enzyme system. A Tribute to Lester Reed written by Tom Roche, Head of the Department of Biochemistry at Kansas State University and a former Reed postdoc, is in the appendix. It shows the extent to which Reed's laboratory has contributed to our understanding of the -keto acid dehydrogenases. The complexes have been isolated, their composition and organization determined, their base sequences are being elucidated, and their amino acid sequences and crystallographic patterns are being deduced. The mechanisms of regulation of the activities of these complexes have been established. A FASEB symposium reviews the topic (2). Several other reviews have appeared (3-6). At the 1994 ASBMB meeting in Washington, DC, Reed was given the ASBMB-Merck Award and presented a review of "A Trail of Research: From Lipoic Acid to Multienzyme Complexes". An American Institute of Nutrition Symposium "-Keto Acid Dehydrogenase Complexes: Nutrient Control, Gene Regulation, and Genetic Defects" was also held in 1994. A review paper from that symposium has appeared (7). Reed has recalled "From lipoic acid to multienzyme complexes" for Protein Science (7a). (See appendix). For the Journal of Biological Chemistry Centennial collection Reed (7b) traced "a trail of research from lipoic acid to -keto acid dehydrogenase complexes". The molecular understanding of the -keto acid dehydrogenases began in the 1950s with the isolation and determination of the structure of lipoic acid (8-10). The next key finding was the enzymatic mechanism by which lipoic acid was converted to the enzyme-bound functional form (11, 12). From that point until the current application of molecular biology techniques the emphasis of study has been on the isolation, characterization, determination of structure/function relationships, and regulation of the -keto acid dehydrogenase complexes (13-15). This has matured into recent and current determinations of the amino acid and base sequences for many components of these complexes (see refs. 1-7). The reaction (sum) catalyzed by the -keto acid dehydrogenases is: TPP, Lipoic acid, FAD RCOCO2H + NAD+ +CoASH &emdash;&emdash;&emdash;&emdash;&emdash;&emdash;&emdash;> RCO-SCoA + CO2 + NADH +H+ The component steps in this overall reaction are: CH3COCO2- + E1TPP + H+ <==> CO2 + E1 CH3C(OH)=TPP (1) E1 CH3C(OH)=TPP + E2_LipS2 <==> E1TPP + E2-Lip(SH)-S-COCH3 (2) E2-Lip(SH)-S-COCH3 + CoASH <===> E2_Lip(SH2) + CH3COSCoA (3) E2-Lip(SH2) + E3FAD <===> E2-LipS2 + Dihydro-E3FAD (4) Dihydro-E3FAD + NAD+ <===> E3FAD + NADH + H+ (5) Lewisite (CHCl=CHAsCl2) is a poison gas that was synthesized too late to be used in World War I. Rudolph Peters headed the Oxford University laboratory that searched for antidotes to chemical warfare agents. They developed BAL (British anilewisite, 2,3 dimercaptopropanol, and for security reasons, OX 217) in 1940. The target of the arsenical was lipoic acid. See (16). Lipoic acid, 1,2-dithiolane-3-pentanoic acid (6,8-dimercapto-octanic acid), functions in transacylation, redox, and transport reactions. It plays a central role in oxidative metabolism: the oxidative decarboxylation of pyruvate, branched chain amino acid metabolism, glycine decarboxylation, and in the citric acid cycle. Lipoic acid is formed from octanoic acid via an enzymatic S-insertion. Additional details have been learned about lipoic acid synthesis. The question of why mitochondria synthesize fatty acids has been answered: the synthesis of lipoic acid. Wada, Shintani, and Ohlrogge (17) established that pea mitochondria can acyl carrier protein and the enzymes to synthesize fatty acids. Radioactivity from labeled malonic acid was found in the H protein, a lipoylcontaining enzyme involved in glycine metabolism. In Saccharomyces cerevisiae Brody, Oh, Hoja, and Schweizer (18) found that absence of the yeast gene ACP1, resulted in a decreased lipoic acid content. They conclude that the mitochondrial ACP is invovled in the synthesis of octanoate which is a lipoic acid precursor. Jordan and Cronan (19) found that the acyl carrier protein of lipid synthesis could donate lipoic acids to the pyruvate dehydrogenase complex in both E. coli and mitochondria. Self, Tsai, and Stadtman (20) have prepared the selenotrisulfide derivatives of lipoic acid and lipoamide. The selenotrisulfide derivative of lipoic acid was an effective substrate for thioredoxin reductase. The lipoamide derivative was reduced by dihydrolipoamide dehydrogenase. The selenium analogs of lipoic acid had been used earlier by Reed, Morris, and Cronan (21) to isolate E. coli mutants. Replacement of either the C-6 or C-8 sulfur atom with Se gave lipoic acid derivatives with unaltered biological properties. The replacement of both S's with Se producing selenolipoic acid that was a growth inhibitor of E. coli. When radioactive 75Se was used, the selenolipoic acid was found incorporated in the -ketoacid dehydrogenases. Resistant mutants were isolated. These mutations were traced to lipoate-protein ligase and to an unknown function in the synthesis of lipoic acid. The E. coli LipA is a lipoyl synthase that forms lipoyl groups from octanoyl-ACP (22). This enzyme as well as biotin synthase contains (2Fe-2S) centers that can combine to form a (4Fe-4S) center. The iron-sulfur center is involved in the formation of a C-S bond (23). The enzymology of sulfur activation during thiamine and biotin biosynthesis has been discussed by Begley, Xi, Kinsland, Taylor, and McLafferty (24). The reactions for lipoic acid activation and its covalent attachment to the pyruvate dehydrogenase complex are: E1 + ATP + Lipoic Acid &emdash;&emdash;&emdash;&emdash;&emdash;> E1lipoyl-AMP + PP E1-lipoyl-AMP + E2 &emdash;&emdash;&emdash;&emdash;> Lipoyl-E2 + AMP + E1 Lipoyl-E2 + apo-PDHC &emdash;&emdash;&emdash;&emdash;&emdash;&emdash;> Lipoyl -PDHC + E2 Where E1 and E2 are the two enzymes of the Streptococcus faecalis lipoic acid-activating system. The lipoic acid is covalently bound to the -amino group of lysines of E2 of the pyruvate dehydrogenase complex (PDHC). PDHC consists of three distinct enzymes designated as E1, E2, and E3 - note that the distinction between the lipoic acid-activating enzymes and the -keto acid dehydrogenases made by a subscript number for the former and an on the line number for the latter. The reaction involved in lipoic acid removal (lipoamidase reaction) is: Lipoyl-PDHC + H2O &emdash;&emdash;&emdash;&emdash;&emdash;&emdash;&emdash;> apoPDHC + Lipoic acid Background The -keto acid dehydrogenases are large enzyme complexes that serve essential roles in metabolism (25). The pyruvate dehydrogenase (PDHC) provides the link between glycolysis and the citric acid cycle and produces acetyl-CoA for the citric acid cycle and acetyl groups for acetylcholine synthesis; in omnivores 50-80% of metabolism goes through the PDHC (26). The -ketoglutarate dehydrogenase functions in the citric acid cycle. The branched chain -keto acid dehydrogenase is important in regulation of nitrogen metabolism (26). These enzyme complexes involve five cofactors: thiamine pyrophosphate (TPP), lipoic acid (LA) in the form of enzyme-bound lipoamide (the amide between lipoic acid and the -amino group of lysine), nicotinamide dinucleotide (NAD+), coenzyme A (CoA), and flavin adenine dinucleotide (FAD) shown in Fig. 1 where E1 is the carboxylase (in the case of pyruvate, pyruvate dehydrogenase, EC # 1.2.4.1), E2 is the transacylase-reductase (in the case of pyruvate, dihydrolipoamide acetyltransferase, EC # 2.3.1.61), and E3 is dihydrolipoamide dehydrogenase (EC # 1.8.1.4). The activity of the mammalian forms of these enzymes is regulated by inhibition by products and by a phosphorylation-dephosphorylation cycle (involving insulin among other factors) (14). There are a series of 5 reactions indicated by the Arabiac numbers that constitute the complete reaction sequence. Reaction 1 is the decarboxylation of pyruvate with the production of CO2 and hydroxyethythiamine pyrophosphate. The hydroxyethylthiamine pyrophosphate is then oxidzed to acetylthiamine pyrophosphate with the reduction of lipoic acid. The acetyl group is then transferred to lipoic acid yielding the 8-S-acyl compound in reaction 2. The third reaction is the transfer of the acyl group to CoA yielding the acyl CoA derivatives. Lipoic acid is now in the dihydro form and must be reoxidized. This occurs in reaction 4. The reduced dihydrolipoyl dehydrogeanse is then oxidzed in reaction 5 producing NADH. TPP FAD Lipoic Acid NAD+ CoA The following scheme shows the reaction mechanism in detail. Complex Structure There are two polyhedral forms of E2: cubic and dodecahedral (8). The components of the mammalian complexes and E. coli are summarized in Table 1. Enzyme Abbreviation Mr x 10-6 Subunits # # # Mr x 10-3 Part A. Mammalian Bovine heart Native complex PDHC 8.5 Pyruvate dehydrogenase E1 0.154 E1 2 41 E1 2 36 60 60 6 50 2 55 E2 3.1 X Dihydrolipoyl dehydrogenase 60 E3 0.11 1 12 Kinase Phosphatase PDHk 0.1 PDHk 1 48 PDHk 1 48 PDHp 1 97 PDHp 1 50 PDHp 0.15 Part B. Bacteria E. coli Native complex PDHC 4.6 Pyruvate dehydrogenase E1 0.19 2 99 24 Dihydrolipoyl transacetylase E2 1.7 24 66 24 Dihydrolipoyl dehydrogeanse E3 0.112 2 where k is for kinase and p is for phosphatase. The amino acid sequence of protein X differs from that of E2, but both contain acetylable lipoamide. Protein X may contribute to assembly of the complex (27). Protein X is also called E3BP now that its function has been established There is an unique structural organization of the Saccharomyces cerevisiae pyruvate dehydrogenase complex (28). The Reed group used truncated E2, BP and various physical techniques to determine the arrangement. The showed that there were 12 large openings in the E2 core multimer that permitted entrance of BP into the central cavity. Various model structures are depicted. Regulation The -keto acid dehydrogenase complexes are regulated by end-product inhibition by NADH and the appropriate acyl CoA. In addition there is regulation by phosphorylationdephosphorylation (14,15). This covalent modification cycle is in turn regulated by many components, as is shown in Fig. 2; this occurs for the pyruvate and branched chain ketoacid complexes. The pyruvate dehydrogeanse complex of yeast is regulation by phosphorylation (29). Olson and his group at UT San Antonio have reviewed the regulation of pyruvate dehydrogenase multienzyme complex in the Annual Review of Nutrition (30). 51 24 Figure 2. Regulation of the mammalian and yeast pyruvate dehydrogenase complexes by phosphorylation/dephosphorylation. Whole Genomes A cluster of genes that encode the branched-chain -keto acid dehydrogenase from Streptomyces avermitilis has been cloned and sequenced (31). ORF1 has E1 with 1,146 nucleotides encoding a 381 amino acid protein of MW 40,969 Da. ORF2 (E1) 1,005 nucleotides would code for a 334 amino acid protein of MW 35,577. The inner genic distance is 73 nucleotides. The ATG start codon of ORF3 overlaps the stop codon of ORF2; ORF3 has part of an E2-like sequence. The sequence and organization of the genes encoding enzymes involved in pyruvate metabolism in Mycoplasma capricolum has been analyzed (32). Three operons were found: 1) naox encoding a NADH-oxidase and lplA coding for lipoyl protein ligase, 2) odpA for E1 and odpB for E1, and 3) odp2 encodes E2 with a single lipoyl domain and dldH a modified E3 that contains a lipoyl domain. The cloning, structure, chromosomal localization, and promoter of human 2-oxoglutarate dehydrogenase gene has been reviewed by Koike (33). The cDNA contains a 3006-bp open reading frame encoding a 40-amino acid leader peptide and a 962-amino acid mature protein with Mr of 108,878. There are 22 exons spanning 85 kb. The gene is located on chromosome 7 at p13-p14. There are two 10-bp cis-acting elements and two trans-acting elements with a nuclear factor binding to region -63 to -24 that includes the two cis-acting elements involved in the control of synthesis. The gene and subunit unit organization of the bacterial pyruvate dehydrogenase complexes has been reviewed by Neveling, Bringer-Meyer, and Sahm (34). Componet Parts E1. The E1 component contains TPP and catalyzes the decarboxylation of the -keto acid with the generation of reduced and acylated E2. A tightly bound enzyme intermediate in the process is 2-(1-hydroxyethylidene)-thiamine pyrophosphate when the substrate is pyruvate (35). The nucleotide sequence for the ace E gene of Escherichia coli has been determined (36). The ace E structural gene contains 2,655 base pairs coding for 885 amino acids excluding the initiator. The relative molecular mass of 99,474, the amino-terminal residue, and carboxyl-terminal sequence predicted from the nucleotide sequence are in excellent agreement with published information on E1. The upstream gene A produces protein A of Mr 27,049 which has the helix-turn-helix structure characteristic of a positive regulator (37). There is little similarity between the sequences of the E1 enzymes of E. coli and bovine heart. The amino acid sequences of the E1b subunit [the subunit of E1 for the branch chain complex] of rat liver (30), the E1b subunit of human liver (39), the E1p subunit of human liver (40-42), the E1b subunit of the bovine liver (43), the E1b and E1b of Pseudomonas putida (44), and the E1p subunit of human liver (42) have all been determined from cDNAs. All three reported human E1 cDNA sequences have significant differences still to be be resolved (4). The ace E gene encoding the E1 for pyruvate (36) and the suc A gene (45) encoding the 2-oxoglutarate dehydrogenase have been sequenced for E. coli. The yeast E1p has 333 amino acids and 36,486 kD (46). A cDNA has been cloned and its amino acid sequence deduced for the E1p from Arabidopsis thaliana (47). It has about 50 % sequence identity and the phosphorylation site and active site cysteines are conserved. Tripatara, Korotchkina, and Patel (48) analyzed human point mutations in E1 and found R349 is critical for activity, M181 is involved in TPP binding, and P188 is necessary for structural integrity of E1. The crystal structure of 2-oxoisovalerate heterotetrameric (22) E1 has been solved (49) for the enzyme from Pseudomonas putida. This is available as PDB file 1QS0. The TPP cofactor is bound at the phosphate end by the -subunit and the aminopyrimidine end by the -subunit. The amino acid around the binding site include Y133, R134, G182, L184, D213, A215, N242, W244, I246 H312, I60', Y88', and H131'. The lipoyl moiety of the E2 visit either H312 or H131', residues that are probably involved in the catalytic mechanism. The determination of the E1 structure allows for the first time the construction of a model of the 2-oxo acid dehydrogenase multienzyme complexes. The core in P. putida branched-chain dehydrogenase is a cubic arrangement of 24 E2 subunits. During a functional cycle, the lipoyl domain swings between the three enzyme components communicating among the three active sites. At the beginning of the cycle, the disulfide at the tip of the lipoyl domain is in the S-S or oxidized form. The 2-oxo acid is decarboxylated by E1 using the TPP cofactor. The substrate is oxidized to an acyl group and the lipoyl is reduced and then acylated. The CoA enters the active site of E2 from inside the complex and then accepts the acyl group. The lipoyl moiety is fully reduced. It is in turn oxidized by E3. See http://www.bmsc.washington.edu/people/hol/WimFigs5.html for the illustrations. The crystal structure of the human branched-chain -ketoacid dehydrogenase has also been solved at a resolution of 2.7 Å (50). This E1b is a 170 kDa 22 heterotetramer. The subunit is a 45.5 kDa protein containing 400 amino acid residues and the subunit is a 37.8 kDa protein. This structure differs from the P. putida one by having a 30-resiude N-terminal tails that intertwin in a firm handshake whereas the P. putida structure extents to opposite sides on the tetramer far from crossing paths. There are two K+ binding sites involving L164, T165, Q167, S161, and S162 for the first and G128', N183, L130, C178, T131, and D181. The small C-terminal domain of the human subunit is 16 residues lower than the counterpart from P. putida this gives a longer last helix and an irregular tail. There is an important mutation in this region Y393N- causes one form of MSUD. The TPP binding site involved E76', S162, Y102', A195, G194, L74', Y113, R114, R220, H291, and I226 E2. The E2 component is the central part of the complex and both E1 and E3 bind to it. E2 contains covalently bound lipoyl moieties and participates in the generation of the acyl groups and their subsequent transfer to CoA. Three domains are defined in E2 as seen in Fig. 3 below There are one, two, or three lipoyl domains each consisting of about 100 amino acid residues at the N-terminus depending on the E2 (51). The domains are joined by flexible regions rich in Ala and Pro. The NMR spectrum of a 32-residue synthetic peptide corresponding to the flexible region is similar to those of the intact complex (52). Thus, the three flexible regions are exposed to solvent and enjoy considerable conformational flexibility. There are separate domains on E2 for the binding E1 and E3 (53). The acyl transferase activity is located toward the C-terminus. The lipoyl moieties are bound through the -amino group of lysine and there is considerable sequence conservation around the actual lipoic binding sites among the various E2s consensus sequence for 12 different lipoyl domains from E. coli (54), Azotobacter vinelandii (55), human and bovine branched-chain dehydrogenase (56), bovine kidney (57), bovine heart (58), and chicken liver H protein (59) is: L4E9S5D9 K12A10S8M6 E7V6P8 The specific peptide sequences on E1 that are phosphorylated are shown below (14). Tyr-His-Gly-His-Ser-Met-Ser(P)-Asn-Pro-Gly-Val-Ser(P)-Tyr-Arg Tyr-Gly-Met-Gly-Thr-Ser(P)-Val-Glu-Arg . Figure 3. The domains of the E2 enzymes. The actual sequences for several enzymes is shown in Table 2 below. Table 2. Amino Acid Sequences Among the Lipoyl Moities. Escherichia coli (40) E2pL 1 E Q S L I T V E G D K* A S M E V P S P Q A E2pL 2 E Q S L I T V E G D K* A S M E V P A P F A E2pL 3 E Q S L I T V E G D K* A S M E V P A P F A E2oL 1 D E V L V E I E T D K* V V L E V P A S A D Azotobacter vinelandii (41) E2p---L 1 Q G L V V L E S A K* A S M E V P S P K A E2pL 2 Q S L I V L E S D K* A S M E I P S P A E2p------L 3 Q S L I V L E S D K* A S M E I P S P A A G Human and bovine (42) E2bL 1 F D S I C E V Q S D K* A S V T I T S R Y D Bovine kidney (43) E2oL 1 I E T D K* T S V Q V P S P A N G Bovine heart (44) E2pL 1 V Q T D K* A T V G F E2p-----------L 2 K* A T I G F Chicken liver H protein 45) D D E F G A L E S V K* A A S E L Y S P L T NMR analysis has been applied to the lipoyl domains (60 and 61). There are two antiparallel -sheets of four strands each. The lipoyl-lysine residue is found in a type-I turn connecting two -strands. There is a high structural similarity of the lipoyl domains in spite of only 25 % sequence identity. The inner lipoyl domain of E2 is invovled in the interaction of pyruvate kinase with the complex (62). The lipoyl-lysine residues have been postulated to swing between E1 and E3 subunits while accomplishing their functions. X-ray crystallographic evidence from an examination of the Bacillus stearothermophilus complex structure has supplied additional information for a model of Perham and Hol and their colleagues (63) that shows how the lipoyl group can visit the active sites of the E2 and E3 during catalysis. The lipoyl domain bearing subunit, E2, serves as the central protein in forming the multienzyme complex, communicates between subunits, is involved in active-site coupling, conformational mobility, substrate specificity, and metabolic regulation (64). Jones, Stott, Reche, and Perham (65) found that the lipoyl domains of the E2 subunit undergo conformational changes when interacting with their homologous E1 but not heterologous E1s. It is evident that recognition of the protein domain is the ultimate determinant of whether reductive acylation of the lipoyl group occurs, and this is ensured by a mosaic of interaction with the E1. Roche's group (66) evaluated the contribution of particular amino acids of the L2 lipoyl domain of human PDHC. The specificity loop contains L140, S141, and T143 whose mutagenesis influencing enzyme activity. Other residues that markedly reduce activity are E162, D172, A174, and E179. The lipoylated K is K173. The influential residues are spread over >24Å of one side of the L2 domain and this side would support extensive contacts between the E1 and L2 domain. Thus surface residues contribute to the unique surface patterns that enable recognition. The structure of the lipoyl domains has been determined by multidimensional NMR. Perham and his group (67) have concluded that there is a greater restriction in the motion of the lipoyl-lysine swinging arm of the E. coli pyruvate dehydrogenase complex than previously thought. Reductive acetylation of the lipoyl moiety gave larger chemical shifts than expected and multiple resonant forms. These observation imply a change in conformation upon acetylation and multiple conformations which may stabilized this catalytic intermediate. The E2 of maize pyruvate dehydrogenase complex contains a single lipoyl domain (68). There are two distinct E2s 50-54 kDa and 76 kDa. Arabidopsis thaliana aslo contains a single lipoyl domain E2 suggesting that all plant mitochondrial PDHs contain an E2 with a single lipoyl domain. The nucleotide sequences have been determined for aceF encoding the E2p of E. coli K12 (69), the sucB encoding the E2o of E. coli K12 (70), the E2b of human and bovine (56), the E2b from placenta (71), the E2p of human (72), the E2b of P. putida (73), and the E2p of A. vinelandii (55). The amino acid sequence of the lipoyl domain has been determined for Bacillus stearothermophilus (74). The E2 inner core domain of E2s (bovine E2b, human E2p, E. coli E2p, and E. coli E2o) is conserved (75). The crystal structure of the catalytic domain of A. vinelandii E2p has been determined (76). His B610, Ser C558, Tyr B608, Ile C571, Phe C568, and Leu C580 are closer to the CoA binding site. Further details have been published (77). CoA and lipoate are found in extended conformation at the two opposite entrances of the 30 Å long channel which runs at the interface between two 3-fold-related subunits and forms the catalytic center. The reactive groups of both (-SHs) are within hydrogen bond distance of the side chain of His 610. There is suggestion of a direct hydrogen bond between Ser 558 to one of the two peptide bonds in CoA. Site-directed mutagenesis studies (78) have revealed that in bovine E2b His 391 and Ser 338 chagnes modified catalytic activity. Ala 348 presumably contacts CoA and plays a key role in the substrate preference. The solution structure of the lipoyl domain of the 2-oxoglutarate dehydrogenase complex from Azotobacter vinelandii has been determined (79). The lipoyl domains are solvent exposed. A recognition of the lipoyl domain-containing surface loop underlies the substrate channelling in the PDHC (80). Trp-22 plays a central role in anchoring two four-stranded sheets which positions the lipoyl attached to lys-43 at the tip of an exposed loop (81). The flexibility of the linker segment of E2 is thought to be a detriment toward crystallization. Smaller parts of E2 such as 24-mer and 60-mer have been analyzed using NMR and x-ray crystallography. The E2o of E. coli occurs as trimer when expressed with a C-terminal [His6] tag (82). Using molecular replacement the structure has been solved to 3.0 Å resolution. The conserved (5 sequences) amino acids in the E2o lipoyl domain are P-X3-ES-X13-G-X5-E-X4-IETDK-X3-V-X5-G and for the catalytic domain part 1 are M-X-R-X-R-X3-A-X-RL-X-E and part 2 D-X3-AV-X4-GLV-X-PV-X-R. X, aka E3BP. Associated with the E2 core is another lipoyl-containing component, Protein X, that undergoes reduction and acetylation (83). The region of protein X that contains the lipoyl moiety is structurally and antigenically related to the lipoyl-bearing portions of E2. However, the bulk of protein X is distinct in sequence from the structure of E2 (84, 85). Protein X is found in liver, kidney, heart, adipose tissue, spleen, skeletal muscle, testes, uterus, red blood cells, and brain of the rat. Thus, it is not just a tissuespecific isozyme of E2 (86). The outer domain of X binds and facilitates regulation of the catalytic subunits of the kinase (87). It is believed that E3 associates with X. The conditions for reconstitution of mammalian pyruvate and 2-oxoglutarate dehyrogenase complexes have been established (88). Protein X is selectively cleaved using arg C and reconstitution activity is decreased. Adding a large excess of E3 gives a more effective reconstitution. Reed and colleagues (89) find that E3BP anchors E3 homodimers inside each of the 12 pentagonal faces of the 60-mer E2. Steric hinderance by the lipoyl and E3 binding domains limits the binding of E2. A novel dihydrolipoyl dehydrogenase binding protein that lacks the amino terminal lipoyl domain has been found in Ascaris suum (90). The stoichiometry of E3BP interaction is 12 mol per mol of PDHC and 60 mol of E2 per mol of PDHC (91). The lipoyl domains of E3BP can substitute for the lipoyl domains of E2 in overall complex catalytic activity. It may have an unique catalytic function. E3. One dihydrolipoyl dehydrogenase gene, lpd, codes for the E3 that serves each of the -keto acid dehydrogenases. The gene is comprised of 1,419 base pairs (473 codons excluding the initiating AUG). The composition, Mr (50,554 or 51,274 if the FAD cofactor is included), the amino-terminal sequence, and the carboxyl-terminal sequence predicted from the nucleotide sequence are in excellent agreement with previous biochemical studies on the enzyme (92). E3 is similar in structure to glutathione reductase (EC # 1.6.4.2), a related disulfide oxido-reductase (93). The crystal structure of glutathione reductase has been determined (94). The sequence of cDNAs encoding E3 has been determined for porcine and human (95) and yeast (96, 97) genes. The lipoamide dehydrogenase gene of A. vinelandii has been cloned in E. coli (98). This protein has 40% conservation of amino acid residues when compared with the E. coli enzyme. When compared with the three-dimensional structure of glutathione reductase, all of the essential residues in four domains are conserved. E3 may have other functions such as in sugar transport in E. coli (99-101). E3 has been reviewed by Patel and his colleagues (102). A lipoamide dehydrogenase that contains a lipoyl domain has been found in Neisseria menginitidis (103´). This protein is membrane-associated and may be invovled in transport processes. Relation of Lipoic Acid to Oxidative Damage Dihydroipoic acid reduces neuronal injury after cerebral ischemic (104). Expression of fos and jun genes were delayed in the presense of dihydrolipoic acid but were accelerated in presence of lipoic acid (105) suggestion a role of redox. Dihydrolipoic acid prevents hypoxic/reoxygenation and peroxidative damage in rat heart mitochondria (106). Both lipoate and dihydrolipoate prevented singlet oxygen-induced damage of DNA (107). Lipoic acid can prevent symptoms of vitamin E deficeincy (108) suggesting that many of the reductive (antioxidative) elements are in communication. Lipoate prevents serum albumin glycation (109). It improves memory in aged mice (110). The naturally occurring optical isomer of lipoic acid is more active in improbing blood during reoxygenation (111). Lipoic acid protects against cerebral ischemia-reperfusion (112). Ischemic-reperfusion injury in humans occurs in conditions such as strokes, cardiac arrest, subarachnoid hemorrhage or head trauma. Oxidative injury due to oxygen free radicals occurs. Lipoic acid protects rats against reperfusion injury following cerebral ischemia (113). Lipoic acid is also neuroprotective in focal cerebral ischemia in rodents (114). In insulin-resistant rat skeletal muscle addition of lipoic acid enchaned insulinstimulated glucose metabolism (115). Ziegler and colleagues (116) evaluated the efficacy and safety of oral treatment with the antioxidant lipoic acid in non insulin dependent diabetes mellitus patients with autonomic neuropathy. Treatment with lipoic acid was well-tolerated and may slightly improved the patients with cardiovascular autonomic neuropathy symptoms. Packer and Cardenas (117) edited a monograph that considered Biothios in Health and Disease. This is an excellent summary of the status in 1992 (published in 1995). Relation to Medicine Structural basis in humans. Malfunctioning of any of the three -keto acid dehydrogenases leads to clincal manifestations. Deficiency of the pyruvate dehydrogenase complex predominantly leads to lactic acidosis, impaired neurological function, and delayed growth and development. Deficiency in the branched chain enzymes leads to maple urine disease. There is an association between both Alzheimer disease and Parkinson disease and the-ketoglutarate dehydrogenase genes. The structural basis of these effects have been reviewed by Hengeveld and de Kok (117a). Therapeutic potential. The role of lipoic acid in liver metabolism and disease has been reviewed (118). It serves as an antioxidant at 600 mg doses. It has been used to treat alcohol-induced damage to the liver, mushroom poisoning, metal intoxication, and CCl4 poisoning. Lipoic acid has a well-defined molecular role in biochemistry and a still diffusely defined role in pharmacology. Ischemic Heart Disease. Clofibrate, an antilipidemic agent, is useful in primary prevention of ischemic heart disease. The branched-chain ketoacid dehydrogenase complex, BCKADHC, is displaced from serum albumin by clofirate and enzyme activity in the heart is increased activity by clofibrate treatment (119, 120). Feeding clofibrate to rats increases the activity of BCKADHC 3-fold (121). Administration of clofibrate to rats both activates and induces the BCKADHC. The increased synthesis would require additional lipoic acid and possibly the activating enzymes. Induction has been demonstrated for carboxylase (E1) and dihydrolipoamide transacetylase (E2), but not dihydrolipoamide dehydrogenase (E3) (122). The BCKADHC increased by clofibrate is mitochondrial and not peroxisomal. Clofibrate causes peroxisomal proliferation (123). The activation is due to kinase inhibition (124). Lactic acidosis and related diseases. Inborn errors of energy metabolism as a group affect 1/5000. The cause of lacticacidemia represents a significant diagnosis problem. A cell tries to maintain its ATP level at all cost and a universal consequence of failure of mitochondria to produce adequate ATP is excessive lactic acid production. Defects of PDHC lead to fatal neonatal lacticacidosis, psychomotor retardation with or without neurodegeneration and a male-only syndrome of ataxia, mild mental retardation and carbohydrate sensitivity (109). Most tissues can survive with little or no PHDC activity when they can use other energy substrates, but in the brain cells that have PDHC defects usually die. Clinical improvement was achieved by oral administration of lipoic acid to an 8-monthold boy who had primary lactic acidosis due to a deficiency of E3 (125). Lipoic acid treatment may be useful in reducing serum-mediated cytotoxicity in patients with acute or chronic alcohol toxicity (126). PDHC abnormalities have been reported in over 50 patients; these patients had ataxia, psychomotor retardation, Leigh's disease, and/or some had lactic acidosis (127). There can be tissue specificity in the deficiency since the deficiency of both subunits of E1 was not expressed in some fibroblasts when the activity of the PDHC was reduced by 70% in the kidney (128). E1 deficiency is associated with lactic acidosis and central nervous system dysfunction (129). Several mutations account for the molecular heterogeneity of PDHC deficiencies. For example, with 11 patients (129), 7 had E1a and E1b proteins and mRNAs, 2 had mRNAs but no proteins, and 2 had only E1b mRNA. The cDNA coding for E1ap was isolated from a patient with lactic acidosis. Endo et al. (130) showed that there was a deletion of four nucleotides upstream from the normal termination codon which made a new termination codon 33 bases downstream. mRNA was present in this patient. This study was the first cloning of a defective gene of PDHC. The effects of mutations in the E1p subunit are extremely varied: lactic acidosis, hydrocephaly, enlarged ventricles, Leigh disease, psychomotor retardation, microcephaly, developmental delay, agenesis of the corpus callosum, dilated ventricles, cortical thinning, ataxia, mental retardation, and comatose episodes. Robinson and colleague analyzed these 14 patients for PDH-complex activity (ranged from 3.5 % to 100 %). The individual with 100 % activity had a 46-bp repeat (frame-shift mutation results) and premature termination (28 amino acids less). The activity in the brain was undoubtedly reduced (131). Even an amino acid substitution in the mitochondrial import sequence of the precursor protein can reduce the PDH-complex activity in fibroblasts (132). Because the E1ap gene is located on the X chromosome the effects of mutations differ in males and females (133). A patient with a defect in the X-lipoyl-containing component of the PDHC can cause neonal lactic acidemia (134). Thiamine treatment of a male child who was PDH deficient and had lactic acidosis (135) improved his condition The A199T E1 mutation decreases the affinity for TPP and yields a complex with 25 % of normal activity and a 10-fold increase in the Km for pyruvate (136). It is suggested that the addition of lactic dehydrogenase inhibitors would be useful in treatment. Lactic acidosis is a broad-anion gap metabolic acidosis caused by either lactic acid overproduction or underutilization. Treatments of the underlying cause of the lactic acidosis are ideal (136a). Maple syrup urine disease (MSUD) and branched chain-related diseases. Deficiency in the branched-chain -keto acid dehydrogenase cause hypervalinemia, hyperleucineisoleucinemia, maple syrup urine disease, isovaleric acidemia, glutaric aciduria type II, ethylmalonic-adipic aciduria, 3-methylcrotonyl CoA carboxylase deficiency, 3-hydroxy3-methylglutaryl CoA lyase deficiency, and 3-ketothiolase deficiency (137). Danner et al. (138) established the immunologic absence of the E2 protein in an individual with MSUD. Another study (139) on cell lines derived from patients with MSUD showed a marked decrease in E1b and faint immunostaining of E1a. Fisher et al. (140) found five types of defects: type I, reduced E1 activity but normal amounts of E1a and E1b; type II, reduced amounts of E1a and E1b; type III, E1a mRNA reduced; type IV, E2 mRNA reduced, and type V, E2 reduced or absent. Thus, MSUD is a complex disease at the molecular level. A history of MSUD from 1954 when it was recognized to 1993 has appeared (141). The incidence is 1:200,000 live births. Numerous mutations in the BCKD have been noted. Retroviral gene transfer has been used to correct the disease in lymphoblasts from a Mennonite MSUD patient where Tyr 393 has been converted to Asn in E1b. There are gender differences in the regulation of the branched-chain 2-oxo acid dehydrogeanses (142). In male and female rats there were differences in response to light-dark cycles. The females showed a profound diurnal rhythm while the males did not. Danner and Doering at Emory have an on-line review of human mutations that affect BCKD (143). His group (144) also the transport of the three enzyme components from the cytoplasm to the mitochondria. E1b transport is rate limiting. There are three groups of MSUD mutations (50): 1) mutations that affect cofactor binding (R114W, T166M, R220W, and N222S; 2) mutations that affect the hydrophobic core (M64T, G204S, A208T, T265R, and I281T; and 3) mutations that affect subunit association (N126Y, Q134K, H156R, A209D, A240P, G245R, R252H, F364C, Y368C, and Y393N. In Turkish patients 3 missense disease specific mutations Q80E, C213Y, and T106M in the E1 gene and a polymorphism F280F were new mutations that produced MSUD (144a). Also there was a splice site mutation in the E2 gene. Some 36 children with MSUD were treated with a protocol that 1) inhibits endogenous protein catabolism, 2) sustains protein synthesis, 3) prevents deficiencies of essential amiono acids and 4) maintains normarl serum osmolarity (144b). All survived but infection or injuries can lead to deteroriation with life-threating neurological function at any time. Leigh's necrotizing encephalomyelopathy. In the case of an infant with lactic acidosis and developmental delay with neuropathological changes consistent with Leigh's necrotizing encephalomyelopathy, there was systemic deficiency of both subunits of E1 (145). A patient with well-documented clinical and biochemical PDHC deficiency was found on postmortem examination to have the specific CNS pathology of Leigh's disease (146). The PDHC activity was about 20% of that of normal individuals. Therefore, a subgroup of Leigh's disease is due to a PDHC deficiency. 20% of that of normal individuals. Therefore, a subgroup of Leigh's disease is due to a PDHC deficiency. Blass and coworkers (147) restored PDHC activity to extracts from skin fibroblast cells from two patients by adding E3. This shows that Leigh's disease can result from abnormalities in either the E1 or E3 components of the complex. A defect of X has been identified in two patients with encephalomyelopathy (148). Alzheimer's disease. The activity of 2-ketoglutarate dehydrogenase complex is reduced by more than 75% in patients with Alzheimer's disease (149). A 50% deficiency in PDHC activity has also been shown (150). In 20 patients with early-onset dementia of the Alzheimer type there was a 44% reduction in the cerebral metabolic rate for glucose (151). This breakdown in glucose metabolism appeared to involve the PDHC, but it is not clear if it is the primary defect. Both PDHC and KGDHC are reduced in Alzheimer diseased brains (152). Brains that are aging or have Alzheimer disease switch from glucose as their fuel source to ketone bodies. Heininger (152a) postulates that a soluble factor Abeta or deprivin is the metabolic switch and inhibits pyruvate dehydrogenase. In the older or Alzheimer brain the ketone bodies are an insufficient energy source. Therefore, outer regions of the brain are shut down to conserve fuel for survival of the inner core. Primary biliary cirrhosis. The autoantibodies of primary biliary cirrhosis recognize E2 and inhibit its function (153). In 10/10 patients there was 30-100 % inhibition of PDHC activity and the inhibition was directly proportional to the ELISA value (151). Another group of investigators has identified the 70-kD M2 autoantigen as the E2 of mitochondrial PDHC. In this study 38/40 patients' sera reacted positively with E2 (154) A synthetic peptide containing the lipoic acid binding site, KATIGF, absorbed the reactivity of sera from patients having primary biliary cirrhosis (155). The target for the autoantibodies corresponds to a functional site of dihydrolipoamide acetyltransferase. Would these sera also block lipoic acid activation since they are directed toward the same amino acid sequence? All 40 patients with primary biliary cirrhosis have autoantibodies directed against at least one of the E2 components of the -keto acid dehydrogenases. Treatment of this disease accounts for about 80 % of European liver transplants (156). The primary structure of the human M2 mitochondrial autoantigen E2 has been determined (156a). There are two presumed lipoyl bearing sequences: ETKATVGFE and ETKATIGFE. Avidin, the protein that avidly binds biotin, interacts with the lipoyl domains in PDHC (157). Both lipoic acid and biotin are bound to -amino groups of lysines and there is similarity in the amino acid sequence around these lysines. An octadecapeptide from amino acid residue 167-184 was specifically recognized by the anti-M2 autoantibodies from patients with primary biliary cirrhosis (158). A flow cytometric method has been developed to detect the anti-PDH antibody in primary biliary cirrhosis (159). The presence of lipoyl residues is crucial for recognition of human PDCE2 by antisera from patients with primary biliary cirrhosis (160). A library of 21 monoclonal antibodies to the E. coli PDH-complex and its components have been developed (161). All the antibodies were bound to the complex; 17 were bound to the E1 subunit. The degree of inhibition of enzymatic activity varied from 0 to > 98 %. A new assay for E1 activity based on N-acetyl-4-thiopyridine was developed. Competitive epitope mapping revealed that there were at least six separate binding regions (162). One antibody counteracted the GTP regulation of PDH-complex. GTP is an allosteric inhibitor. A semiautomated PDH-complex assay based on use of microtiter plates has been developed for the diagnosis of primary biliary cirrhosis (163). The results of assays following cyclosporin treatment showed a decrease in antibodies that correlated with the improvement of liver function. Halothane hepatitis has associated antibodies that are directed against the lipoyl-containing components of the 2-oxoacid dehydrogenase complex (164). The mechanisms of autoimmune liver disease have been reviewed (164a). Chronic viral infections could initiate primary biliary cirrhosis (164b). There isonly one treatment that is effective other than liver transplant and it is ursodeoxycholic acid (164c). The diagnosis usually uses three criteria: 1) elevated alkaline phosphatase, 2) antimitochondrial antibodies, and liver biospy. Systemic sclerosis. Many patients with have the multisystem connective tissue disorder of systemic sclerosis produce antibodies to the E1 (165). Systemic sclerosis is also an autoimmune disease. Lipoic Acid Activation and the Lipoamidase Reaction Lipoic acid is enzymatically activated in an ATP-dependent reaction before being covalently incorporated into the E2 and protein X (E3BP) units of the -keto acid dehydrogenases. The reaction is E1 + ATP + Lipoic Acid &emdash;&emdash;&emdash;&emdash;&emdash;&emdash;&emdash;&emdash; > E1-lipoyl-AMP + PP E1-lipoyl-AMP + E2 &emdash;&emdash;&emdash;&emdash;&emdash;&emdash;&emdash;&emdash; &emdash;> Lipoyl-E2 + AMP + E1 Lipoyl-E2 + apo-PDHC &emdash;&emdash;&emdash;&emdash;&emdash;&emdash;&emdash;&emdash; &emdash;&emdash;> Lipoyl-PDHC + E2 Yang and Frey (166) found that only the R-enantiomer of lipoic acid is a substrate for the enzymes of the complexes. Morris et al (167) have identified the gene encoding the lipoate-protein ligase A in E. coli. It codes for a 337 amino acid residue protein of 38 kD that attaches radioactive lipoic acid to apopyruvate dehydrogenase. A lipoyltransferase that catalyzes the second reaction above involving the lipoyladenylate was purified and characterized by Fujiwara, et al. (168) from bovine liver mitochondria. The same group (168a) purified the lipoic acid activating enzyme to homogeneity and showed that GTP was 1000 times more active than ATP. The product was lipoyl-GMP. cDNA clones encode a precusor of the enzyme. The amino acid sequence is identical to that of the medium-chain fatty acid CoA ligase. The lipoate protein ligase from E. coli has been purified and characterized by Guest and colleagues (169). The purified enzyme is monomeric with a Mr of 38000. It is inactivated by forming an intramolecular disulfide. Studies with mutants that lack this enzyme have another lipoylation system. The lipoamide arm of the glycine decarboxylase is not freely swinging (170). Lipoic acid is the prosthetic group for the H-component of the glycine decarboxylase that catalyzes the oxidative decarboxylation and deamination of glycine with the formation of CO2, NH3, and N5,N10-methylene-5, 6, 7, 8-tetrahydropteroyl-glutamate. One of the components, L-protein, is dihydrolipoamide dehydrogenase. These observations are based on the X-ray crystallographic analysis of two forms of the H-protein. The Hprotein has been crystallized and its structure determine to a 2.6 Å resolution (171). The 131-amino acid residues form seven -strands arranged into two antiparallel -sheets forming a sandwich structure. One -helix is observed at the C-terminal end. The lipoyl moiety points toward the H34 and D128 resiudes. Guest and colleagues (172) have engineered an E. coli that has PDH-complexes with lower numbers of lipoyl domains. They show that the organism has a reduced growth rate and yield. While not explaining why three lipoyl domains are in the E2 of E. coli, these results show that more than one is required for maximum growth efficiency. Properties of the complexes, the growth under limiting glucose conditions is a more drastic condition that shows a functional debilitation. Additional study (173) has revealed that although there are three lipoyls on E2p only one lipoyl domain is sufficient to give full catalytic activity in vitro. Only the outermost lipoyl domain must be lipoylated for activity. The presence of the other lipoyl domain provides a structural extension rather than extra catalytically active cofactors. Constructs with three lipoyl domains give maximum growth rate while only one domain is required to give the greatest in vitro specific activity. The E. coli lipoyl-ligase can lipoylate the H protein of the glycine decarboxylase complex when exogenous lipoic acid is supplied (174). Selenolipoic acid, a lipoic acid analog, where both S atoms are replaced by Se, can be attached to the H protein. The enzyme is 26 % as active as the S-containing one for glycine decarboxylation (175). In reactions where redox is required it is poorly active. The changed redox potential is responsible. There are redundant pathways for lipoyl ligation in E. coli (176). lipA encodes a ligase that attaches exogenously supplied lipoic acid. lipB is required for the functioning of a second ligase which uses the lipoyl group from the biosynthetic pathway. The lipoyltransferase has been cloned from bovine liver (177). It contains 373 amino acids, a mitochondrial leader and has a MW of 39,137 Da. The sequence is 35 % identical with that of E. coli lipoate-protein ligase A. Lipoyltransferases I and II from bovine liver mitochondria can lipoylate E2p and E2o effectively but E2b had a low rate of lipoylation (178). By in vitro mutagensis studies it was found that G-54 of E2o was important for the ligation reaction. The lipB gene, lipoyl-protein ligase, from yeast has 27 % sequence identity with the E. coli protein (179). The yeast has also been cloned by Marvin, Williams, and Cashmore (179a). The biotinylating and lipoylating enzymes are part of an evolutionarily related protein family that contains a homologous catalytic module (180). There is a single conserved lysine that is expected to contribute to the binding of lipoic acid in the LplA and LipB enzymes. There is no three-dimensional structure available for any of the liopoylating enzymes. The modeling has been done using the biotinylating enzyme BirA from E. coli. The human lipoyltransferase gene is located on chromosome band 2q11.2. It is 88 % identical to the bovine enzyme and 31 % identical to the E. coli one (180a). The biotinylating and lipoylating enzymes look at different keys in determining what is a proper substrate for posttranslational modification. The lipoylating enzyme, LplA, recognized the MKM sequence and responds to structural cues in the flanking -strand of the substrate protein (181). The surface antigentic protein P64K fromthe pathogenic Neisseria meningitidis is a chimera of the lipoyl domains of E2, the linker regions of E2, and E3. Lipoyl protein ligase attached a lipoyl group (181a). The enzyme lipoamidase hydrolytically removes the bound lipoic acid from the E2 and protein X units of the -keto acid dehydrogenases. The reaction is: Lipoyl-PDHC + H2O &emdash;&emdash;&emdash;&emdash;&emdash;&emdash;&emdash;&emdash; > apo-PDHC + Lipoic acid These two reactions may be another means of regulating the activity of the -activating system has been little studied since its original characterization in the 1950s and 1960s. There have been more studies on lipoamidase because of its usefulness in studying the complexes, but many questions remain. The molecular biology (base sequence, etc.) of the complexes is being elucidated. However, there have been no such studies on the lipoic acid-activating system and lipoamidase. Oizumi and Hayakawa (182) have purified lipoamidase 295-fold to near homogeneity from guinea pig liver. Lipoamidase was found in the microsomal membrane fraction and was labile. The subunit molecular mass was 68 kDa; under nondenaturing conditions the molecular mass was 120 kDa, suggesting a homodimer. Lipoamidase has a pI of 5.7, a pH optimum of 8.0, and is not inhibited by PCMB. It is inhibited by DFP and is thus a serine peptidase. Recent Results on the Regulatory Enzymes The catalytic subunit of bovine pyruvate dehydrogenase phosphatase has been cloned and sequenced (183). The protein has 467 amino acid residues with a Mr of 52,625. The activity of enzyme expressed in E. coli was near that of the native bovine PDP. The primary structure of the pyruvate dehydrogenase kinase has been determined and it establishes a new family of eukaryotic protein kinases (184). The cDNA codes for 434 amino-acid protein. T he protein lacks the motifs usually associated with ser/thr-protein kinases. There is homology with the prokaryotic histidine kinase. The lipoyl group is involved in the interaction with the kinase (185). Reed's group (186) has developed a one-step purification for the catalytic subunit of the recombinant bovine pyruvate dehydrogenase phosphatase. The PDPc binds to the inner lipoyl domain of the E2 of mammalian pyruvate dehydrogenase. To conserve carbohydrate reserves, the reaction of PDH-complex is down-regulated when the citric acid cycle is provided with sufficient acetyl~CoA. The E1 kinase activity is increase by increased NADH/NAD+ and acetyl~CoA/CoA ratios. Roche and associates (187) showed that the lipoyl moities on E2 were required for the control by the coenzymes. The control of the kinase is changed according to the molecular state of the inward lipoyl group: oxidized, reduced, or acetylated. There are at least three isoenzymatic forms of PDpK (PDpK1 and PDpK2) (188). cDNAs for these enzymes were cloned from human liver. The PDpK3 is almost exclusively expressed in heart and skeletal muscle suggesting a muscle-specialized function. PDpK2 has the highest expression in most tissues and is probably the one responsible for the majority of regulation. Although the rat branched-chain -ketoacid dehydrogenase kinase has sequence similarity to histidine-protein kinase, autophosphorylated kinase is not active (189). Using site-directed mutagenesis Korotchkina and Patel the three phosphorylation sites were changed from ser to ala to allow analysis of the phosphorylation reaction (190). The rates of phosphorylation and inactivation were site specific. Phosphorylation of each site resulted in complete inactivation of the E1p. The rates of dephosphorylation of the various sites were similar and there is a random dephosphorylation mechanism. The same authors ((190a) have reviewed the phosphorylation by four kinases on the three sites. There appears to be site specificity and independence. The roles of amino acids residues surrounding the phosphorylation site of BCKDH have been analyzed (191). Changing of arg-288, his-292, or asp-291 resulted in inactive enzymes. The his-292 and a S-293 mutation prevented TPP binding. The arg-288 was not phosphorylated, but all others were. Roche's group (192) has implicated tryptophan-135 in the TPP binding site of E1p. Two metal ions, Ca2+ and Mg2+, independently modulate -ketoglutarate dehydrogenase complex activity (193). There are tertiary and quaternary interactions that regulate the yeast 2-oxo acid decarboxylases using the substrate pyruvate and the cofactor TPP. These studies have been reviewed by Guest and colleagues (194). Roche and colleagues (195) have reviewed the regulation by the four pyruvate dehydrogenase kinases and two pyruvate dehydrogenase phosphatases. Tissue-specific and metabolic state-specific control is acheived by the selective expression and distinct regulatory properties of the enzymes. One or more of the lipoyl domains in E2 selectively bind each PDK and PDP. Web connections 1. Pyruvate dehydrogenase complex Complex http://www.biochemtech.unihalle.de/PPS2/course/section11/complexes.html Structure http://w3pharm.u-shizuokaken.ac.jp/~bioorg/macromol/pdh/pdh_complex.html Deficiency http://www.nlm.nih.gov/mesh/jablonski/syndromes/syndrome548.html Architecture http://www.bmsc.washington.edu/people/hol/WimFigs5.html & word description but must navigate back to section 5 from above. 2. NMR structure of the lipoyl domain http://coli.polytechnique.fr/NMR.html 3. Reactions of the complex http://fig.cox.miami.edu/Faculty/Tom/bil255/pyruvate.gif mechanism http://www.chem.umd.edu/biochem/jollie/462/enzymes/glycol/pyrdhm.htm 4. Pyruvate dehydrogenase E1 polypeptide 1, small deletions http://www.uwcm.ac.uk/uwcm/mg/ns/4/118895.html 5. Crystallography, sequence, and comparisons E1 http://heme.gsu.edu/glactone/PDB/Proteins/Krebs/1pyd.html E1 http://web1.ebc.uu.se/molev/publications/cfg2000/descrip tions/PDA1.html E1 http://web1.ebc.uu.se/molev/publications/cfg2000/de scriptions/PDB1.html E1o http://web1.ebc.uu.se/molev/publications/cfg2000/de scriptions/KGD1.html X http://web1.ebc.uu.se/molev/publications/cfg2000/de scriptions/PDX1.html E2 http://chemistry.gsu.edu/glactone/PDB/Proteins/Krebs/1iyu.html Various http://web1.ebc.uu.se/molev/publications/cfg2000/de scriptions/LAT1.html E2o http://web1.ebc.uu.se/molev/publications/cfg2000/de scriptions/KGD2.html E3 http://www.scripps.edu/pub/goodsell/interface/interface_images/1ebd.html Various listed http://web1.ebc.uu.se/molev/publications/cfg2000/de scriptions/LPD1.html 6. PDB files search for variations http://www.rcsb.org 7. Lipoic acid synthase http://web1.ebc.uu.se/molev/publications/cfg2000/descriptions/LIP5.html Literature Cited 1. Roche, T. E., &. Patel, M.S., (Eds.) (1989) -Keto Acid Dehydrogenase Complexes: Organization, Regulation, and Biomedical Aspects, Ann. NY Acad. Sci. 573, 474 pp. 2. Patel, M.S., & Roche, T.E. (1990) FASEB J., 4, 3224-3233. 3. Bisswanger, H., & Ullrich, J. (Eds.) (1991) Biochemistry & Physiology of Thiamin Diphosphate Enzymes, VCH, Weinheim, 453 pp. 4.Yeaman, S.J. (1989) Biochem. J., 257, 625-632. 5. Perham, R.N., Packman, L.C., & Radford, S.E. (1987) Biochem. Soc. Symp., 54, 67-81. 6. Reed, L.J., & Hackert, M.D. (1990) J. Biol. Chem., 265, 8971-8974. 7. Patel, M.S., & Harris, R.A.. (1995) FASEB J., 9, 1164-1172. 7a. Reed, L. J. (1998) Prot. Sci., 7, 220-224. 7b. Reed, L. J., (2001) J. Biol. Chem., 276, 38329-38336. 8. Reed, L. J., Gunsalus, I.C., Schnakenberg, G.H.F., Soper, Q.F., Boaz, H.E., Kern, S.F., & Parke, T.V. (1953) J. Am. Chem. Soc., 75, 1267-1270. 9. Reed, L. J., DeBusk, B.G., Hornberger, C.S., & Gunsalus, I.C. (1953) J. Am. Chem. Soc., 75, 1271-1273. 10. Hornberger, C. S., Heitmiller, R.F., Gunsalus, I.C., Schnakenberg, G.H.F., & Reed, L.J. (1953) J. Am. Chem. Soc., 75, 1273-1277. 11. Leach, F. R., Yasunobu, K., & Reed, L.J. (1955) Biochim. Biophys. Acta, 18, 297-298. 12. Reed, L. J., Leach, F.R., & Koike, M. (1958) J. Biol. Chem., 232, 123-142. 13. Reed, L. J. (1974) Acc. Chem. Res., 7, 10-16. 14. Wieland, O. H. (1983) Rev. Physiol. Biochem. Pharmacol., 96, 123-170. 15. Reed, L. J. & Yeaman., S.J. (1987) in The Enzymes (Boyer, P. D. & Krebs, E.G., Eds.) Vol. 18, Academic Press, New York, pp. 77-95. 16. Ord, M.G., & Stocken, L.A. (2000) TIBS, 25, 253-256. 17. Wada, H., Shintani, D., & Ohlrogge, J. (1997) Proc. Nat. Acad. Sci. USA, 94, 1591-1596. 18. Brody, S., Oh, C., Hoja, U., & Schweizer, E. (1997) FEBS Lett., 408, 217-220. 19. Jordan, S.W., & Cronan, Jr., J.E. (1997) J. Biol. Chem., 272, 17903-17906. 20. Self, W.T., Tsai, L., & Stadtman, T.C. (2000) Proc. Nat. Acad. Sci. U.S.A., 97, 12481-12486. 21. Reed, K.E., Morris, T.W., & Cronan, J.E. (1994) Proc. Nat. Acad. Sci.U.S.A., 91, 3720-3724. 22. Miller, J.R., Busby, R.W., Jordan, S.W., Cheek, J., Henshaw, T.F., Ashley, G.A., Broderick, J.B., Cronan, J.E., & Marletta, M.A. (2000) Biochemistry, 39, 15166-15178. 23. Choudens, S.O-d, Sanakis, Y., Hewitson, K.S., Roach, P., Baldwin, J.E., Munck, E., & Fontecave, M. (2000) Biochemistry, 29, 4165-4173. 24. Begley, T.P., Xi, J., Kinsland, C., Taylor, S., & McLafferty, F. (1999) Cur. Opin. Chem Biol., 3, 623629. 25. Stryer, L. (1995) Biochemistry , W.H. Freeman, New York. 26. Harper, A. E. (1989) in ref. 1. 27. Roche, T. E., Rahmatullah, M., Powers-Greenwood, S.L., Radke, G. A., Gopalakrishnan, S., & Chang, C.L. (1989) in ref. 1. 28. Stoops, J.K., Cheng, R.H., Yazdi, M.A., Maeng, C.-Y., Schroeter, J.P., Kueppleberg, U., Kolodziej, S.J., Baker, T.S., & Reed, L.J. (1997) J. Biol. Chem., 272, 5757-5764. 29. James, JA.G., Cook, R.M., West, S.M., & Lindsay, J.G. (1995) FEBS Lettr., 373, 111-114. 30. Behal, R. H., Buxton, D.B., Robertson, J.G., & Olson, M.S. (1993) Annu. Rev. Nutri., 13, 497-520. 31. Skinner, D.D., Morgenstern, M.R., Fedechko, R.W., & Denoya, C.D. (1995) J. Bacteriol., 177, 183190. 32. Zhu, P.-P., & Peterkofsky, A. (1996) Prot. Sci., 5, 1719-1736. 33. Koike, K. (1998) Biochim. Biophys. Acta, 1385, 373-384. 34. Neveling, U., Bringer-Meyer, S.,& Sahm, H. (1998) Biochim. Biophys. Acta, 1385, 367-372. 35. Frey, P. A. (1989) in ref. 1. 36. Stephens, P. E., Darlison, M.G., Lewis, H.M., & Guest, J.R. (1983) Eur. J. Biochem., 133, 155-162. 37. Guest, J. R., & Russell, G.C. (1989) in ref. 1. 38. Zhang, B., Kuntz, M.J., Goodwin, G.W., Harris, R.A., & Crabb, D.W. (1987) J. Biol. Chem., 262, 15220-15224. 39. Fisher, C.W., Chuang, J.L., Griffin, T.A., Lau, K.S., Cox, R.P., & Chuang, D.T. (1980) J. Biol. Chem., 264, 3448-3453. 40. Dahl, H. M., Hunt, S.M., Hutchison, W.M., & Brown, G.K. (1987) J. Biol. Chem., 262, 7398-7403. 41. De Meirleir, L., MacKay, N., Wah, A.M.L.H., & Robinson, B.H.(1988) J. Biol. Chem., 263, 19911995. 42. Ho, L., Javed, A.A., Pepin, R.A., Thekkumkara, T.J., Raefsky, C., Mole, J.E., Caliendo, A.M., Kwon, M.S., Kerr, D.S., & Patel, M.S. (1988) Biochem. Biophys. Res. Commun., 150, 904-908. 43. Hu, C.-W.C., Lau, K.S., Griffin, T.A., Chuang, J.L., Fisher, C.W., Cox, R.P., & Chuang, D.T. (1988) J. Biol. Chem., 263, 9007-9014. 44. Burns, G., Brown, T., Hatter, K., Idriss, J.M., & Sokatch, J.R. (1988) Eur. J. Biochem., 176, 311-317. 45. Darlison, M. G., Spencer, M.E., & Guest, J.R. (1984) Eur. J. Biochem., 141, 351-359. 46. Miran, S.G., Lawson, J.E., & Reed, L.J. (1993) Proc. Nat. Acad. Sci. USA, 90, 1252-1256. 47. Luethy, M.H., Miernyk, J.A., & Randall, D.D. (1995) Gene, 164, 251-254. 48. Tripatara, A., Korotchkina, L.G., & Patel, M.S. (1999) Arch. Biochem. Biophys., 367, 39-50. 49.Aevarsson, A., Seger, K., Turley, S., Sokatch, J.R., & Hol, W.G.J. (1999) Nature Struc. Biol., 6,785792. 50.Aevarsson, A., Chaung, J.L., Wynn, R.M., Turley, S., Chuang, D.T., & Hol, W.G.J. (2000) Structure, 8, 277-291. 51. Reed, L. J., Browning, K.S., Niu, X-a., Behal, R.H., & Uhlinger, D.J. (1989) in ref. 1. 52. Radford, S.E., Laue, E.D., Perham, R.N., Martin, S.R., & Appella, E. (1989) J. Biol. Chem., 264, 767776. 53. Perham, R. N. (1989) in ref. 1. 54. Spencer, M. E., Darlison, M.G., Stephens, P.E., Duckenfield, I.K., & Guest, J.R. (1984) Eur. J. Biochem., 141, 361-374. 55. Hanemaaijer, R., Janssen, A., de Kok, A., & Veeger, C. (1988) Eur. J. Biochem., 174, 593-599. 56. Lau, K. S., Griffin, T.A., Hu, C-W.C., & Chuang, D.T. (1988) Biochemistry, 27, 1972-1981. 57. Bradford, A. P., Aitken, A., Beg, F., Cook, K.G., & Yeaman, S.J. (1987) FEBS Lett., 222, 211-214. 58. Bradford, A. P., Howell, S., Aitken, A., James, L.S., & Yeaman, S.J. (1987) Biochem. J., 245, 919-922. 59. Fujiwara, K., Okamura-Ikeda, K., & Motokawa, Y. (1986) J. Biol. Chem., 261, 8836-8841. 60. Berg, A., de Kok, A., & Vervoort, J. (1994) Eur. J. Biochem., 221, 87-100. 61. Berg, A., Smits, O., de Kok, A., & Vervoort, J. (1995) Eur. J. Biochem., 234, 148-159. 62. Liu, S., Baker, J.C., & Roche, T.E. (1995) J. Biol. Chem., 270, 793-800. 63. Mande, S.S., Sarfaty, S., Allen, M.D., Perham, R.N., & Hol, W.G.J. (1996) Structure, 4, 277-286. 64. Berg, A., & de Kok, A. (1997), Hoppe-Syler Biol. Chem., 378, 617-634. 65. Jones, D.D., Stott, K.M., Reche, P.A., & Perham, R.N. (2000) J. Mol. Biol., 305, 49-60. 66. Gong, X., Peng, T., Yakhnin, A., Zolkiewski, M., Quinn, J., Yeaman, S.J., & Roche, T.E. (2000) J. Biol. Chem., 275, 13645-13653. 67. Jones, D.D., Stott, K.M., Howard, M.J., & Perham, R.N. (2000) Biochemistry 39, 8448-8459. 68. Thelen, J.J., Muszyncki, M.G., David, N.R., Luethy, M/H/. Elthon, T.E., Miernyk, J.A., & Randall, D.D., (1999) J. Biol. Chem., 274, 21769-21775. 69. Stephens, P. E., Darlison, M.G., Lewis, H.M., & Guest, J.R. (1983) Eur. J. Biochem., 133, 481-489. 70. Spencer, M. E., Darlison, M.G., Stephens, P.E., Duckenfield, I.K., & Guest, J.R. (1984) Eur. J. Biochem., 141, 361-374. 71. Nobukuni, Y., Mitsubuchi, H., Endo, F., & Matsuda, I. (1989) Biochem. Biophys. Res. Commun., 161, 1035-1041. 72. Thekkumkara, T.J., Ho, L., Wexler, I.D., Pons, G., Liu, T.-C., & Patel, M.S. (1988) FEBS Lett., 240, 45-48. 73. Burns, G, Brown, T. Hatte, K., & Sokatch, J.R. (1988) Eur. J. Biochem., 176, 165-169. 74. Packman, L.C., Borges, A., & Perham, R.N. (1988) Biochem. J., 252, 79-86. 75. Griffin, T.A., Lau, K.S, & Chuang, D.T. (1988) J. Biol. Chem., 263, 14008-14014. 76. Mattevi, A., Obmolova, G., Schylze, E., Kalk, K.H., Westphal, A.H., de Kok, A., & Hol, W.G.J. (1992) Science, 255, 1544-1550. 77. Mattevi, A., Obmolova, G. Kalk, K.H., Tepyakov, A., & Hol, W.G.J. (1993) Biochemistry, 32, 38873901. 78. Meng, M., & Chuang, D.T. (1994) Biochemistry 33, 12879-12885. 79. Berg, A., Vervoot, J., & de Kok, A. (1996) J. Mol.Biol., 261, 432-442. 80. Wallis, N.G., Allen, M.D., Broadhurst, R.W., Lessard, I.A.D., & Perham, R.N. (1996) J. Mol. Biol., 263, 463-474. 81. Ricaud, P.M., Howard, M.J., Roberts, E.L., Broadhurst, R.W., & Perham, R.N. (1996) J. Mol. Biol., 264, 179-190. 82. Knapp, J.E., Carroll, D., Lawson, J.E., Ernst, S.R., Reed, L.J., & Hackert, M.L. (2000) Prot. Sci., 9, 3748. 83. Hodgson, J. A., de Marcucci, O.G., & Lindsay, J.G. (1986) Eur. J. Biochem., 158, 595-600. 84. Jilka, J. M., Rahmatullah, M., Kazemi, M., & Roche, T.E. (1986) J. Biol. Chem., 261, 1858-1867. 85. Rahmatullah, M., Gopalakrishnan, S., Andrews, P.C., Chang, C.L., Radke, G.A., & Roche, T.E. (1989) J. Biol. Chem., 264, 2221-2227. 86. Hodgson, J. A., de Marcucci, O.G.L., & Lindsay, J.G. (1988) Eur. J. Biochem., 171, 609-614. 87. Rahmatullah, M., Gopalakrishnan, S., Radke, G.A., & Roche, T.E. (1989) J. Biol. Chem., 264, 12451251. 88. Sanderson, S.J., Kahn, S.S., McCartney, R.G., Miller, C., & Lindsay, J.G. (1996) Biochem. J., 319, 109-116. 89. Maeng, C.-Y., Yazdi, M.A., & Reed, L.J. (1996) Biochemistry, 35, 5879-5882. 90. Klingbeil, M.M., Walker, D.J., Arnette, R., Sidawy, E., Hayton, K., Komuniecki, P.R., & Komuniecki, R. (1996) J. Biol. Chem., 271, 5451-5457. 91. Sanderson, S.J., Miller, C., & Lindsay, J.G. (1996) Eur. J. Biochem., 236, 68-77. 92. Stephens, P. E., Lewis, H.M, Darlison, M.G., & Guest, J.R (1983) Eur. J. Biochem., 135, 519-527. 93. Rice, D. W., Schulz, G.E., & Guest, J.R. (1984) J. Mol. Biol., 174, 483-496. 94. Karplus, P. A., & Schulz, G.E. (1987) J. Mol. Biol., 195, 701-729. 95. Otulakowski, G., & Robinson, B.H. (1988) J. Biol. Chem., 262, 17313-17318. 96. Browning, K. S., Uhlinger, D.J., & Reed, L.J. (1988) Proc. Nat. Acad. Sci. U.S.A., 85, 1831-1834. 97. Ross, J., Reid, G.A., & Dawes, I.W. (1988) J. Gen. Microbiol., 134, 1131-1139. 98. Westphal, A. H., & de Kok, A. (1988) Eur. J. Biochem., 172, 299-305. 99. Owen, P., Kaback, H.R., & Graeme-Cook, K.A. (1980) FEMS Microbiol. Lett., 7, 345-348. 100. Richarme, G. (1985) J. Bacteriol., 162, 286-293 101. Danson, M. J. (1988) Biochem. Soc. Trans., 16, 87-89. 102. Carothers, D.J., Pons, G., & Patel, M.S. (1989) Arch. Biochem. Biophys., 268, 409-425. 103. Bringas, R., & Fernandez, J. (1996) Proteins, 21, 303-306. 104. Prehn, J.H.M., Karkoutly, C., Nuglisch, J., Peruche, B., & Krieglstein, J., J. Cere. Blood Flow Metab., 12, 78-87 (1992). 105. Mizuno, M., & Packer, L., Biochem. Biophys. Res. Commun., 200, 1136-1142 (1994). 106. Scheer, B., & Zimmer, G., Arch. Biochem. Biophys., 302, 385-390 (1993). 107. Devasagayam, T.P.A., Subbaranian, M., Pradhan, D., & Sies, H., Chem.-Biol. Interactions, 86, 79-92 (1993). 108. Podda, M., Tritschler, H.J., Ulrich, H., & Packer, L., Biochem. Biophys. Res. Commun., 204, 98-104 (1994). 109. Kawabata, T., & Packer, L., Biochem. Biophys. Res. Commuin., 203, 99-104 (1994). 110. Stoll, S., Hartmann, H., Cohen, S.A., & Muller, W.E., Pharnacol. Biochem. Behavior, 46, 799-805 (1993). 111. Zimmer, G., Beikler, T.-K., Schneider, M., Ibel, J., Tritschler, H., & Ulrich, H. (1995) J. Mol. Cell. Cardiol., 27, 1895-1903. 112. Cao, X., & Phillis, J. W. (1995) Free Rad. Res., 23, 365-370. 113. Panigrahi, M., Sadguna, Y., Shivakumar, B.R., Kolluri, S.V.R., Roy, S., Packer, L., & Ravindranath, V. (1996) Brain Res., 717, 184-188. 114. Wolz, P., & Krieglstein, J. (1996) Neuropharmacology, 35, 369-375. 115. Jacob, S., Streeper, R.S., Fogt, D.L., Hokama, J.Y., Tritschler, H.J., Dietze, G.J., & Henriksen, E.J. (1996) Diabetes, 45, 1024- 1029. 116. Ziegler, D., Schatz, H., Conrad, F., Gries, F.A., Ulrich, H., & Reichel, G. (1997) Diabet. Care, 20, 369-373. 117. Packer, L., & Cadenas, E., eds. (1995) Biothiols in Health and Disease, Dekker, New York, 520+ pp. 117a. Hengeveld, A.F., & de Kok, A. (2002) Curr. Med. Chem., 9, 499-520. 118. Bustamante, J., Lodge, J.K., Marcocci, L., Tritschler, H.J., Packer, L., & Rihn, B.H. (1998) Free Rad. Biol. Med., 24, 1023-1039. 119. Wagenmakers, A.J., Veerkamp, J.H., Schepens, J.T., & van Moerkerk, H.T., Effect of Clofibrate on Branched-Chain Amino Acid Metabolism, Biochem. Pharmacol., 34, 2169-2173 (1985). 120. Ono, K., Shioya, H., Hakozaki, M., Honda, K., Mori, T., & Kochi, H., Regulation by Induction of Branched-Chain 2-Oxo Acid Dehydrogenase Complex in Clofibrate-Fed Rat Liver, Biochem. Biohys. Res. Commun., 172, 243-248 (1990). 121. Honda, K., Ono, K., Mori, T., & Kochi, H., Both Induction and Activation of the Brached-Chain 2Oxo Dehydrogenase Complex in Primary-Cultures Rat Hepatocytes by Clofibrate, J. Biochem., 1098, 822897 (1991). 122. Paul, H.S., Sekas, G., & Adibi, S.A., Investigation of the Presence of Branched-Chain -Keto Acid Dehydrogenase in Mammalian Peroxisomes, Int. J. Biochem., 24, 617-629 (1992). 123. Harris, R.A., Popov, K.M., Shimomura, Y., Zhao, Y., Jaskiewicz, J., Nanaumi, N., & Suzuki, M., Purification, Characterization, Regulation, and Molecular Cloning of Mitochondrial Protein Kinase, Adv. Enz. Reg., 32, 267-284 (1992). 124. Robinson, B.H. (1993) Biochim. Biophys. Acta, 1182, 231-244. 125. Matalon, R., Stumpf, D.A., Michals, K., Hart, R.D., Parks, J.K., & Goodman, S.I. (1984) J. Pediat., 104, 65-69. 126. Wickramasinghe, S.N., & Hasan, R., In Vitro Effects of Vitamin C, Thioctic Acid and Dihydrolipoic Acid on the Cytotoxicity of Post-Ethanol Serum, Biochem. Pharmacol., 43, 407-4111 (1992). 127. Blass, J. P. (1985) in The Metabolic Basis of Inherited Diseases (Stanbury, J. B., Wyngaarden, J.B., Fredrickson, D.S., Goldstein, J.L., & Brown, M.S., Eds.) McGraw Hill, New York, p. 193-203. 128. Kerr, D. S., Berry, S.A., Lusk, M.M., Ho, L., & Patel, M.S. (1988) Pediat. Res., 24, 95-100. 129. Wexler, I.D., Kerr, D.S., Ho, L., Lusk, M.M., Pepin, R.A., Javed, A.A., Mole, J.E., Jesse, B.W., Thekkumkara, T.J., Pons, G., & Patel, M.S. (1988) Proc. Nat. Acad. Sci. U.S.A., 85, 7336-6340. 130. Endo, H., Hasegawa, K., Narisawa, K., Tada, K., Kagawa, Y., & Ohta, S. (1989) Am. J. Hum. Genet., 44, 358-364. 131. Chun, K., MacKay, N., Petrova-Benedict, R., Federico, A., Fois, A., Cole, D.E.C., Robertson, E., & Robinson, B.H. (1995) Am. J. Hum. Genet., 56, 558-569. 132. Takakubo, F., Cartwright, P., Hoogenraad, N., Thorburn, D.R., Collins, F., Lithgow, T., & Dahl, H.-H. M. (1995) Am. J. Hum. Genet., 57, 772-780. 133. Dahl, H.-H. M. (1995) Am. J. Hum. Genet., 56, 553-557. 134. Geoffroy, V., Fouque, F., Benelli, C., Poggi, F., Saudubray, J.M., Lissens, W., Meirleir, L.D., Marsac, C., Lindsay, J.G., & Sanderson, S.J. (1996) Pediatics, 267-272. 135. Pastoris, O., Savasta, S., Foppa, P., Catapano, M., & Dossena, M. (1996) Acta Paediatr., 85, 625-628. 136.Wu, Y.-G., Widjaja, S.L., Huang, C.-Y., Li, W., Nixon, P.F., & Duggleby, R.G. (2001) Molec. Genet. Metab., 72, 269-272. 136a. Luft, F.C. (2001) J. Am. Soc. Nephrol., Suppl. 17, S15-S19. 137. Tanka, K., & Rosen, L.E. (1985 ) in The Metaboic Basis of Inherited Disease (Stanbury, J. B., Wyngaarden, J.B., Fredrickson, D.S., Goldstein, J.L., and Brown, M.S., Eds.) McGraw Hill, New York, p. 440-473. 138. Danner, D.J., Armstrong, N., Heffelfinger, S.C., Seweh, E.T., Priest, J.H., & Elsas, L.J. (1985) J. Clin. Invest., 75, 858-860. 139. Indo, Y., Kitano, A., Endo, F., Akaboshi, I., & Matsuda, I. (1987) J. Clin. Invest., 80, 63-70. 140. Fisher, C.W., Chuang, J.L., Griffin, T.A., Lau, K.S., Cox, R.P., & Chuang, D. (1989) J. Biol. Chem., 264, 3448-3453. 141. Peinemann, F., & Danner, D.J. (1994) J. Inher. Metab. Dis., 17, 3-15. 142. Kobayshi, R., Shimomura, Y., Murakami, T., Nakai, N., Fujitsuka, N., Otsuka, M., Arakawa, N., Popov, K.M., & Harris, R.A. (1997) Biochem. J., 327, 449-453. 143. http://www.bioscience.org/1998/v3/d/danner/d517-524.htm 144. Sitler, T.L., McKean, M.C., Peinemann, F., Jackson, E., & Danner, D.J. (1998) Biochim. Biophys. Acta, 1404, 385-392. 144a. Dursun, A., Henneke, M., Ozgul, K., Gartner, J., Cooskun, T., Tokatli, A., Kalkanoglu, H.S., Demirkol, M., Wendel, U., & Ozalp, I. (2002) J. Inherit. Metab. Dis., 25, 89-97. 144b. Morton, D.H., Strauss, K.A., Robinson, D.L., Puffenberger, E.G., & Kelley, R.I. (2002) Pediatrics, 109, 999-1008. 145. Kerr, D. S., Ho, L., Berlin, C.M., Lanoue, K.F., Towfight, J., Hoppel, C.L., Lusk, M.M., Gondek, C.M., & Patel, M.S. (1987) Pediat. Res., 22, 312-318. 146. Kretzschmar, H. A., DeArmond, S.J., Koch, T. K., Patel, M.S., Newth, C.J.L., Schmidt, K.A., & Packman, S. (1987) Pediatrics, 79, 370-373. 147. Kinman, L.M., Sheu, K.-F.R., Baker, A.C., Kim, Y.T., & Blass, J.P. (1989) Neurology, 39, 70-75. 148. Marsac, C., Stansble, D., Bonne, G., Cousin, J., Jehenson, P., Benelli, C., Lerous, J.-P., & Lindsay, G. (1993) J. Pediatr., 123, 915-920. 149. Gibson, G. E., Sheu, K-F., Blass, J.P., Baker, A., Carlson, K.C., Harding, B., & Perrino, P. (1988) Arch. Neurol., 45, 836-840. 150. Sheu, K. R., Kim, Y.T., Blass, J.P., & others (1985) Ann. Neurol., 17, 444-449. 151. Hoyer, S., Oesterreich, K., & Wagner, D. (1988) J. Neurol., 235, 143-148. 152. Gibson, G.E., Sheu, K.F., & Blass, J.P. (1998) J. Neural Transm., 105, 855-870. 152a. Heininger, K. (2000) Rev. Neurosci., 11, Spec No:23-328. 153. Van de Water, J., Fregeau, D., Davis, P., Ansari, A., Danner, D., Leung, P., Coppel, R., & Gershwin, M.E. (1988) J. Immunol., 141, 2321-2324. 154. Yeaman, S. J., Danner, D.J., Mutimer, D.J., Fussey, S.P.M., James, O.F.W., & Bassendine, M.F. (1988) Lancet, 1067-1070. 155. Van de Water, J., Gershwin, M.E., Leung, P., Ansari, A., & Coppel, R.L. (1988) J. Exp. Med., 167, 1791-1799. 156. Yeaman, S. J., Bassendine, M.F., James, O.F.W., & Fussey, S.P.M. (1989) in ref. 1. 156a. Fussey, S.P., Guest, J.T., James, O.F., Bassendine, M.F., & Yeaman, S.J. (1988) Proc. Nat. Acad. Sci. USA, 85, 8654-8658. 157. Hale, G., Wallis, N.G., & Perham, R.N. (1992) Proc. Roy. Soc. London B, 248, 247-253. 158. Tuaillon, N., Andre, C., Briand, J.-P., Penner, E., & Muller, S. (1992) J. Immunol., 148, 445-450. 159. Elkhalifa, M.Y., Kiechle, F.L., Gordon, S.C., Chen, J., & Poulik, M.D. (1992) Am. J. Clin. Pathol., 97, 202-208. 160. Quinn, J., Diamond, A.G., Palmer, J.M., Bassendine, M.F., James, O.F.W., & Yeaman, S.J. (1993) Hepatology, 18, 1384-1391. 161. McNally, A.J., Motter, K., & Jordan, F. (1995) J. Biol. Chem., 270, 19736-19743. 162. McNally, A.J., Mattsson, L., & Jordan, F. (1995) J. Biol. Chem., 270, 19744-19751 163. Teoh, K.-L., Rowley, M.J., Zafirakis, H., Dickson, E.R., Wiesner, R.H., Gershwin, M.E., & Mackay, I.R. (1994) Hepatology, 20, 1220-1224. 164. Frey, N., Christen, U., Jeno, P., Yeaman, S.J., Shimomura, Y., Kenna, J.G., Gandolfi, A.J., Ranek, L., & Gut, J. (1995) Chem. Res. Toxicol., 8, 736-746. 164a. Mabee, C.L., & Thiele, D.L. (2000) Clin. Liver Dis., 4, 431-445. 164b. Sutton, I., & Neuberger, J. (2002) Gut, 50, 743-746. 164c. Nishio, A., Keeffe, E.B., & Gershwin, M.E. (2001) Clin. Exp. Med., 1, 165-178. 165. Fujimoto, M., Sato, S., Ihn, H., Kikuchi, K., Tamaki, K., & Takehara, K. (1995) Arthr. Rheum., 38, 985-989. 166. Yang, Y.-S., & Frey, P.A. (1989) Arch. Biochem. Biophys., 268, 465-474. 167. Morris, T.W., Reed, K.E., & Cronan, Jr., J.E. (1994) J. Biol. Chem., 269, 16092-16100. 168. Fujiwara, K., Okamura-Ikeda, K., & Motokawa, Y. (1994) J. Biol. Chem., 269, 16605-16609. 168a. Fujiwara, K., Takeuchi, S., Okamura-Ikeda, K., & Motokawa, Y. (2001) J. Biol. Chem., 276, 2881928823. 169. Green, D.E., Morris, T.W., Green, J., Cronan, Jr., J.E., & Guest, J.R. (1995) Biochem. J., 309, 853862. 170. Cohen-Addad, C., Pares, S., Sieker, L., Neuburger, M., & Douce, R. (1995) Struct. Biol., 2, 63-68. 171. Pares, S., Cohen-Addad, C., Sieker, L., Neuburger, M., & Douce, R. (1994) Proc. Nat. Acad. Sci. USA, 91, 4850-4853. 172. Dave, E., Guest, J.R., & Attwood, M.M. (1995) Microbiology, 141, 1839-1849. 173. Guest, J.R., Attwood, M.M., Machado, R.S., Matqi, K.Y., Shaw, J.E., & Turner, S.L. (1997) Microbiology, 143, 457-466. 174. Macherel, D., Bourguignon, J., Forest, E., Faure, M., Cohen-Addad, C., & Douce, R. (1996) Eur. J. Biochem., 236, 27-33. 175. Fujiwara, K., Okamura-Ikeda, K., Packer, L., & Motokawa, Y. (1997) J. Biol. Chem., 272, 1988019883. 176. Morris, T.W., Reed, K.E., & Cronan, J.E. (1995) J. Bacteriol., 177, 1-10. 177. Fujiwara, K., Okamura-Ikeda, K., & Motokawa, Y. (1997) J. Biol. Chem., 272, 31974-31978. 178. Fujiwara, K., Okamura-Ikeda, K., & Motokawa, Y. (1996) J. Biol. Chem., 271, 12932-12936. 179. Chen, X.J. (1997) Mol. Gen. Genet., 255, 341-349. 179a. Marvin, M.E., Williams, P.H., & Cashmore, A.M., FEMS Microbiol. Lett., 199, 131-136. 180. Reche, P.A. (2000) Prot. Sci., 9, 1922-1929. 180a. Fujiwara, K. Suzuki, M., Okumachi, Y., Okamura-Ikeda, K., Fujiwara, T., Takahashi, E., & Motokawa, Y. (1999) Eur. J. Biochem., 260, 761-767. 181. Reche, P., & Perham, R.N. (1999) EMBO J., 18, 2673-2682. 181a. Tozawa, K., Broadhurst, R.W., Raine, A.R., Fuller, C., Alvarez, A., Guillien, G., Pardon, G., & Perham, R.N. (2001) Eur. J. Biochem., 268, 4908-4917. 182. Oizumi, J., & Hayakawa, K. (1989) Biochim. Biophys. Acta, 991, 410-414. 183. Lawson, J.E., Niu, X.-D., Browning, K.S., Trong, H.L., Yan, J., & Reed, L.J., Biochemistry, 32, 89878993 (1993). 184. Popov, K.M., Kedishvilli, N.Y., Zaho, Y., Shimomuaru, Y., Crabb, D.W., & Harris, R.A., J. Biol. Chem., 268, 26602-26606 (1993). 185. Ono, K., Radke, G.A., Roche, T.E., & Rahmatullah, M., J. Biol. Chem., 268, 26135-26143 (1993). 186. Choi, W.S., Yan, J., McCarthy, D.B., Park, S.H., & Reed, L.J. (2000) Prot. Exp. Puri., 20, 128-131. 187. Ravindran, S., Radke, G.A., Guest, J.R., & Roche, T.E. (1995) J. Biol. Chem., 271, 653-662. 188 Gudi, R., Bowker-Kinley, M., Kedishvili, N.Y., Zhao, Y., & Popov, K.M. (1995) J. Biol. Chem., 270, 28989-28994. 189. Davie, J.R., Wynn, R.M., Meng, M., & Huang, Y.-S. (1995) J. Biol. Chem., 270, 19861-19867. 190. Korotchina, L.G., & Patel, M.S. (1995) J. Biol. Chem., 270, 14297-14304. 190a. Patel, M.S., & Korotchina, L.G. (2001) Exp. Mol. Med., 33, 191-197. 191. Ali, M.S., Shenoy, B.C., Eswarant, D., Amdersson, L.A., Roche, T.E., & Patel, M.S. (1995) J. Biol. Chem., 270, 4570-4574. 192. Panov, A., & Scarpa, A. (1996) Biochemistry, 35, 427-432.95. Simonot, C., Lerme, F., Louisot, P., & Gateau-Roesch, O. (1997) FEBS Lett., 401, 158-162. 193. Simonot, C., Lerme, F., Louisot, P., & Gateau-Roesch, O. (1997) FEBS Lett., 401, 158-162. 194. Jordan, F., Nemeria, N., Guo, F., Baburina, I., Gao, Y., Kahyaoglu, A., Li, H., Wang, J., Yi, J., Guest, J.R., & Furey, W. (1998) Biochim. Biophys. Acta, 1385, 287-306. 195. Roche, T.E., Baker, J.C., Yan, X., Hiromass, Y., Gong, X., Peng, T., Dong, J., Turkan, A., & Kasten, S.A. (2001) Prog. Nucleic Acid Res. Mol. Biol., 70, 33-75. Appendix Lester James Reed A Tribute to Lester J. Reed From Ref (1). This conference and volume [Roche, T.E. & Patel, M.S. (eds) (1989) -Keto Acid Dehydrogenase Complexes: Organization, Regulation, and Biomedical Aspects, Ann. NY Acad. Sci. 573, 474 pp.] are a tribute to Dr. Lester J. Reed for his many outstanding contributions to the field of -keto acid dehydrogenase complexes. Lester gouged out of the forest of biochemical distractions a highway that runs from lipoic acid chemistry to exquisite information on the structure, function, and regulation of -keto acid dehydrogenase complexes. We wish to pay tribute to Lester for his dedication in this brilliant effort and for his integrity and quiet leadership, which have made lasting impressions on those who have had the privilege to be associated with him. A Brief Biography Lester J. Reed was born on January 3, 1925, in New Orleans, Louisiana. He received his B.S. degree at Tulane University in 1943 and completed his Ph.D. under Reynold C. Fuson at the University of Illinois in 1946. Having achieved the latter degree at the ripe old age of 21, he took a position as a postdoctoral research associate with Vincent duVigneaud at Cornell University Medical College from 1946 to 1948. In 1948, he joined the University of Texas at Austin, where he has been a professor since 1958, director of the Clayton Foundation Biochemical Institute since 1963, and Ashbel Smith Professor since 1984. Affiliations and Honors Professor Reed is a member of several professional societies, has served on many advisory councils and editorial boards, and has been the recipient of several honors. Memberships. Phi Beta Kappa, Sigma Xi, American Chemical Society, American Society for Biochemistry and Molecular Biology, the Protein Society, American Association for the Advancement of Science (Fellow), National Academy of Sciences, American Academy of Arts and Sciences. Advisory Councils and Editorial Boards. Biochemistry Study Section of the National Institutes of Health; Editorial Board, Archives of Biochemistry and Biophysics; Nominating Committee and Executive Committee, Division of Biological Chemistry, American Chemical Society; Membership Committee and Nominating Committee, American Society of Biological Chemists; Editorial Board, Journal of Biological Chemistry; Editorial Board, Biofactors; U.S. National Committee for the International Union of Biochemistry. Honors. Eli Lilly and Co. Award in Biological Chemistry (American Chemical Society), 1958; election to the National Academy of Sciences, U.S.A., 1973; Honorary Doctor of Science Degree, Tulane University, 1977; election to the American Academy of Arts and Sciences, 1981; Ashbel Smith Professor, University of Texas, 1984. Major Contributions to Research on the Structure, Function, and Regulation of KetoAcid Dehydrogenase Complexes Lester joined the University of Texas at Austin, following his productive studies in synthetic organic chemistry with Fuson and in intermediary metabolism with duVigneaud. In 1949, the pioneering studies of Roger Williams in characterizing B vitamins were being extended in Williams's laboratory to characterizing the "acetatereplacing factor." The latter studies were initiated by Esmond Snell at the University of Wisconsin and continued in Texas after Snell moved to Austin. Dr. Williams invited Lester to undertake the characterization of this factor. The following is a selected list of landmark contributions of Lester Reed to the field of keto acid dehydrogenase complexes: Isolation and characterization of lipoic acid Identification of the functional form of lipoic acid Resolution of Escherichia coli pyruvate and -ketoglutarate dehydrogenase complexes and characterization of the components Utilization of electron microscopy to analyze the structural organization of the bacterial and mammalian complexes Regulation of the mammalian pyruvate dehydrogenase complex by phosphorylation dephosphorylation Isolation and characterization of the pyruvate dehydrogenase kinase and pyruvate dehydrogenase phosphatase Identification of inner and outer domain structure of the transacylase components Isolation and characterization of the branched-chain -keto acid dehydrogenase phosphatase and its inhibitor protein While establishing the fundamental organization of these quintessential multienzyme complexes, Lester also had a seminal role in formulating concepts concerning the unique properties attendant to the organization of enzymes in a clustered state. The following listing of selected major contributions (arbitrarily divided as convenient) gives a rough time-frame for the construction of the Lester J. Reed "highway," as well as a road map for traveling the route. To be honest, my intention was to make a shorter list, but I could not delete from, but only expand on, these significant discoveries. Here, then, is a selected list of Lester's major contributions. 1949-l954 Isolation, characterization, and synthesis of -lipoic acid: structural and physical characterization, broad distribution and enrichment in mitochondria, high-yield synthesis, 35 S-labeled cofactor, analogs. 19S4-l9S8 Functional form of lipoic acid: attachment to -amino group of lysine, ATP requiring (lipoyl-adenylate intermediate) reaction of lipoic acid-activating enzyme, lipoyl-X hydrolase reaction. 1957-1963 Purification of E. coli pyruvate and -ketoglutarate dehydrogenase complexes, flavoprotein nature of the dihydrolipoyl dehydrogenase component, resolution and reconstitution of the E. coli pyruvate debydrogenase complex, characterization of component enzymes, sequence around the -amino lipoyllysine group, purification of lipoamidase. model reactions, formation of 2-acetyl-thiamin pyrophosphate (AcTPP), other aspects of reaction mechanism of components. 1964-1968 Electron microscopic characterization of complexes, lipoyllysine swinging arm activesite coupling mechanism, transacylase cores composed of 24 E2 subunits arranged with 432 symmetry in cube-like particle, location of the El component on the edges and E3 component on the faces of the cubic core, resolution and reconstitution of the E. coli ketoglutarate dehydrogenase complex, presence of the same E3 component in the pyruvate and the -ketoglutarate dehydrogenase complexes, purification and molecular organization of the pyruvate and -ketoglutarate dehydrogenase complexes from bovine kidney, development of concepts concerning unique properties of enzymes that are organized into complexes. 1968-1970 Regulation of the E. coli pyruvate dehydrogenase complex by phosphoenolpyruvate, acetyl-CoA, and guanine nucleotides; regulation of bovine kidney and heart and porcine liver pyruvate dehydrogenase complex by phosphorylation and dephosphorylation, ATPMg2+ dependent PDH kinase tightly associated with the complex and inhibited by ADP and pyruvate, Mg2+-dependent PDH phosphatase weakly associated with the complex. 1971-1974 X-ray crystallography of inner core of E. coli dihydrolipoyl transsuccinylase establishing octahedral (432) symmetry; preparation of the component enzymes of the pyruvate dehydrogenase complexes from bovine kidney and heart, and characterization of their physical and chemical properties; stoichiometry of subunits in the mammalian complexes; subunit ratios based on sedimentation equilibrium molecular weights of purified components of E. coli pyruvate and -ketoglutarate dehydrogenase complexes; multiple phosphorylation sites in mammalian pyruvate dehydrogenase (PDH) and sequence around phosphorylation sites; characterization of PDH kinase: separation of the kinase from the transaeetylase, direct pyruvate inhibition, thiamin PP inhibition, transaeetylase effect on Vmax and Km for PDHa, and monovalent cation effects on ADP inhibition; role of Ca2+ in activating PDH phosphatase by increasing its association with the transacetylase and lowering its Km for PDH; first studies on modulation of steadystate phosphorylation and dephosphorylation; kinetic data supporting multisite ping-pong mechanism of kidney pyruvate dehydrogenase complex. 1975-1979 Kinetic mechanism of bovine kidney transacetylase, data supporting acetyl and electronpair relay system between lipoyl moieties, purification and properties of bovine kidney branched-chain -keto acid dehydrogenase complex, acetyl-CoA/CoA and NADH/NAD+ effects on PDH kinase and PDH phosphatase activities, kinetic properties determined with peptide substrates of PDH kinase 1982-1987 Two-domain structure of the transacetylase from E. coli and mammalian sources, transacetylation and E2-subunit binding role of inner domain; extended lipoyl-bearing outer domain, capacity of transacetylase domains to bind other components, contribution of structure in the lipoyl domain to the reductive acetylation reaction catalyzed by the E I component; X-ray crystallography of the inner domain of the E. coli transacetylase, establishing 432 symmetry; regulatory properties of the kinase and the phosphatase, utilizing peptide substrates. 1982-1987 Purification and properties of: the pyruvate dehydrogenase phosphatase (FAD-containing 90-kDa regulatory subunit, stimulation by polyamines); the pyruvate dehydrogenase kinase; the branched-chain -keto acid dehydrogenase phosphatase and a potent protein inhibitor of this phosphatase; and a distinct cation-independent, spermine-stimulated mitochondrial phosphatase. Computer model analysis supporting a multiple random coupling mechanism for active-site coupling through the lipoyl. domains of the E. coli pyruvate and -ketoglutarate dehydrogenase complexes. 1986-1988 Purification of bakers' yeast pyruvate dehydrogenase complex and phosphorylationdephosphorylation by mammalian kinase and phosphatase; cloning of genes of components of the yeast pyruvate dehydrogenase complex. It should be readily apparent from these latest contributions that Lester is still building the highway and opening new frontiers for further research. During preparations for this conference, Lester gave me a list of 60 students and collaborators who have contributed to the above work. For readers wanting more details about any of this work, I would note that, in addition to his numerous research papers, Lester has written 38 review articles which describe with great clarity the above work and related studies by other researchers. Thomas E. Roche Kansas State University Manhattan, Kansas October 1988 RECOLLECTIONS From lipoic acid to multi-enzyme complexes Lester J. Reed 1, 2 1 Department of Chemistry and Biochemistry, The University of Texas at Austin, Austin, Texas 78712 Adapted from Protein Science (1998) 7: 220-224 I shall retrace the high points of a trail of research that I have had the pleasure of establishing in association with many collaborators. This trail has led from the isolation and identification of a microbial growth factor to the structure, function, and regulation of -keto acid dehydrogenase complexes. The high points of this trail in the 1950s were the isolation, characterization, and synthesis of lipoic acid and identification of its functional form. In the late 1950s and into the 1960s the trail led to the isolation, resolution, and reconstitution of the Escherichia coli pyruvate and -ketoglutarate dehydrogenase complexes, characterization of their component enzymes, and elucidation of their macromolecular organization. In the late 1960s part of our research effort was directed toward isolating and characterizing the bovine pyruvate dehydrogenase (PDH) complex. In the early stage of this investigation we found that the complex is regulated by phosphorylation and dephosphorylation. Resolution of the mammalian PDH complex and characterization of its component enzymes, including the kinase and the phosphatase, continued in the 1970s and early 1980s. We also obtained evidence that the dihydrolipoamide acetyltransferase components of the E. coli and bovine PDH complexes possess a multi-domain structure. In the 1980s we isolated and characterized the bovine branched-chain -keto acid dehydrogenase complex and the phosphatase that regulates its activity. In the late 1980s and early 1990s we cloned and disrupted the genes encoding the components of the Saccharomyces cerevisiae PDH complex and used protein engineering techniques to study structure-function relationships. In the mid-1990s we cloned, sequenced, and expressed cDNAs encoding the two subunits comprising bovine PDH phosphatase and gained a deeper understanding of their nature and regulation (Fig. 1). This trail of discovery started in the spring of 1949, about six months after I joined the faculty of the Department of Chemistry at The University of Texas. At that time I started working on the isolation of a factor that replaced acetate in the growth medium for certain lactic acid bacteria. Research on the acetate-replacing factor was initiated by Esmond Snell at the University of Wisconsin, continued with a graduate student, Beverly Guirard, after Esmond moved to The University of Texas, and then pursued by Milton Getzendaner, a graduate student under Roger Williams' supervision. I inherited this project in the spring of 1949. We established that this factor is widely distributed in animal, plant, and microbial cells and that animal liver is a rich source. The factor is tightly bound to liver protein and is released only after hydrolysis in acid or base. At that time pharmaceutical companies were processing large amounts of pork and beef liver to obtain extracts suitable for treatment of pernicious anemia. Fresh liver was extracted with warm water, and the residual liver proteins and fatty material were dried and sold as an animal feed supplement. Arrangements were made with Eli Lilly and Company to obtain liver residue, and we developed procedures for extracting and purifying the acetatereplacing factor. In the late 1940s and early 1950s several other groups were trying to isolate factors that were similar to, if not identical with, the acetate-replacing factor. These factors included the "pyruvate oxidation factor, POF" of O'Kane and Gunsalus that was necessary for oxidation of pyruvate to acetate and carbon dioxide by Streptococcus faecalis; "protogen," an unidentified growth factor for a protozoan, Tetrahymena geleii, that was being purified by Stokstad, Jukes, and associates at Lederle Laboratories; and the "B. R. factor" of Kline and Barker required for growth of Butyribacterium rettgeri with lactate as the fermentable carbon source. In the fall of 1950, a collaboration with Gunsalus and the Lilly Research Laboratories was undertaken to isolate the acetate-replacing/pyruvate oxidation factor. The Lilly group adapted and scaled up isolation procedures developed by us, and concentrates of the growth factor that were 0.1 to1% pure were sent to us for further processing. One of the most exciting times in my life occurred on or about March 15, 1951, when I obtained the first pale-yellow crystals of the factor. The amount was minute, only about 3 mg. It was partially characterized and given the trivial name alpha-lipoic acid (Reed et al., 1951). The isolation procedure involved a 300,000-fold purification. A total of approximately 30 mg of crystalline lipoic acid was eventually isolated. We estimated that approximately 10 tons of liver residue were processed to obtain this small amount of the pure substance. NMR and mass spectrometers were not available in those days, but it was possible to establish that lipoic acid is either 6,8-, 5,8-, or 4,8-dithiooctanoic acid. That the correct structure is 6,8-dithiooctanoic acid (1,2-dithiolane-3-valeric acid) was established by synthesis of DL-lipoic acid, first achieved by the Lederle group (Reed, 1957). I was intrigued by this simple, yet unique substance and wanted to know more about its biological function, i.e., with what and how did it function in living cells. We, therefore, set about establishing this part of the trail, which turned out to be even more exciting than the isolation and characterization of lipoic acid. Elucidation of the mechanism of oxidative decarboxylation of -keto acids is a fascinating chapter in modern biochemistry. I shall review briefly the major developments in this story. The equation shown below represents the coenzyme A and NAD+-linked oxidative decarboxylation of -keto acids. In addition to CoA and NAD+, thiamin diphosphate, a divalent metal ion, protein-bound lipoic acid, and FAD are required. TPP, Lipoic acid, FAD RCOCO2H + NAD+ +CoASH &emdash;&emdash;&emdash;&emdash;&emdash;&emdash;&emdash;&emdash;&emdash;&emdash;&emdash;&emdash;&emdash ;&emdash;> RCO-S-CoA + CO2 + NADH +H+ With a few notable exceptions, prior to 1950 pyruvate and -keto-glutarate oxidation had been studied with particulate preparations from animal tissues and micro-organisms that were unsuitable for detailed analysis. However, these studies, notably those of Peters and his associates at Oxford, including Ochoa, had shown that thiamin diphosphate is required by enzymes that catalyze a decarboxylation of -keto acids. Other important developments in the late 1940s and early 1950s were Lipmann's discovery of coenzyme A, Stadtman's discovery of phosphotransacetylase and elucidation of the reaction catalyzed by this enzyme, and Lynen's demonstration of the thioester linkage in acetylCoA. Solubilization of bacterial and animal -keto acid oxidation systems in the early 1950s in the laboratories of Ochoa and Green was a significant advance. Korkes, Gunsalus, and Ochoa demonstrated that dismutation of pyruvate by enzyme preparations from E. coli and S. faecalis required a divalent metal ion, thiamin diphosphate, CoA, and NAD+. They succeeded in separating the pyruvate oxidation system of E. coli into two components, designated Fraction A and Fraction B. Jaganathan and Schweet isolated a pyruvate oxidation system from pigeon breast muscle in a highly purified state, with an apparent molecular weight of about 4 million. These preparations were shown subsequently to reduce NAD+ and to acetylate CoA. I remember Dick Schweet telling me about the skepticism expressed by some well-known enzymologists concerning the nature and purity of his "pyruvic oxidase" preparations. One prominent enzymologist suggested that Schweet had isolated a membrane fragment, and that if he continued with the purification he would eventually obtain a soluble enzyme with a respectable molecular weight. Seymour Kaufman showed that dismutation of -ketoglutarate by soluble preparations from pig heart required NAD+ and CoA and that one of the products was succinyl CoA. Sanadi and Littlefield isolated the -ketoglutarate oxidation system from pig heart as a highly purified preparation with an apparent molecular weight of 2 million and showed that NAD+ and CoA were the natural electron and acyl acceptors. The next important development was the isolation and characterization of lipoic acid described above. The presence of a disulfide linkage in lipoic acid recalled the interesting results of Peters and co-workers, who had observed a rather specific inhibition of the pigeon brain pyruvate oxidation system by trivalent arsenicals, particularly Lewisite, and a reversal of this toxic action by the dithiol 2,3-dimercaptopropanol (British antiLewisite, BAL), but not by monothiols. They postulated the existence of a dithiol structure as part of the pyruvate oxidation system. These results were duplicated by Gunsalus and associates with S. faecalis cells, and interpreted as indicating the involvement of dihydrolipoic acid in pyruvate oxidation. Gunsalus proposed that lipoic acid underwent a cycle of reactions in -keto oxidation comprising reductive acylation, acyl transfer, and electron transfer. Lipoic acid was visualized as functioning after TPP and before CoA and NAD+. Gunsalus, Hager, and associates obtained evidence for this proposal using lipoic acid and derivatives thereof in substrate amounts. They demonstrated that E. coli Fraction A contained a lipoyl transacetylase and that Fraction B contained a lipoyl dehydrogenase. In the late 1950s, Vince Massey showed that the lipoyl dehydrogenase component of the pig heart -ketoglutarate dehydrogenase complex is identical with Straub diaphorase, a flavoprotein described in 1939. [Franklin Leach observed that he could ascertain when the pyruvate complex was precipitated by its yellow color which should have been a clue to the flavin involvement]. Mechanistic studies by Massey and later by Charles Williams elucidated the catalytic mechanism. Model experiments conducted by Ronald Breslow with thiamin and analogs thereof led him to propose a mechanism of thiamin diphosphate action. 2-(1-Hydroxyethyl)thiamin diphosphate was proposed to be "active acetaldehyde." This hypothesis was confirmed and extended by enzymic studies carried out by Lester Krampitz and by Helmut Holzer and their associates. In my laboratory, we developed mild procedures for purification of the pyruvate and alpha-ketoglutarate oxidation systems from E. coli By the late 1950s, Masahiko Koike succeeded in isolating these enzyme systems as highly purified functional units with molecular weights in the millions (Koike et al., 1960). It was very exciting to see in the analytical ultracentrifuge of my friend and collaborator at NIH, Bill Carroll, a major symmetrical peak for each of the two highly purified preparations, and that the boundary of the yellow color of the flavoprotein was associated with the main peak. The molecular weights of these multi-enzyme units were determined to be 4.8 and 2.4 million, respectively. By careful, and persistent work over a period of several years, we dissected the pyruvate and alpha-ketoglutarate dehydrogenase complexes into their component enzymes and reassembled the large functional units from the isolated enzymes (Koike et al., 1963). We demonstrated that each of these functional units is composed of multiple copies of three enzymes, a pyruvate or alpha-ketoglutarate decarboxylase-dehydrogenase (E1), a dihydrolipoyl acetyltransferase or succinyltransferase (E2), and the flavoprotein, dihydrolipoyl dehydrogenase (E3). T hese three enzymes, acting in sequence, catalyze the reactions shown below. CH3COCO2- + E1TPP + H+ <==> CO2 + E1 CH3C(OH)=TPP (1) E1 CH3C(OH)=TPP + E2_LipS2 <==> E1TPP + E2-Lip(SH)-S-COCH3 (2) E2-Lip(SH)-S-COCH3 + CoASH <===> E2_Lip(SH2) + CH3COSCoA (3) E2-Lip(SH2) + E3FAD <===> E2-LipS2 + Dihydro-E3FAD (4) Dihydro-E3FAD + NAD+ <===> E3FAD + NADH + H+ (5) Fig. 2 Reaction sequence in -keto acid oxidation. Abbreviations: TPP, thiamin diphosphate; LipS2 and Lip(SH)2, lipoyl moiety and its reduced form. E1 catalyzes both the decarboxylation of the -keto acid (reaction 1) and the subsequent reductive acylation of the lipoyl moiety that is covalently bound to E2 (reaction 2). E2 catalyzes the acyl transfer to CoA (reaction 3), and E3 catalyzes the re-oxidation of the dihydrolipoyl moiety with NAD+ as the ultimate electron acceptor (reactions 4 and 5). Hayao Nawa showed in the late 1950s that the lipoyl moiety in the E. coli pyruvate and -ketoglutarate dehydrogenase complexes is attached in amide linkage to the epsilonamino group of a lysine residue (Nawa et al., 1960). An enzyme that hydrolyzes the lipoyllysyl linkage, lipoamidase, as well as an ATP-dependent enzyme that reincorporates the lipoyl moiety, i.e., a lipoate-protein ligase, were detected in S. faecalis extracts and partially purified. [This was the lipooic acid activating story which has been told above and which was the subject of Franklin Leach's thesis]. We proposed that this linkage provides a flexible arm, about 14 Å in length, for the reactive 1,2-dithiolane ring, permitting the lipoyl moiety to rotate among the catalytic sites of the three component enzymes of each complex. This is the so-called "swinging-arm" active-site coupling mechanism. Some 15 years later, Richard Perham and Cees Veeger and their associates attached a spin label to the protein-bound lipoyl moieties and showed by ESR spectroscopy that the lipoyl moieties exhibited considerable rotational mobility. These were exciting times for us in the late 1950s and early 1960s. We visualized the E. coli PDH complex as an organized mosaic of enzymes. To obtain evidence for this hypothesis, we turned to electron microscopy. I contacted Humberto Fernandez-Moran, who was then at the Massachusetts General Hospital, and arranged to bring a sample of the PDH complex to his laboratory. This was in January of 1962, and was indeed a memorable occasion. When our sample was negatively stained with phosphotungstate to provide contrast and then examined in the electron microscope, we saw a beautifully organized structure. The particles seen in the electron microscope had a diameter of about 300 Å, and there was a definite indication of subunits arranged in a regular manner (Fernandez-Moran et al., 1964). Within about two years, we set up an electron microscopy laboratory in the Biochemical Institute at The University of Texas. Electron microscopy studies were carried out by my associate Robert Oliver, X-ray crystallographic studies by collaborators David DeRosier and Marvin Hackert, and sedimentation equilibrium molecular weight determinations by Petr Munk. The results demonstrated that both the acetyltransferase and the succinyltransferase (E2) consist of 24 apparently identical polypeptidechains arranged as eight trimers in a cube-like particle exhibiting octahedral (432) symmetry. Multiple copies of E1 and E3 are attached to E2 by noncovalent bonds. In the PDH complex, 12 E1 dimers and 6 E3 dimers are apparently arranged, respectively, on the 12 edges and in the six faces of E2 (Reed, 1974). In the late 1960s part of our research effort was directed toward isolation and characterization of the mammalian pyruvate and -ketoglutarate dehydrogenase complexes, which are localized to mitochondria, within the inner membrane-matrix compartment. Procedures were developed for preparation of mitochondria on a large scale from bovine kidney and heart (with the advice and assistance of my friend and colleague, Dan Ziegler), and relatively mild procedures were developed to isolate the pyruvate and -ketoglutarate dehydrogenase complexes from the mitochondrial extracts. In the course of attempts to stabilize these complexes in crude extracts of bovine kidney mitochondria, Tracy Linn observed that the PDH complex, but not the alphaketoglutarate dehydrogenase complex, underwent a time-dependent inactivation in the presence of ATP. A systematic investigation revealed that the bovine kidney and heart PDH complexes are regulated by a phosphorylation-dephosphorylation cycle (Linn et al., 1969). Phosphorylation and concomitant inactivation of the complex is catalyzed by an ATPdependent kinase, which is tightly bound to the complex, and dephosphorylation and concomitant reactivation is catalyzed by a Mg2+-dependent phosphatase, which is loosely attached to the complex. It seemed curious at the time (1968) that inactivation of the PDH complex by phosphorylation had not been detected earlier. The explanation may lie in a remark by Henry Lardy after receiving a preprint of our paper on the phosphorylation and inactivation of the PDH complex. (This finding) "explains why we have never been able to get pyruvate to be oxidized in submitochondrial particles, because we invariably add ATP to keep things in the 'optimum' state." This control mechanism was subsequently confirmed in the laboratories of Otto Wieland, Philip Randle, S.E. Severin, and other investigators with preparations of the PDH complex from other mammalian tissues and from pigeon breast muscle, plant tissue, and Neurospora crassa. Over a period of several years our group separated the bovine kidney and heart PDH complexes into their component enzymes, including the kinase and the phosphatase, and characterized the individual enzymes (Linn et al., 1972). The bovine heart PDH complex has a molecular weight of about 9.5 million. Its subunit composition is now known to be 60 E2 subunits, 30 E1 tetramers (alpha2beta2), and 12 E3 dimers, which are positioned on the E2 core by 12 E3-binding protein (protein X) monomers. The E1alpha subunit undergoes phosphorylation and dephosphorylation. The appearance of E2 in the electron microscope is that of a pentagonal dodecahedron, and its design is based on icosahedral (532) symmetry. We proposed that the E1 tetramers are located on the 30 edges and the E3 dimers in the 12 faces of the pentagonal dodecahedron. A novel architectural feature of dihydrolipoamide acyltransferases was revealed initially in our laboratory by limited proteolysis of the E. coli acetyltransferase and by electron microscopy. Dennis Bleile found in the late 1970s that trypsin cleaved the acetyltransferase, which contained radioactive lipoyl moieties, into two large fragments (Bleile et al., 1979). One fragment, designated the lipoyl domain, contained the covalently bound lipoyl moieties and exhibited an extended structure. The other tryptic fragment exhibited a compact structure and contained the active site, the intersubunit binding sites, and the binding sites for E1 and E3. The assemblage of compact catalytic domains constitutes the inner core of E2, conferring the cube-like appearance in the electron microscope. The two domains are connected by a trypsin-sensitive hinge region. We suggested that movement of lipoyl domains and not simply rotation of lipoyllysyl moieties provides the means to span the physical gaps between catalytic sites on the complex. These early findings on the domain structure of dihydrolipoamide acyltransferases were confirmed and extended by studies involving molecular genetics, limited proteolysis, and proton NMR spectroscopy in the laboratories of John Guest and Richard Perham. Briefly, the amino-terminal segment possesses one, two, or three lipoyl domains, followed by a domain that is involved in binding E3 and/or E1, and then by a catalytic domain that contains the active site as well as additional subunit binding sites. The domains are connected by flexible segments or hinge regions that are rich in alanine, proline, and charged amino acid residues. Recently, Wim Hol and associates determined the crystal structure at 2.6 Å resolution of the cube-like inner core of the dihydrolipoamide acetyltransferase from the Azotobacter vinelandii PDH complex. Richard Perham and associates used multi-dimensional NMR to determine the threedimensional solution structures of the lipoyl domain and the E1/E3-binding domain of the acetyltransferase from Bacillus stearothermophilus. Hol and associates also determined the crystal structure of this E3-binding domain complexed with an E3 dimer. These structures provide a deeper understanding of how the lipoyl domain can move between the active sites of E2 and E3 in the PDH complex. In the late 1970s Flora Pettit and Steve Yeaman purified to apparent homogeneity and characterized the bovine branched-chain -keto acid dehydrogenase complex. In the 1980s Zahi Damuni isolated and characterized the phosphatase that participates in the regulation of this complex, and he also isolated and characterized a potent heat-stable inhibitor of the phosphatase. Hormonal regulation of the mammalian PDH complex is particularly fascinating because it involves signal transduction not only across the cell membrane but also across the inner mitochondrial membrane to target the PDH phosphatase and, consequently, the PDH complex, located in the mitochondrial matrix. It is now known that the major regulators of the phosphatase activity are Ca2+ and Mg2+, which involve the hormones epinephrine and insulin, respectively. In the early 1970s our group partially purified PDH phosphatase from bovine heart and kidney mitochondria and showed that it requires Mg2+ or Mn2+ for activity. Denton, Randle, and Martin subsequently reported that Ca2+ stimulated the activity of the phosphatase in the presence of Mg2+. Flora Pettit and Tom Roche in our group showed that Ca2+ mediates translocation of the phosphatase to the E2 component of the PDH complex, presumably in proximity to its substrate, phosphorylated E1, thereby increasing the rate of dephosphorylation. This Ca2+-mediated translocation apparently is the molecular basis of the epinephrine-induced activation of PDH phosphatase observed by Richard Hansford, Richard Denton, and other investigators. In the early 1980s Martin Teague, Flora Pettit, and co-workers purified PDH phosphatase to near homogeneity and showed that it consists of a Mg2+-dependent and Ca2+stimulated catalytic subunit (50 kDa; PDPc) and a flavoprotein of unknown function (100 kDa; later designated PDPr) (Teague et al., 1982). Zahi Damuni showed that polyamines, particularly spermine, increase the sensitivity of PDH phosphatase to Mg2+. Denton and associates subsequently showed that insulin stimulates the activity of PDH phosphatase in adipose tissue by increasing the sensitivity of the phosphatase to Mg2+. Spermine apparently mimics the insulin effect. The function of PDPr remained a mystery until Janet Lawson recently cloned and expressed cDNA encoding PDPc. By comparing the properties of recombinant PDPc and the native PDH phosphatase heterodimer (PDPc bound to PDPr), we obtained insight into the function of PDPr. Jiangong Yan found that PDPr decreases the sensitivity of PDPc to Mg2+ and that spermine increases the sensitivity of PDH phosphatase but not PDPc to Mg2+, apparently by interacting with PDPr (Yan et al., 1996). We interpret these observations to indicate that PDPr blocks or distorts the Mg2+-binding site of PDPc and that spermine produces a conformational change in PDPr (allosteric effect) that reverses its inhibitory effect. These observations raise the intriguing prospect that an insulin-induced allosteric effect on PDPr may underlie its stimulation of PDH phosphatase activity. To gain further understanding of structure-function relationships in eukaryotic PDH complexes, we initiated in the late 1980s molecular genetic studies of the PDH complex in the yeast Saccharomyces cerevisiae. The genes encoding the five proteins comprising the complex (E1 , E1, E2, E3BP, and E3) were cloned, sequenced, expressed, and disrupted. Studies on E3-binding protein (E3BP) confirmed and extended previous studies of Tom Roche and of Gordon Lindsay and their associates with the protein X component of the bovine PDH complex. E3BP and E2 apparently evolved from a common ancestor. E3BP possesses an amino-terminal lipoyl domain, followed by an E3binding domain, and then by a carboxyl-terminal domain that is involved in anchoring E3BP to the inner core of E2. Binding studies in conjunction with cyroelectron microscopy and three-dimensional image reconstruction in collaboration with James Stoops and Timothy Baker and their associates revealed a unique structural organization of the S. cerevisiae PDH complex and, by analogy, of the mammalian complex (Stoops et al., 1977). E2 consists of 20 cone-shaped trimers at the vertices of a pentagonal dodecahedron. There are 12 large openings that lead into a central cavity. It was generally believed that the other components of these complexes are bound on the outside of the E2 scaffold. By contrast, our results show that E3BP binds near the tips of the E2 trimers within the central cavity and anchors an E3 dimer inside each of the 12 pentagonal faces of E2. Our finding that the E2 structure, with 532 molecular symmetry, can physically accommodate only one BP-E3 complex in each of its 12 pentagonal-shaped faces provides a satisfactory explanation of the unique polypeptide chain ratio in the S. cerevisiae and mammalian PDH complexes (60 E1:60 E1:60 E2:12 BP:24 E3). I hope these recollections have given some appreciation of the thrill and excitement I have experienced in establishing this trail of research from lipoic acid to the structure, function, and regulation of the -keto acid dehydrogenase complexes. I have been accompanied in the various stages of this journey by excellent associates, including undergraduate, graduate, and postdoctoral students, technicians, and members of the senior staff of the Biochemical Institute, and by collaborators at other universities and institutes. Acknowledgments I am pleased to acknowledge the Clayton Foundation for Research and the National Institutes of Health for generous financial support. References Bleile DM, Munk P, Oliver RM, Reed LJ. 1979. Subunit structure of dihydrolipoyl transacetylase component of pyruvate dehydrogenase complex from Escherichia coli. Proc Natl Acad Sci USA 76:43854389. Fernandez-Moran H, Reed LJ, Koike M, Willms CR. 1964. Electron microscopic and biochemical studies of pyruvate dehydrogenase complex of Escherichia coli. Science 145:930-932. Koike M, Reed LJ, Carroll WR. 1960. -Keto acid dehydrogena tion complexes I. Purification and properties of pyruvate and alpha-ketoglutarate dehydrogenation complexes of Escherichia coli. J Biol Chem 235:1924-1930. Koike M, Reed LJ, Carroll WR. 1963. -Keto acid dehydrogenation complexes IV. Resolution and reconstitution of the Escherichia coli pyruvate dehydrogenation complex. J Biol Chem 238:30-39. Linn TC, Pettit FH, Reed LJ. 1969. -Keto acid dehydrogenase complexes X. Regulation of the activity of the pyruvate dehydrogenase complex from beef kidney mitochondria by phosphorylation and dephosphorylation. Proc Natl Acad Sci USA 62:234-241. Linn TC, Pelley JW, Pettit FH, Hucho F, Randall DD, Reed LJ. 1972. -Keto acid dehydrogenase complexes XV. Purification and properties of the component enzymes of the pyruvate dehydrogenase complexes from bovine kidney and heart. Arch Biochem Biophys 148:327-342. Nawa H, Brady WT, Koike M, Reed LJ. 1960. Studies on the nature ofprotein-bound lipoic acid. J Am Chem Soc 82:896-903. Reed LJ. 1957. The chemistry and function of lipoic acid. Adv Enzymol 18:319-347. Reed LJ. 1974. Multienzyme complexes. Acc Chem Res 7:40-46. Reed LJ, DeBusk BG, Gunsalus IC, Hornberger CS Jr. 1951. Crystalline -lipoic acid: A catalytic agent associated with pyruvate dehydrogenase. Science 114:93-94. Stoops JK, Cheng RH, Yazdi MA, Maeng C-Y, Schroeter JP, Klueppelberg U, Kolodziej SJ, Baker TS, Reed LJ. 1997. On the unique structural organization of the Saccharomyces cerevisiae pyruvate dehydrogenase complex. J Biol Chem 272:5757-5764 Teague WM, Pettit FH, Wu T-L, Silberman SR, Reed LJ. 1982. Purification and properties of pyruvate dehydrogenase phosphatase from bovine kidney and heart. Biochemistry 21:5585-5592. Yan J, Lawson JE, Reed LJ. 1996. Role of the regulatory subunit of bovine pyruvate dehydrogenase phosphatase. Proc Natl Acad Sci USA 93:4953-4956. 2 Lester J. Reed was born in New Orleans, Louisiana, in 1925 and received a B.S. in chemistry from Tulane University in 1943. He was awarded a Ph.D. in organic chemistry from the University of Illinois in 1946 under R.C. Fuson. He was a research associate in biochemistry under Vincent du Vigeaud at Cornell University Medical College from 1946 to 1948. He joined the Department of Chemistry at The University of Texas in 1948 as an Assistant Professor. He is currently Ashbel Smith Professor of Chemistry and Biochemistry. He was Director of the Clayton Foundation Biochemical Institute from 1963 to 1996. Dr. Reed is a member of the National Academy of Sciences and the American Academy of Arts and Sciences. He received the Eli Lilly & Co. Award in Biological Chemistry of the American Chemical Society in 1958, an honorary doctorate from Tulane University in 1977, and the Merck Award of the American Society for Biochemistry and Molecular Biology in 1994.