* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Structural
Nucleic acid analogue wikipedia , lookup
Expression vector wikipedia , lookup
Fatty acid synthesis wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Magnesium transporter wikipedia , lookup
Ancestral sequence reconstruction wikipedia , lookup
Interactome wikipedia , lookup
Western blot wikipedia , lookup
Point mutation wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Homology modeling wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Genetic code wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Biosynthesis wikipedia , lookup
Peptide synthesis wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
Ribosomally synthesized and post-translationally modified peptides wikipedia , lookup
Metalloprotein wikipedia , lookup
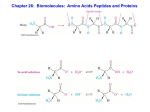
Biochemistry Sixth Edition ביוכימיה היא מדע החוקר את התכונות הכימיות המאפיינות מולקולות המיוצרות בתאים חיים . ומאפשרות להם לבצע את תפקידם,וייחודיות להם Berg • Tymoczko • Stryer Chapter 2 Protein Composition and Structure Copyright © 2007 by W. H. Freeman and Company Functions of some Proteins Catalysis (enzymes) Structural (collagen) Contractile (muscle) Transport (hemoglobin) Storage (myoglobin) Electron transport (cytochromes) Hormones (insulin) Growth factor (EGF) DNA binding (histones) Ribosomal proteins Toxins and venoms (cholera & melittin) Vision (opsins) Immunoglobins Levels of Protein Structure • Primary structure (1o) – sequence of amino acids starting from the N-terminus of the peptide. • Secondary structure (2o) – conformations of the peptide chain from rotation about the a-Cs, e.g. a-helices and b-sheets, etc. • Tertiary structure (3o) – three dimensional shape of the fully folded polypeptide chain. • Quaternary structure (4o) - arrangement of two or more protein chains into multisubunit molecule Levels of Protein Structure Bonding: 1o = covalent 2o = H-bond 3o = covalent & noncovalent 4o = noncovalent Amino Acids The 20 common are those used in making protein on a ribosome using mRNA and tRNA. These are called a-amino acids since each has a carboxyl group and an amino group attached to an a-carbon atom. They differ by the sidechain or “R” group. a +NH -CH-COOH 3 l R Amino Acid Classification Classification is made using the structure of the side chain, R. (* = essential) 1. None (hydrogen): 2. Non-polar: Aliphatic: Alicyclic: 3. Aromatic: 4. Polar uncharged: 5. Thiol: 6. Acidic: 7. Basic: Gly Ala, Val*, Leu*, Ile*, Met* Pro Phe*, Tyr, Trp* Ser, Thr*, Asn, Gln Cys Asp, Glu Lys*, His*, Arg* ball & stick model wedge Fischer projection Amino Acid Names & Codes All are Chiral at the a-carbon atom except Gly D, L Assignments The convention for making D, L assignments is to draw a Fischer projection with the carboxyl at the top and the R at the bottom. +NH 3 to the left = L to the right = D COO H + NH3 H CH 2-OH COO H H NH3+ CH 2-OH L stereochemistry = S, for this case Acids and Bases, pH, pKa • pH is defined as the negative logarithm of the concentration of H+ pH = - log [H+] = log 1/ [H+] • pKa is defined as the negative logarithm of an acid ionization constant Ka. pKa = - log [Ka] = log 1/ [Ka] Acetic Acid is a Weak Acid • Weak acids and bases do not ionize (dissociate) completely in H2O. CH3COOH <==> CH3COO- + H+ acetic acid acetate anion conjugate acid form conjugate base form Acid Ionization Constant [CH3COO-][H+] Ka = -----------------[CH3COOH] [CH3COO-] -log Ka = -log [H+] + -log ---------------[CH3COOH] [CH3COO-] pH = pKa + log ---------------[CH3COOH] Henderson-Hasselbalch Written in a more general form this expression is called the Henderson-Hassebalch equation. This relates the pH of a solution to the pKa of the weak acid in solution and the ratio of its conjugate base/conjugate acid forms. [A-] pH = pKa + log -----[HA] Buffers Resist Change in pH • Buffer capacity is the ability of a solution to resist changes in pH. • Most effective buffering occurs where: solution pH = buffer pKa • At this point: [conjugate acid] = [conjugate base] • Effective buffering range is usually at pH values equal to the pKa ± 1 pH unit Amino Acid Ionization, pKas Each of the 20 common a-amino acids has two pKa values, for the carboxyl group and the amino group attached to the a-carbon. +NH +NH -CH -COO- + H+ -CH -COOH < === > 3 2 3 2 +NH - < === > NH -CH -COO- + H+ -CH -COO 3 2 2 2 Seven of the 20 have an ionizable sidechain and therefore have a third pKa value. Amino Acid pKas Gly Ala Val Leu Ile Met Pro Phe Trp Ser Thr Asn Gln Cys Asp Glu Tyr Lys His Arg 2.18 a-COOH 2.34 2.34 2.32 2.36 2.36 2.28 1.99 1.83 2.83 2.21 2.63 2.02 2.17 1.71 2.09 2.19 2.20 8.95 1.82 2.17 a-+NH3 9.60 9.69 9.62 9.60 9.60 9.21 10.60 9.13 9.39 9.15 10.43 8.80 9.13 10.78 9.82 9.67 9.11 R (sidechain) 8.33 3.86 4.25 10.07 10.79 9.17 9.04 6.00 12.48 Peptide 3.86 D, 4.25 E Peptide 10.0 Effect of Change in pH The Structure of an Amino Acid An amino acid can never exist as an uncharged compound. © 2014 Pearson Education, Inc. The pI of Alanine The isoelectric point (pI) of an amino acid is the pH at which it has no net charge. © 2014 Pearson Education, Inc. The pI of Lysine © 2014 Pearson Education, Inc. The pI of Glutamic Acid © 2014 Pearson Education, Inc. Formation of a Peptide Two amino acids are joined together to form a peptide (amide) bond with a loss of HOH. After becoming part of a peptide or protein these are called “residues” due to loss of HOH. Primary Structure (1o) The sequence of amino acids (N-term to C-term) in a peptide or protein is its primary structure. A pentapeptide Disulfide Bond Formation Disulfide Bridges Disulfide bridges contribute to the overall shape of the protein. © 2014 Pearson Education, Inc. Straight and Curly Hair © 2014 Pearson Education, Inc. Insulin Insulin has •two interchain disulfide bridges and •one intrachain disulfide bridge. © 2014 Pearson Education, Inc. Peptide Bond Resonance Due to resonance participation of the unshared pair of electrons on N, amides are neutral. .. Peptide Bond Planarity 6 atoms are coplanar Peptide Bond Structures Less crowded in the favored trans arrangement Rotation Sites, y and f The rotational arrangements about a-carbons of a peptide or protein gives its secondary (2o) structure. f View from N-term to a-carbon y View from a-carbon to C-term Ramachandran Plot Secondary Structure (2o) The two most common secondary structures are the a-helix and the b-sheet. Each of these 2o structures have fairly specific y and f angles. All other rotational angles represent “random” secondary structure. Secondary structure is maintained by hydrogen bonding. a-helix by intramolecular H-bonds b-sheet by intermolecular H-bonds a-helix in a Ramachandran Plot y = -47 f = -57 The a-helix, a 3.613 helix The a-helix Hydrogen Bond Contacts a-helix in a Protein b-sheet in a Ramachandran Plot y = +135 f = -139 A b-sheet strand Anti-parallel b-sheet Parallel b-sheet Anti-parallel b-sheet b-sheet in a Protein b-turn or hairpin turn A turn is 4-5 aa residues. A b-turn (hairpin) is 4 aa residues. Loops are usually larger than turns. >5 aa residues a-Keratin, a fibrous protein, forms an a coiled coil Each strand is a modified a-helix, 3.5 residues/turn. Right-handed helices form a left-handed supercoil. Collagen, a triple helix Gly every third residue; Gly-Pro-HPro is frequent. Interstrand H-bond at Gly. No Cys, so, no -S-S- Tertiary Structure (3o) The three dimensional folding of a polypeptide is its tertiary structure. Both the a-helix and b-sheet may exist within the tertiary structure. Generally the distribution of amino acid sidechains in a globular protein finds mostly nonpolar residues in the interior of the protein and polar residues on the surface. Tertiary structure is maintained by noncovalent interactions and disulfide bonding. Myoglobin, a globular protein Myoglobin Surface Blue = charged Cross-section Yellow = hydrophobic Porin, a membrane spanning protein Domain A domain is a discrete globular area within protein. There are four general types: All a All b Mixed a/b a+b only a- helices and loops only b- sheet & loops or turns alternating abab cluster of a then cluster of b Domain Multiple domains exist in the protein below. Quaternary Structure (4o) is an assembly of 3o structures (two or more subunits). A dimer Hemoglobin, a tetramer Quaternary structure is maintained by noncovalent interactions Denaturing Proteins Denaturing agents destroy the protein 3o structure (causes the protein to unravel). Methods: Heat; Extremes of pH; Detergents; Mechanical agitation; Mercaptoethanol – breaks -S-S- bonds; 6M guanidine HCl or 10M urea: these are chaotropic agents that break up noncovalent interactions. Denaturing agents Disulfide oxidation-reduction Oxidized Reduced Ribonuclease 4 disulfide bonds. -S-S- Then removing urea and ME permits reoxidation. Reoxidation reforms –S-S- but not necessarily in the correct place. A trace of ME allows reduction/oxidation to occur until the low free energy form is found and >98% of activity is restored. Sharp Transition suggests an all or nothing effect in denaturing. Lysozyme Ribbon diagrams 3o structure Active site & -S-S- Protein Cleavage Protein sequencing is most manageable with small polypeptides. Therefore, in order to sequence a large protein, it must be cleaved into smaller pieces. Cleavage is conducted using either chemical or enzymatic methods. The pieces must be separated and purified before sequencing. Chemical and Enzymatic Cleavage CNBr Cleavage at Met Enzymatic cleavage by Trypsin שאלה :לפניך תוצרי הביקוע של פפטיד באמצעות טריפסין וכימוטריפסין: כימוטריפסין: Val-Ser-Arg-Gly-Trp Met-Asn-Lys-Phe טריפסין: Phe-Val-Ser-Arg Met-Asn-Lys Gly-Trp הערות :טריפסין חותך בצד הקרבוקסילי של Lys, Arg : וכימוטריפסין חותך בצד הקרבוקסילי של . Phe,Trp,Tyr : מהו רצף החומצות האמיניות בפפטיד. תשובה: Met-Asn-Lys-Phe-Val-Ser-Arg-Gly-Trp 82 An Example, Peptide overlap