* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Lecture 1
Implicit solvation wikipedia , lookup
Rosetta@home wikipedia , lookup
List of types of proteins wikipedia , lookup
Protein design wikipedia , lookup
Bimolecular fluorescence complementation wikipedia , lookup
Structural alignment wikipedia , lookup
Protein moonlighting wikipedia , lookup
Homology modeling wikipedia , lookup
Protein domain wikipedia , lookup
Protein purification wikipedia , lookup
Western blot wikipedia , lookup
Intrinsically disordered proteins wikipedia , lookup
Protein mass spectrometry wikipedia , lookup
Circular dichroism wikipedia , lookup
Protein folding wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
Protein–protein interaction wikipedia , lookup
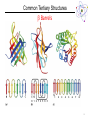
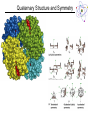
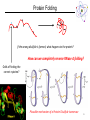
Lecture 6 Protein Structure and Properties 1 Torsion Angles When both peptide bonds are in trans conformation: F = -180 Y = 180 2 Globular Proteins Exist as compact, roughly spherical molecules Highly Solvated 3 Superseconday Structure Some common groupings of Secondary Structural Motifs bab b hairpin aa Helix-turn-helix Greek Key 4 Superseconday Structure Find the Common Structural Motifs bab b hairpin aa Helix-turn-helix 5 Common Tertiary Structures b Barrels 6 Common Tertiary Structures Multi-domain Proteins 7 Protein Stability Calculations show that proteins are NOT very stable molecules Net stabilization of ~0.4 kJ mol-1 for each amino acid Electrostatic Hydrogen Bonding Hydrophobic Covalent (disulfide) The cumulative stabilizing energy of each of these forces can be in the thousands of kJ mol-1 8 Protein Stability – Electrostatics 𝑘𝑞1 𝑞2 𝐸= 𝐷𝑟 Protein cores are approximated to have a dielectric constant of 4 What residues have a potential for ionic interactions? 9 Protein Stability – Hydrogen Bonds Proteins maximize H-bonding by selecting a local H-bond network Side Chain – Backbone H-bonds are clustered @ Helix termini H – bonding position Type of 2˚ Structure Percentage of H bonds nn-4 a helix ~25% nn-3 310 helix and turns ~25% b sheets ~25% 10 Protein Stability – dipoles Which amino acids can be involved in dipole-dipole interactions? What else is an important dipole interaction? d+ dd+ d- 11 Protein Stability – The Hydrophobic Effect Protein Stability – Hydrophobicity This scale takes the cumulative hydropathy for 9 consecutive amino acids Quaternary Structure and Symmetry Measuring Protein Stability AbsCW - AbsCCW Heat Circular Dichroism Spectroscopy Absorbance of circularly polarized light is dependent on the chirality of the sample How could we denature a protein without heating it up? Protein Folding If the wrong disulfide is formed, what happens to the protein? How can we completely reverse RNase A folding? Odds of finding the correct cysteine? Plausible mechanism of a Protein Disulfide Isomerase 16 Protein Folding – Energy Landscapes Native conformation Local Energy minimum = Wrong Structure 17 Overcoming the Local Energy Minima Molecular Chaperones bind to unfolded proteins and prevent non-native conformations 18 How well do you understand protein structure? 19 Which of the following statements about the peptide bond is true? A.It is a phosphoamide bond. B.It is a double bond. C.Atoms of the peptide bond are located in a single plane. D.The peptide bond is charged. Which of the following statements is FALSE with respect to the ribbon diagrams shown below ? 1 A. B. C. D. 2 3 Protein 2 contains parallel b-sheet strands. Protein 3 contains a prosthetic group. The secondary structure of protein 3 is all a-helix. These proteins have both secondary and tertiary structure. 21 a-helix and b-sheet secondary structure is located mainly in the interior of folded proteins, whereas irregular loops occur on the outside. Why? Which of the following series of amino acids is MOST likely to be at the surface of a watersoluble globular protein? A. ……..met-phe-pro-ile-leu…….. B. ……..tyr-phe-gly-asn-leu……... C. ……..glu-asn-ser-thr-arg……… D. ……..val-ala-val-glu-val……… E. ……..met-cys-pro-ala-tyr……... Which of the following amino acid residues form hydrogen bonds with Ala residues located in an ahelix? A. Residues in a neighbouring a-helix. B. Residues located within the same a helix. C. None, because Ala is unable to form hydrogen bonds. The following sequence of amino acids is very unlikely to fold into a stable ahelix. Why? Asp-Glu-Ser-Asp-Glu-Glu-Arg-Val-Asp-Ala-Glu-Asp Would a b-sheet be more likely? Why? What Secondary Structures correspond with each label? What is the symmetry of the structure shown? A. C2 B. C3 C. C6 D. C12