* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download emboj7601952-sup
Genomic imprinting wikipedia , lookup
Public health genomics wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Gene desert wikipedia , lookup
Gene nomenclature wikipedia , lookup
Genome (book) wikipedia , lookup
Gene expression programming wikipedia , lookup
Nicotinic acid adenine dinucleotide phosphate wikipedia , lookup
Genome evolution wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Designer baby wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Gene expression profiling wikipedia , lookup
History of genetic engineering wikipedia , lookup
Microevolution wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
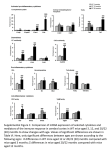
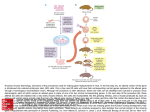
Supplemental Information Supplemental Figure 1. The MURF gene family and the KO mouse models used in this study. (A) MuRFs (brief for Muscle specific RING Finger; see Spencer 2000) correspond to a gene family coding for three highly homologous RING fingers proteins, referred to as MuRF1,2 and 3; also called RNF 28-30; or TRIM 63, 55, 54). The three MuRFs share highly conserved N-terminal RING finger domains, followed C-terminally by a MuRF family conserved motif (MFC), a B-box, and a coiled-coil region. The MuRF1Bcc fragment expressed in E. coli corresponds to residues 109-315, MuRFcc corresponds to residues 146-315. (B) MuRF1 and MuRF2 were inactivated by a homologous recombination into exon 2 and exon 5 respectively (stop codons in all three frames are provided by the Neo cassette; gene targeting essentially as described in Witt et al., 2006). (C) Genotyping with the primer pairs indicated in (B). (D) RT-PCR detects loss of MuRF1 and MuRF2 transcription in KO mice. MuRF3 transcription is intact. Aldolase: Control for input mRNA/cDNA quantity and quality. Primer for RT-PCR: MuRF1 for: AGAAGTCGGGGGTCAGGGGACG; MuRF1 rev: GGTCCATGATCACTTCATGGCGGCACGAGG; MuRF2 for: GAGCACTTCTCTGAATTACAAGTCTTTCTCC; MuRF2 rev: GCTGAGAGCCAATGGGCACCACCTCCCC; MuRF3 for: GCCCCATCTGCCTGGAGATGTTCTCCAAGCC; MuRF3 rev: GCCGCTCCTCCTCCAGCCCCGCATTGCCC Aldolase for: AGCTGTCTGACATCGCTCACCG; Aldolase rev: CACATACTGGCAGCGCTTCAAG 1 Supplemental Figure 2. Post mortem histology of the lung of dKO animals. dKO mice at day 13-15 of age found dead in their cages were histologically examined post mortem. A. Left: Wildtype mice show intact alveoles and no signs of lung compression and edema. Right: Young dKO animals show a partial compression of the lung (arrow) while the peripheral parts of the lung which are not hypostatic are filled with air. B. Left: Severe congestion (Hyperemia) in dKO lungs with acute edema (arrow) and partial effacement of the alveolar cells. Right: The myocardium shows microthombi (arrow) but no areas of necrosis and no signs of fibrosis. Supplemental Figure 3. Comparison of WT and dKO heart at months 14-18. ECG Eindthoven scans of one WT (A), one MuRF1KO (B), and one dKO mouse (D) indicates intact excitation conduction in dKO hearts. Studied mice were 12 months old (n=2 for each genotype). Supplemental Video 4-7. Time-resolved MRI imaging of a WT (Video Sup_4 and Sup_5) and a dKO mouse (Video Sup_6 and Sup_7), showing massive ventricular hypertrophy in dKO. Supplemental Figure 8. Absolute weights of bodies, hearts (ventricles) and quadriceps. Mean absolute weights of WT and dKO mice are compared in three groups: young animals (2 month, WT: n=4, dKO: n=3), adult animals (6-8 months, WT: n=15, dKO: n=10), old animals (16-18 months, WT: n=14, dKO: n=5; significant p-values: BW old animals p=0.003, HW young animals p=0.001, HW adult animals p=0.003, HW old animals p=0.009). Supplemental Table 9. Summary of microarray data. 2 The table shows 40 upregulated genes including those genes mentioned in the text (red: alphaactin, ANP type A and B, Myosin Light Chain, CARP, p62/sequestosome1, and atogrin/MafBx. MuRF1 is the most downregulated gene in the microarray. MuRF2 is not represented on the chip (Affymetrix Mouse Genome 430 2.0). Genes described in the text but showing no dysregulation are presented in the bottom. Supplemental Figure 10. (A) Western Blots of young mice hearts. As in old mice (figure 5) the MuRF1/MuRF2 binding protein CARP is upregulated. No clear trend can be seen with FHL2 antibody. In agreement with the results in old mice, cTroponin I does not show differential expression in our MuRF1/MuRF2 KO panel. In contrast, the hypertrophy marker ANP is strikingly upregulated in young dKO mice. (B) In immunofluorescence, the protein level of the early response gene c-fos (stained with Cy3, red color) is found elevated in the nuclei (blue DAPI staining) of dKO mice. Size bar is 10 µm. 3