* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Protein synthesis
Survey
Document related concepts
Homology modeling wikipedia , lookup
Protein domain wikipedia , lookup
Protein design wikipedia , lookup
List of types of proteins wikipedia , lookup
Intrinsically disordered proteins wikipedia , lookup
Protein folding wikipedia , lookup
Protein structure prediction wikipedia , lookup
Bimolecular fluorescence complementation wikipedia , lookup
RNA-binding protein wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Western blot wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
Protein mass spectrometry wikipedia , lookup
Transcript
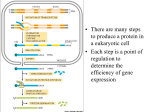
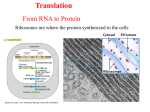
Protein Synthesis Prof.Dr. Gönül Kanıgür Protein synthesis • Protein synthesis requires more than a hundred macromolecules [mRNA,tRNA,activating enzymes,protein factors,ribosomes] • There are three steps in protein synthesis • -initiation • Elongation • termination Activation of aminoacids • +Activated precursors of prot.synthesis are aminoacil-tRNAs • The carboxyl group of an aa is joined to the 3terminus of a tRNA • The linking of aa to its coresponding tRNA is catalyzed by an aminoacyl-tRNA synthetase • ATP • Reaction aa+ATP+tRNA---aa-tRNA+AMP+PPi • İn the first step,the aa is joined to AMP,forming an aminoacyl AMP intermediate • İn the second step,the aa is transfered to the 3’CCA terminus of the acceptor tRNA and AMP is released. • Both step of the reaction are catalyzed by aminoacyl tRNA synthetases. İnitiation of protein synthesis • İn both eucaryotic and procaryotic cells,translation always initiates with the aa methionine usually encoded by AUG. • Alternative initiation codon such as GUG are used in bacteria. • İn most bacteria protein synthesis is initiated with a modified methionine(N-formyl methionine),whereas unmodified methionine initiate prot.synt.in eucaryotes Formation of initiatiator tRNA • Methionyl-tRNAsynthetaze adds methionine to the methionyl tRNAs • Second step is the formylation • Formylation is catalized by transformylase (N10- formyltetrahydrofolate) • This product is called formyl methionyl tRNA RİBOSOMES • Ribosoms in procaryotes consist of a large and small subunit. • 30S subunit contains 21 proteins and 16S ribosomal RNA. • 50S subunit contains 34 proteins and 2 rRNA molecules. (23S and 5S) The bacterial ribosome Protein factors: • Initiation factor 1 (IF1) prevents the reassociation of diassociated 50S and 30S subunits. • IF2 is necessary to the formation of IF2.GTP.fMet tRNAf complex (ternary complex) • IF3 is similar to IF1 initiation of protein synthesis: • 1) Ternary complex formation (IF2.GTP.initiator tRNA) • This complex binds to mRNA to form 30S initiation complex • The intereacting components are(mRNA+30S subunit+fMet tRNAf+GTP+Initiation factors) • The fmet-tRNAf is located to the AUG (initiator)codon • 50S subunit joins to 30S initiation complex to form a 70S initiation complex. GTP is hydrolysed, all these factors are released. • The 70S initiation coplex is ready for elongation step of protein synhesis. • tRNA-mRNA-rRNA base-pairing interactions • determine accuracy of protein synthesis. • • • • • • • İnitiation Formation of a 30S initiation complex 30S ribosomal subunit mRNA Formylmethionyl-tRNA GTP İnitiation factors [E,P,A] three tRNA binding sites on 30S subunit ELONGATİON CYCLE IN PROTEIN SYNTHESIS • This cycle consist of three steps: • binding of aa-tRNA to the A site[codon recognation] • Peptid bond formation • Translocation • The ribosome has three tRNA binding sites; P [peptidyl],A [aminoacyl],E [exit] • The first step starts with the insertion of aa-tRNA into the empty A site • The initiator tRNA is located to the P site • aa-tRNA is brought to the A site by EF-Tu complexed with GTP • Following GTP hydrolysis ,EF-Tu.GDP leaves the ribosome,with aa-tRNA correctly placed at the A site Peptid bond formation • aa-tRNA is located to the A site,initiator tRNA is in P site • A peptid bound is formed,resulting in the transfer of formyl- methionine to the aa-tRNA at the A site • This reaction is catalyzed by peptidyltransferase,is located in the 50S • Dipeptidyl-tRNA occupies the A site,an uncharged tRNA occupies P site Elongation : Peptide Bond Synthesis [peptide bond formed and growing peptide moves from P-site to A-site] The next step is translocation • Three movements occur • The uncharged tRNA leaves the P site • The peptidyl-tRNA moves from A site to the P site • The ribosome moves three nucleotides along the mRNA • This process requires EF-G (translocase) and GTP 50S Peptidyl transferase : A ribozyme activity • 23S RNA-catalyzed peptide bond format Termination • Termination codons are UAA;UAG;UGA • Aa-tRNA does not bind to the A site if the codons are stops • These codons are recognized by release factor • RF-1 recognized UAA or UAG • RF-2 recognized UAA or UGA Termination: • Protein release factor(s) recognizes a stop codon. • Stimulates release of new protein Antibiotic inhibitors of protein synthesis Protein synthesis in eucaryotes • İt is very complex process • Protein synthesis requires,ATP,GTP,tRNAs,mRNAs,aminoacids,a minoacyltRNA-synthetases,two sets of enzymes • One set,is for the process of initiation,the other for elongation and release of the nascent of peptid chain Protein synthesis in Eucaryotes • İnitiation factors[10 different factors] • eIF-1,eIF-1A,eIF-2,eIF-2B,eIF-3,eIF-4A,eIF4B,eIF-4E,eIF-4G,eIF-5 • Elongation factors • eEF-1alfa,eEF-1Beta,gama,eEF-2 • Termination factors • eRF-1,eRF-3 Ribosomes • 80S ribosome consists of 60S and 40S subunits • 60S contains 50 proteins,and 3 rRNA molecules[5s rRNA,28SrRNA,5.8S rRNA] • 40S contains 33 proteins and28S rRNA molecule initiation Eukaryotic Initiation complex EIF-2(GTP) for start AUG only EIF2-GDP + Pi Elongation 1) Ribosome binds to cap 2) Moves to 1st AUG 3) Large + small subunits associate • • • • eIF-3and eIF-1A,eIF-1,eIF-5 bind to the 40S eIF-2 binds initiator tRNA and GTP the mRNA binds eIF-4E ( binds to the 5 cap) eIF-4G binds to the eIF-4E and PABP (poly A binding protein) • eIF-4A and 4B bring mRNA to 40S. • Ribosome than scans down the mRNA to identify AUG initiation codon.eIF-5, catalizes the binding of the 60S to the 40Sinitiation complex. • Scanning requires energy [ATP] • When AUG is identified eIF-5 triggers the hydrolysis of GTP bound to eIF-2 • eIF-2 .GDP and other factors are released • 60S ribosomal subunit then joins the 40S complex • Elongation and termination steps are similar to procaryotes Elongation • The ribosome has three binding sites(P,A,E) • İnitiator tRNA is located P site,aa-TRNA is then brought to the A site by eEF-1alfa (complexed with GTP) • Following GTP hydrolysis ,EF-1alfa.GDP leaves the ribosome • A peptide bound is then formed • -ribosome moves three nucleotides along the mRNA. • This movement translocates the peptidyltRNA to the P site • The uncharged tRNA to the E site • -empty A site ready for adition to the next aatRNA • Translocation is mediated by eEF-2,coupled to GTP hydrolysis. termination • Eucaryotic cells contain only one relase factor (eRF) which recognize all three termination codons. • Release factor binds termination codon at the A site. • İt stimulates hydrolysis of the bond between the tRNA and the polypeptide chain at the P site. • The tRNa released,and the ribosomal subunits and the mRNA dissociate. Posttranslational Processing • Folding: chaperon protein; inclusion body • Secretion: signal sequence (20~25 AA); outer membrane block in E. coli • Exocytosis: constitutive vs. regulated • Glycosylation: addition of sugar; glycosylation pattern: location, degradation; glycoforms (glycosidase) • Phosphorylation