* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download A Figure S7. A. Standard curve of actin quantification using silver
Immunoprecipitation wikipedia , lookup
Structural alignment wikipedia , lookup
Rosetta@home wikipedia , lookup
Protein domain wikipedia , lookup
Homology modeling wikipedia , lookup
Intrinsically disordered proteins wikipedia , lookup
Protein design wikipedia , lookup
Protein folding wikipedia , lookup
Protein structure prediction wikipedia , lookup
Bimolecular fluorescence complementation wikipedia , lookup
Circular dichroism wikipedia , lookup
Protein mass spectrometry wikipedia , lookup
Gel electrophoresis wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Protein purification wikipedia , lookup
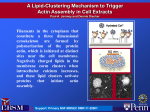
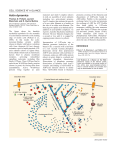
Yap et al - Fig. S7 A B C NAC1-V5 input N3ICD-V5 input 75 kDa 75 kDa Actin in supernatant 42 kDa Figure S7. A. Standard curve of actin quantification using silver staining. Actin standards were prepared by serial dilution and separated using SDS gel electrophoresis. Silver staining was carried out and band quantification was accomplished using the BioRad QuantityOne software. Linear regression revealed that the r 2 value of the curve is 0.9767. B. NAC1 and profilin-1 (PFN1) do not demonstrate a saturable curve in a quantitative pull-down assay. C. Notch3-ICD-V5 protein demonstrated negligible binding to actin at similar molar concentrations to NAC1. This V5 protein with a similar molecular weight as NAC1 was utilized as a low affinity control to demonstrate the specificity of actin for the NAC1-V5 protein.