* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Variant prioritization in NGS studies: Candidate gene prioritization
Genomic imprinting wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Genome evolution wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Gene desert wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Gene therapy wikipedia , lookup
Gene expression programming wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Gene nomenclature wikipedia , lookup
Neuronal ceroid lipofuscinosis wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Genome (book) wikipedia , lookup
Gene expression profiling wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Public health genomics wikipedia , lookup
Microevolution wikipedia , lookup
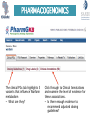
Variant prioritization in NGS studies: Candidate gene prioritization (Practical) " Work through the following practical exercises on your own." " The objective of these exercises is to become familiar with the concepts of variant and candidate gene prioritization." CANDIDATE GENE PRIORITIZATION" We will now look at prioritizing your list of interesting variants further. As a biologist, what would the next logical question be?" " • Ordinarily you would extract the list of gene symbols from this file and examine all of them for possible links to the disease of interest." " • For this example, use the following list of genes from your interesting variants file & pick one or two others that you think are good candidates:" • MYO5C; CYP2C9; TTN; F5; CCDC141! CANDIDATE GENE PRIORITIZATION" For each of these “candidate” genes:" • Use OMIM to get a broad idea of their function & what diseases they might be involved in (if any)" • Use Phenolyzer to see what phenotypes are associated with mouse/rat knockout models?" • Use BioGPS to see which human tissues these genes are expressed in?" • Use KEGG to see if these genes are involved in any pathways which may be linked to our disease features?" Now, do you have any very strong candidates related to your disease?" VARIANT PRIORITIZATION" Go back to your file of interesting variants and find the variant/s from your strong candidate gene." (If you didn’t find any strong candidates, have a close look at the F5 gene!)" " • Is it a known SNP? (does it have a dbSNP rsID?)" • Use SNPedia to look for any known phenotypes associated with this variant." • Is the “correct” allele seen in our patient? i.e. is it concordant with the phenotype?" • Does the text highlight any allele-allele or gene-gene interactions that might be of interest? If so, go back to your unfiltered annotated .vcf to see if they might be present" PHARMACOGENOMICS" So by now you know your patient has the factor V Leiden mutation which is strongly associated with Thrombosis. Typically, treatment for Thrombosis involves the prescription of blood thinners such as Warfarin. As a clinician you are aware of PGx associations with Warfarin metabolism so you want to see if your patient has any of the associated variants." " • Go to PharmGKB and search for “Warfarin”" PHARMACOGENOMICS" The clinical PGx tab highlights 3 variants that influence Warfarin metabolism" • What are they?" Click through to Clinical Annotations and examine the level of evidence for these associations." • Is there enough evidence to recommend adjusted dosing guidelines?" PHARMACOGENOMICS" Go back to your original unfiltered, annotated .vcf file to see if any of these variants are present." " • Why would you check the unfiltered file rather than your filtered file of interesting variants?" " • In conjunction with the information provided by PharmGKB, would you adjust your patients Warfarin dosage prescriptions?"