* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download tnf-alpha stimulated activation of mmp
Survey
Document related concepts
Protein moonlighting wikipedia , lookup
Histone acetylation and deacetylation wikipedia , lookup
Cell culture wikipedia , lookup
Cell-penetrating peptide wikipedia , lookup
Gene expression profiling wikipedia , lookup
Gene expression wikipedia , lookup
Promoter (genetics) wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Secreted frizzled-related protein 1 wikipedia , lookup
Transcriptional regulation wikipedia , lookup
Endogenous retrovirus wikipedia , lookup
Biochemical cascade wikipedia , lookup
List of types of proteins wikipedia , lookup
Gene regulatory network wikipedia , lookup
Proteases in angiogenesis wikipedia , lookup
Transcript
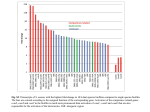
TNFα STIMULATED ACTIVATION OF MMP-2 IN NUCLEUS PULPOSUS CELLS OCCURS THROUGH EGR-1 MEDIATED TRANSCRIPTION OF MT1-MMP * Séguin, C A; *,**Pilliar, R M; +*,**Kandel, R A CIHR BioEngineering of Skeletal Tissues Team, *Mount Sinai Hospital, Toronto, Canada; ** Institute of Biomaterials and Biomedical Engineering, University of Toronto, Canada. [email protected] INTRODUCTION Degeneration of the intervertebral disc (IVD) involves a shift in the balance between catabolic and anabolic processes leading to changes in tissue architecture and function[1]. TNFα is a proinflammatory cytokine expressed within the nucleus pulposus (NP) of degenerate non-herniated IVDs, whose expression correlates with age and the degree of disc degeneration. Although implicated in the intitiation of discogenic pain, the role of TNFα in IVD degeneration remains poorly understood. We have developed a culture system that permits the formation of NP tissue in vitro, thus enabling investigations into the mechanisms regulating tissue degradation [2]. We have demonstrated that when exposed to low levels of TNFα, NP cells demonstrate increased gene expression of MMP-1, -3, -13 and ADAMTS-4 and -5, suggesting that TNFα stimulation of NP cells is a good model to study mechanisms regulating disc degeneration in vivo [3]. Among the enzymes known to be activated by TNFα, MMP-2 (gelatinase-A) is thought to contribute to the progression of degeneration and to the induction of neovascularization that occurs in the early stages of disc degeneration. In this study, we investigated if TNFα functionally activates MMP-2 and the regulatory mechanisms underlying TNFα-mediated MMP-2 activation in NP cells. MATERIALS AND METHODS NP Tissue Formation: The NP was dissected from bovine caudal spines and NP cells were isolated as previously described[2]. NP cultures were maintained for 2 wk to allow tissue formation similar to the native NP. NP Tissue Treatments: Cultures were treated with TNFα (25 ng/ml, Sigma) for 48 h in the presence of mitogen activated protein kinase (MAPK) inhibitors SB220225 (0-20 uM), SP600250 (0-20 uM), PD98059 (0-50 uM) or U0126 (0-10 uM)(Calbiochem). Gelatin Zymography: Conditioned medium was separated on 7.5% gels containing 0.1% gelatin by SDS-PAGE. Gels were incubated in renaturing buffer (30 min) and overnight in developing buffer. Gelatinase activity was visualized by Coomasie blue staining. RT-PCR Analysis: Total RNA was isolated from in vitro-formed tissues by Trizol extraction, reverse-transcribed (Superscript II, Invitrogen), and relative gene expression was examined by PCR. Immunoblot Analysis: Cell lysates were isolated from NP tissues following disruption of tissues by Dounce homogenization in RIPA buffer. 25 ug of protein was separated on 7.5% SDS-PAGE, electroblotted to nitrocellulose, and probed with antibodies reactive with ERK1/2 or Egr-1 (Cell Signaling), MMP-2 or MT1-MMP (Chemicon). Electrophoretic Mobility Shift Assays (EMSA): Nuclear extracts were prepared following disruption of tissues by Dounce homogenization. 10ug of protein were used in EMSA with radiolabelled oligonucleotides containing the Egr-1 consensus or bp -303 to -284 of the MT1-MMP promoter. For supershift reactions, antibodies against Egr-1 were included. Complexes were resolved by electrophoresis on 4% polyacrylamide gels and visualized by autoradiography. Transfections and Luciferase Reporter Analysis: Transient transfections of NP cells were performed using Fugene-6 (Roche) and 2ug of either pGL3-Basic vector or the MT1-MMP(-300)-promoter luciferase reporter construct [4] along with 0.1 ug pRL-SV40 to control for transfection efficiency. Luciferase expression was measured using the Dual Luciferase System (Promega). RESULTS Gelatinolytic activities corresponding to both latent and active forms of MMP-2 were present in the medium of TNFα-treated cultures by 48 h and were not present in untreated cultures. The role of MAPK pathways in the induction of MMP-2 activation was investigated using pharmacological inhibitors. Whereas inhibition of JNK or p38 did not alter MMP-2 activity, inhibition of ERK prevented the TNF -induced activation of MMP-2. As TNF functionally activated MMP-2, the effects of TNF on levels of MMP-2 and its potential regulators MT1-MMP, and TIMP-2 were determined. TNF up-regulated MT1-MMP gene expression in a timedependent manner. Changes in mRNA were associated with increased levels of MT1-MMP protein in NP cells following 24h. There were no significant changes in MMP-2 or TIMP-2 mRNA or protein levels. A GC-rich sequence in the MT1-MMP promoter containing consensus sites for Egr-1 can regulate gene expression [4]. In NP tissues, TNF induced a rapid and transient increase in Egr-1 mRNA by 0.5h which returned to baseline following 8h. A corresponding increase in levels of Egr-1 protein was detected, which peaked by 2h of TNF treatment and returned to basal levels by 24h. Selective blockade of the ERK pathway prevented TNF -induced Egr-1 gene expression suggesting Egr-1 as an effector protein in the ERK-mediated MMP-2 activation. EMSAs were conducted to assess the DNA-binding activity of Egr-1 and complex formation on the MT1-MMP promoter. Activation of Egr-1 and binding to the MT1-MMP promoter were detected by 2h of TNF treatment and returned to constitutive levels by 8h. ERK inhibition prevented Egr-1 activation and complex formation at the MT1-MMP promoter. The complex on the MT1-MMP promoter was confirmed to contain Egr-1 by competition with excess unlabelled oligonucleotides, and supershift reactions containing monoclonal Egr-1 antibodies. Figure1:Analysis of NP cell gene expression following TNFα treatment Transcriptional activity of the MT1-MMP promoter was assessed in NP cells expressing the basal promoter sequence coupled to the luciferase reporter gene (MT1-MMP(-300)-promoter luciferase)[4]. As suggested by PCR analysis, TNFα induced a significant increase in transcriptional activation of the MT1-MMP promoter. In keeping with the regulatory role of Egr-1, transcriptional induction of MT1-MMP was completely prevented by selective ERK inhibition. DISCUSSION As MMP-2 is implicated in the pathogenesis of IVD degeneration, the current study determined the intracellular mechanisms responsible for the induction of MMP-2 activity in a TNFα-induced model of NP degeneration. We demonstrate that in NP cells, TNFα induced MMP-2 gelatinase activity is dependent on a transient ERK-dependent increase in Egr-1 production, which was followed by an increase in the production of MT1-MMP. We confirmed that up-regulation of MT1MMP gene expression was mediated by ERK MAPK and Egr-1 transcriptional regulation of the MT1-MMP promoter by luciferasereporter assays. These findings suggest that TNFα induces the activation of both MMP-2 and MT1-MMP in NP tissues in vitro and in this way may contribute to TNFα-induced NP tissue degradation. As regulation of these enzymes is important for maintenance of IVD homeostasis in vivo, understanding the mechanisms governing their activation may be essential to delineate and eventually prevent the process of IVD degeneration. REFERENCES: [1] Buckwalter. (1995) Spine 20(11) : 1307. [2] Séguin et al. (2004) Spine 29(12):1299. [3] Séguin et al. (2005) Spine (in press). [4] Haas et al. (1999) J.Biol.Chem.274(32):22679. ACKNOWLEDGEMENTS: Supported by the Arthritis Society of Canada and NSERC Canada. 52nd Annual Meeting of the Orthopaedic Research Society Paper No: 0012