* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Heterodimerization of the Two Motor Subunits of the Heterotrimeric
Histone acetylation and deacetylation wikipedia , lookup
Protein phosphorylation wikipedia , lookup
Protein (nutrient) wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Protein moonlighting wikipedia , lookup
Magnesium transporter wikipedia , lookup
List of types of proteins wikipedia , lookup
Homology modeling wikipedia , lookup
Protein structure prediction wikipedia , lookup
VLDL receptor wikipedia , lookup
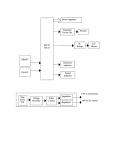
J. Mol. Biol. (1995) 252, 157–162 COMMUNICATION Heterodimerization of the Two Motor Subunits of the Heterotrimeric Kinesin, KRP85/95 Dana J. Rashid, Karen P. Wedaman and Jonathan M. Scholey* Section of Molecular and Cellular Biology, Division of Biological Sciences University of California Davis, CA 95616, USA The heterotrimeric kinesin-related motor protein, KRP85/95 is assembled from two kinesin-related polypeptides, SpKRP85 and SpKRP95, together with an uncharacterized 115 kDa polypeptide. Here we report the deduced amino acid sequence of SpKRP95, a close relative of SpKRP85. Both SpKRP85 and SpKRP95 are predicted to have a tripartite domain organization consisting of an N-terminal motor domain, a central stalk domain capable of coiled-coil formation, and a second globular C-terminal domain. The sequences of the central stalk domains predict that SpKRP85 and SpKRP95 should be capable of forming heterodimeric coiled coils. Furthermore, SpKRP85-SpKRP95 complexes can be immunoprecipitated from a cell-free translation system, providing direct evidence that SpKRP85 and SpKRP95 are capable of heterodimerization. 7 1995 Academic Press Limited *Corresponding author Keywords: heterotrimeric kinesin; KRP85/95 ; motor protein The heterotrimeric kinesin-related motor protein, KRP85/95, is a member of the kinesin superfamily of motor proteins (Bloom & Endow, 1994) that hydrolyzes ATP and transports cargo towards the plus ends of microtubules at approximately 0.4 mm per second (Cole et al., 1992, 1993). Two of the three subunits of this trimeric protein, SpKRP85 (Mr = 85 kDa) and SpKRP95 (Mr = 95 kDa), are thought to function as kinesin-related motor polypeptides, whereas a third 115 kDa subunit is thought to be a non-motor accessory polypeptide, possibly analogous to the kinesin light chains (Cole et al., 1993). KRP85/95 has recently been immunolocalized to detergent-sensitive particles in the metaphase half spindles and anaphase interzone of sea urchin embryonic mitotic apparatuses, leading us to hypothesize that this motor protein transports vesicles that deliver new membrane to the developing cleavage furrow of dividing cells (Henson et al., 1995). A novel feature of KRP85/95 is the apparent presence of two distinct kinesin-related polypeptides in a single holoenzyme (Cole et al., 1992, 1993), raising questions concerning the molecular architecture of KRP85/95. For example, we hypothesize that SpKRP85 and SpKRP95 could bind together within the KRP85/95 holoenzyme, perhaps by forming heterodimeric coiled coils analogous to the homodAbbreviations used: SpKRP, Strongylocentrotus purpuratus kinesin related polypeptide; mAb, monoclonal antibody. 0022–2836/95/370157–06 $12.00/0 imeric coiled coil that binds the two identical motor polypeptides of conventional kinesin (deCuevas et al., 1992). However, it is also possible that SpKRP85 and SpKRP95 do not physically interact, but are held together in the heterotrimeric complex by the 115 kDa subunit acting as a linker protein. Here we report the cloning and sequencing of a cDNA encoding full length SpKRP95 and we describe sequence analysis and co-immunoprecipitation experiments which support the hypothesis that SpKRP85 and SpKRP95 are indeed capable of heterodimerization. A full length SpKRP85 cDNA and a partial cDNA encoding approximately 40% of SpKRP95 were previously described (Cole et al., 1993). The partial SpKRP95 cDNA was used as a probe to obtain a full length cDNA by plaque hybridization screening of an unfertilized sea urchin egg cDNA library. The amino acid sequence of SpKRP95, deduced from the nucleotide sequence of the full length clone, predicts a kinesin-related polypeptide of 742 amino acids (Figure 1(a)) with a molecular weight of 84 kDa. The SpKRP95 sequence displays 51.6% identity to that of SpKRP85, but SpKRP95 is 43 residues longer than SpKRP85 (699 amino acids long, predicted Mr = 79 kDa) with the extra residues located at the C terminus of SpKRP95. Like SpKRP85, SpKRP95 has a tripartite structure consisting of a globular N-terminal motor domain (residues 1 to 356, predicted pI = 9.9), a central stalk domain with a high probability of a-helical structure (residues 357 to 588, predicted pI = 5.04), and a second globular 7 1995 Academic Press Limited (a) (b) Figure 1. (a) Deduced amino acid sequence of full length SpKRP95. A l ZAP Strongylocentrotus purpuratus unfertilized egg cDNA library was screened with a partial SpKRP95 cDNA fragment (Cole et al., 1993). The resulting 3.1 kb clone was sequenced and found to contain an open reading frame encoding 742 amino acids. The X in position 668 is resolved as R. This peptide sequence, and the corresponding nucleotide sequence, is available from GenBank under accession number U00996. Library screening and sequencing were performed by standard methods (Wedaman et al., 1993). (b) Dot matrix comparisons between SpKRP95 and closely related kinesin-related polypeptides. The full length SpKRP95 amino acid sequence (vertical axes) was compared to the full length sequences of S. purpuratus SpKRP85, murine Kif3A, Drosophila Klp68D, Chlamydomonas KHP1, Caenorhabditis elegans Osm-3 and S. purpuratus kinesin heavy chain (SpKHC) (horizontal axes). The plots were generated from the Wisconsin GCG dotplot program, using a window size of 8 and stringency 7. 159 Communication Table 1. Percent identities between SpKRP95 and related motor proteins SpKRP95 Full Motor SpKRP95 SpKRP85 Kif3A KHP1 Klp68D Osm3 SpKRP85 Full Motor Kif3A Full Motor KHP1 Full Motor Klp68D Full Motor Osm3 Full Motor SpKHC Full Motor 51.6 50.7 72.5 47.8 48.2 49.5 47.2 41.6 41.9 38.8 41.0 42.1 39.4 41.3 39.1 33.8 33.6 35.1 35.8 30.6 33.2 72.8 70.1 80.5 68.5 66.1 65.2 60.0 57.2 57.2 54.4 60.7 58.8 55.5 59.5 52.0 46.0 44.9 44.7 43.4 44.3 43.2 The full length or motor region sequences of S. purpuratus KRP95, S. purpuratus KRP85, mouse Kif3A, Chlamydomonas KHP1, Drosophila Klp68D, C. elegans Osm-3, and S. purpuratus conventional kinesin heavy chain are compared. region at the C terminus (residues 589 to 742, predicted pI = 9.62). The pI values of the corresponding domains in SpKRP85 and SpKRP95 are almost identical, but the motor domains of SpKRP95 and SpKRP85 are considerably more basic than the kinesin motor domain (pI = 8). The heterotrimeric kinesin, KRP85/95, was first discovered and purified from sea urchin eggs by Cole et al. (1992), but it is now apparent that its two motor subunits, SpKRP85 and SpKRP95 (Cole et al., 1993; this report) are members of a kinesin subfamily referred to as the kinesin II or Kif3 subfamily (Bloom & Endow, 1994; Goldstein, 1993). Close relatives of SpKRP85 and SpKRP95 in other organisms (Figure 1(b); Table 1) include murine Kif3A (Aizawa et al., 1992; Kondo et al., 1994), Drosophila Klp68D and Klp64D (Pesavento et al., 1994), the Chlamydomonas Fla10 gene product, KHP1 (Walther et al., 1994), and Caenorhabditis elegans Osm-3 (Shakir et al., 1993; Tabish et al., 1995). The motor domains of SpKRP85 and SpKRP95 display approximately 60% or greater sequence identity to the motor domains of these close relatives, compared to approximately 45% sequence identity with the motor domain of sea urchin kinesin heavy chain (SpKHC; Table 1). In addition there is significant sequence identity outside the motor domains of SpKRP85, SpKRP95, Kif3A, Klp68D and KHP1 but not Osm-3 or SpKHC (Figure 1(b)). Thus SpKRP95 is more closely related to mouse Kif3A, fruitfly Klp68D and Chlamydomonas KHP1 than it is to conventional kinesin heavy chain from the same organism. The remarkably high sequence conservation between SpKRP85 and Kif3A suggests that these two proteins are true homologs, and the recently described Kif3B, whose sequence is not yet available, may be a homolog of SpKRP95 (Yamazaki et al., 1994; see also Pesavento et al., 1994). The amino acid sequences of SpKRP85 and SpKRP95 were analyzed in order to evaluate their potential for heterodimerization. The algorithm of Lupas et al. (1991) revealed a high probability of coiled-coil formation extending approximately between residues 340 and 590 of SpKRP85 and SpKRP95, corresponding to the stalk domain (Figure 2(a)). Visual inspection and helical wheel analysis of the sequence itself reveal obvious heptad repeats indicative of coiled-coil formation between residues 354 to 588 of SpKRP95 and residues 357 to 594 of SpKRP85, ending in a conserved helixbreaking FIP-like sequence. This region could heterodimerize to form a coiled coil of approximately 235 residues. Assuming a residue repeat of 0.15 nm, this predicts a 35 nm long rod separating the head from the tail in the heterotrimeric KRP85,95 holoenzyme, which is considerably shorter than the 50 to 70 nm stalk of kinesin. The aforementioned helical wheel analysis revealed that SpKRP85/SpKRP95 heterodimers would display a hydrophobicity pattern that is typical of coiled coil proteins. This pattern is consistent with the formation of one set of hydrophobic bonds between the A positions of SpKRP85 and SpKRP95, and a second set of hydrophobic bonds between the D positions of the two peptides, that would form a hydrophobic core throughout virtually the entire length of the stalk of an SpKRP85/SpKRP95 heterodimer (Figure 2(b)). Interestingly, however, the glycine and proline-rich regions corresponding to the first major discontinuity predicted by the Lupas et al. algorithm in both SpKRP85 and SpKRP95 (asterisks in Figure 2(a)) contain stretches of charged residues that are oppositely charged in corresponding positions of SpKRP85 and SpKRP95 (Figure 2(c)). These regions would be expected to give rise to attractive electrostatic forces within SpKRP85/ SpKRP95 heterodimers and repulsive electrostatic forces within SpKRP85/SpKRP85 or SpKRP95/ SpKRP95 homodimers. Therefore these residues may play a role in stabilizing heterodimers relative to homodimers. In addition, unfavourable electrostatic interactions between residues of like charge in the A and D positions themselves (Figure 2(b)) should preferentially destabilize homodimers. To test the hypothesis that SpKRP85 and SpKRP95 are indeed able to form heterodimers in vitro, we used the cell-free translation system that has been used extensively to examine the heterodimerization of the cellular oncoproteins, fos and jun (Ransone et al., 1989). Thus, cDNAs encoding SpKRP85 and SpKRP95 were transcribed and translated in a rabbit reticulocyte lysate, and an antibody that recognized only the SpKRP95 polypeptide was tested for its ability to immunoprecipitate SpKRP85/SpKRP95 complexes (Figure 3). This was done by expressing a fusion protein consisting of full length SpKRP95 fused to the T7 gene 10 leader peptide recognizable by the T7 tag monoclonal antibody. A partial SpKRP85 construct, missing one-third of the motor domain, was expressed alone, lacking the gene 10 leader peptide. As expected, the T7 tag mAb was able to immunoprecipitate SpKRP95 (Figure 3 lane 4) but not SpKRP85 (Figure 3 lane 3) from lysates (a) (b) (c) Figure 2. Sequence analysis of SpKRP85 and SpKRP95. (a) Probability of a-helical coiled coil formation as predicted by the method of Lupas et al. (1991). The horizontal axes indicate residue number, and vertical axes represent the probability of coiled coil formation. The plots show a striking similarity in the predicted domain organization of SpKRP85 (bottom) and SpKRP95 (top). In particular, both peptides are predicted to form a similar sized coiled-coil, consistent with them forming a heterodimeric coiled-coil. The abundance of turns and proline residues, together with the predicted lack of extensive a-helical structure in the N-terminal heads and C-terminal tails, indicate globular ends on either side of the stalk domains. Asterisks indicate the glycine/proline-rich charged region that may favour heterodimerization over homodimerization. (b) Hydrophobicity of A and D positions in the heptad repeats of the SpKRP85 and SpKRP95 stalk regions. Helical wheels were generated using residues 420 to 594 of SpKRP85 and residues 410 to 588 of SpKRP95 (downstream from the first discontinuity (a)). The helical wheels were adjusted for skip residues; there were two skips in the SpKRP85 sequence and one skip in SpKRP95. The A and D positions show a typical pattern of hydrophobicity found in coiled coil proteins; not every residue is hydrophobic, but there is ample opportunity for hydrophobic bonding between positions A to A and D to D within SpKRP85/SpKRP95 heterodimers. (c) Amino acid region of opposing charge. This region, 33 amino acids in length in both SpKRP85 and SpKRP95, lies within the stalk domains but is not predicted to form an a-helical coiled coil. This segment contains proline residues in both subunits, and may impede homodimerization and/or appropriately align the peptides for heterodimerization by ionic interactions between oppositely charged side-chains. Communication Figure 3. Heterodimerization of SpKRP85 and SpKRP95 in vitro. This autoradiograph depicts 35S-labelled proteins that were expressed in and immunoprecipitated from a rabbit reticulocyte lysate cell-free translation system. Lane 1 shows expression of a truncated SpKRP85 construct missing one-third of its motor domain, and lane 2 shows the expression of T7 tag-SpKRP95 fusion protein in the reticulocyte lysate. Lane 3, control showing that the T7 tag mAb does not immunoprecipitate any radiolabelled polypeptides from lysates expressing only SpKRP85 lacking the T7 tag, whereas lane 4 shows that the T7 tag mAb does immunoprecipitate the T7 tag-SpKRP95 fusion protein from lysates containing T7 tag-SpKRP95 fusion protein but no SpKRP85. Finally lane 5 shows that the T7 tag mAb immunoprecipitates both SpKRP85 and T7 tag-SpKRP95 from lysates containing both these polypeptides. As the SpKRP85 polypeptide was not fused to the T7 tag epitope, it could only have been immunoprecipitated by the T7 tag mAb as a result of its binding to SpKRP95 which was fused to the T7 tag. A truncated SpKRP85 construct was used in these experiments because SpKRP95 fragments tended to mask full length SpKRP85 on the autoradiograms. Methods: The coding sequences of the SpKRP85 and SpKRP95 cDNAs were amplified by PCR using pfu polymerase (Stratagene) from the pBluescript library and subcloned into either pGEM-3Z (Promega) or PRSET (Invitrogen) vectors. The SpKRP95 construct was subcloned into the PRSET A vector for expression as a fusion protein containing the T7 gene 10 leader peptide at its N terminus, which is recognised by the T7 tag antibody. The SpKRP85 construct was subcloned into pGEM-3Z for expression of a truncated SpKRP85 missing one-third of its motor domain, and lacking the T7 gene 10 leader peptide. In vitro transcription of the constructs was performed with the Promega Riboprobe kit following the manufacturer’s protocols. The RNA transcripts were translated in rabbit reticulocyte lysate containing [35S]L-methionine (Promega). For some experiments (not shown) the Promega TNT rabbit reticulocyte lysate expression system was used instead. Immunoprecipitations were carried out as previously 161 containing only SpKRP95 or SpKRP85, respectively. However when SpKRP85 and SpKRP95 were expressed then incubated together in a cell-free lysate, SpKRP85 was immunoprecipitated together with SpKRP95 by the T7 tag antibody (Figure 3 lane 5), suggesting that SpKRP85 must have bound to SpKRP95 in the lysate. This experiment provides direct evidence that SpKRP85 and SpKRP95 are capable of heterodimerization. Two types of control experiment were used to assess the specificity of heterodimerization in these cell-free lysates (data not shown). First we tested the ability of T7 tag-SpKRP85 and T7 tag-SpKRP95 fusion proteins to form complexes with radiolabelled sea urchin egg kinesin heavy chain (SpKHC) that could be immunoprecipitated with the T7 tag mAb. Visual inspection of autoradiograms and scintillation counting of the immunoprecipitates revealed only background levels of radioactivity in the precipitates, suggesting that SpKRP85 and SpKRP95 did not form heterodimers with SpKHC. Therefore SpKRP85 and SpKRP95 do not heterodimerize promiscuously with all kinesin-related polypeptides. Secondly, we investigated the possibility that SpKRP85 and SpKRP95 could homodimerize by testing the ability of unlabelled T7 tag-SpKRP85 or T7 tag-SpKRP95 fusion proteins to bind to radiolabelled SpKRP85 or SpKRP95, forming radioactive complexes that could be immunoprecipitated by the T7 tag mAb. Visual inspection of autoradiograms and scintillation counting of the immunoprecipitates revealed an increase in radioactivity above background consistent with homodimerization of SpKRP85 but not SpKRP95. However, the extent of SpKRP85 homodimerization was estimated to be at least fivefold lower than heterodimerization. These experiments are consistent with the idea that a small fraction of the 300 kDa KRP85/95 holoenzyme might consist of the 115 kDa subunit complexed with SpKRP85 homodimers, but the majority consists of the 115 kDa subunit complexed with SpKRP85/SpKRP95 heterodimers. Based on these and previous studies (Cole et al., 1992, 1993), it seems reasonable to propose that each KRP85/95 molecule consists of one 115 kDa subunit, one SpKRP85 subunit and one SpKRP95 subunit, but additional immunoelectron microscopy studies are being initiated to further test this hypothesis. Are the close relatives of SpKRP85 and SpKRP95 from other organisms also components of heterotrimeric complexes? Holoenzymes assembled from described, with minor modifications (Ransone et al., 1989). Briefly, constructs were expressed separately, then incubated together at 18°C. The lysates were then incubated with the T7 tag monoclonal antibody (Novogen) on ice, and the T7 tag antibody-conjugated peptides were precipitated using Pansorbin cells (Calbiochem), washed in RIPA buffer (50 mM NaCl, 25 mM Tris-HCl (pH 7.5), 0.5% (w/v) NP-40, and 0.1% (w/v) sodium deoxycholate), then resuspended in SDS-PAGE sample buffer. The samples were subsequently electrophoresed on 7.5% (w/v) SDS-PAGE. The acrylamide gels were Coomassie stained, dried, and autoradiographed. 162 the close relatives of SpKRP85 and SpKRP95 described in Table 1 have not yet been purified from native tissue so their subunit composition is unknown. However, in a recent independent study described in abstract form, mouse Kif3A and Kif3B were shown to heterodimerize in vitro, and they could be co-immunoprecipitated with a 115 kDa polypeptide from brain tissue (Yamazaki et al., 1994). In addition, Drosophila Klp64D (whose sequence is not yet published) and Klp68D are reported to have sequences that predict similar sized coiled coils, consistent with the idea that they too may form heterodimeric coiled coils (Pesavento et al., 1994). These results are consistent with the idea that the close relatives of SpKRP85 and SpKRP95 are all components of heterotrimeric complexes that may represent a new heterotrimeric subfamily of kinesins (Cole & Scholey, 1995). In summary, KRP85/95 is a heterotrimeric motor protein composed of two heterodimerized motor subunits, SpKRP85 and SpKRP95, and one 115 kDa subunit. Sequence analysis is consistent with the hypothesis that heterodimerization between SpKRP85 and SpKRP95 could occur by a-helical coiled coil formation within their stalk domains, with a 33 amino acid residue segment rich in glycine, proline and charged residues serving to appropriately align SpKRP85 and SpKRP95 and stabilize heterodimers relative to homodimers. Additional structural studies will be aimed at testing this hypothesis and analyzing the contribution of the 115 kDa subunit to the structure of the KRP85/95 holoenzyme. In a broader context, the ability of distinct kinesin-related polypeptides to hetero-oligomerize may represent a general mechanism by which cells generate diversity in their complement of kinesin motors. For example, hetero-oligomeric assemblies of kinesin-related polypeptides may have properties that differ from the corresponding homo-oligomers, and consequently the combinatorial assembly of different kinesin-related polypeptides into a variety of oligomeric states could broaden functional diversity within the kinesin family. Currently, many kinesin-related polypeptides have been sequenced (Bloom & Endow, 1994; Goldstein, 1993) but only conventional kinesin, the heterotrimeric kinesin KRP85/95, and the homotetrameric kinesin KRP130 have been purified and characterized as native multimeric holoenzymes (Cole & Scholey, 1995). It will now be interesting to determine if any of the kinesin-related polypeptides analyzed only at the primary structural level also form hetero-oligomers in their natural host cell. Acknowledgements We thank members of the Scholey laboratory, Scott Chinn and Dr Sharyn Endow for helpful discussion. This work Communication was supported by NIH grants no. GM46376 and no. GM50718, and no. BE-46 from the American Cancer Society. References Aizawa, H., Sekine, Y., Tekamura, R., Zhang, Z., Nangaku, M. & Hirokawa, N. (1992). Kinesin family in murine central nervous system. J. Cell Biol. 119, 1287–1296. Bloom, G. S. & Endow, S. (1994). Kinesins. Protein Profile, 1, 1059–1116. Cole, D. G. & Scholey, J. M. (1995). Structural variations among the kinesins. Trends Cell Biol. 5, 259–262. Cole, D. G., Cande, W. Z., Baskin, R. J., Skoufias, D. A., Hogan, C. J. & Scholey, J. M. (1992). Isolation of a sea urchin egg kinesin-related protein using peptide antibodies. J. Cell Sci. 101, 291–301. Cole, D. G., Chinn, S. W., Wedaman, K. P., Hall, K., Vuong, T. & Scholey, J. M. (1993). Novel heterotrimeric kinesin-related protein purified from sea urchin eggs. Nature, 366, 268–270. de Cuevas, M., Tao, T. & Goldstein, L. S. (1992). Evidence that the stalk of Drosophila kinesin heavy chain is an alpha-helical coiled coil. J. Cell Biol. 116, 957–965. Goldstein, L. S. B. (1993). With apologies to Sheherazade: tails of 1001 kinesin motors. Annu. Rev. Genet. 27, 319–351. Henson, J. H., Cole, D. G., Terasaki, M., Rashid, D. & Scholey, J. M. (1995). Immunolocalization of the heterotrimeric kinesin related protein KRP85,95 in the mitotic apparatus of sea urchin embryos. Dev. Biol. In the press. Kondo, S., Sato-Yoshitake, R., Noda, Y., Aizawa, H., Nakata, T., Matsuura, Y. & Hirokawa, N. (1994). Kif3A is a new microtubule-based motor in the nerve axon. J. Cell Biol. 125, 1095–1107. Lupas, A., Van Dyke, M. & Stock, J. (1991). Predicting coiled coils from protein sequences. Science, 252, 1162–1164. Pesavento, P. A., Stewart, R. J. & Goldstein, L. S. (1994). Characterization of the Klp68D kinesin-like protein in Drosophila: possible roles in axonal transport. J. Cell Biol. 127, 1041–1048. Ransone, L., Visvader, J., Sassone-Corsi, P. & Verma, I, M. (1989). Fos-jun interaction: mutational analysis of the leucine zipper domain of both proteins. Genes Dev. 3, 770–781. Shakir, M., Fukushige, T., Yasuda, H., Miwa, J. & Siddiqui, S. S. (1993). C. elegans osm-3 gene mediating osmotic avoidance behaviour encodes a kinesin-like protein. Neuroreport, 4, 891–894. Tabish, M., Siddiqui, Z. K., Nishikawa, K. & Siddiqui, S. S. (1995). Exclusive expression of C. elegans Osm-3 kinesin gene in chemosensory neurons open to the external environment. J. Mol. Biol. 247, 377–389. Walther, Z., Vashishtha, M. & Hall, J. L. (1994). The Chlamydomonas Fla10 gene encodes a novel kinesinhomologous protein. J. Cell Biol. 126, 175–188. Wedaman, K. P., Knight, A. E., Kendrick-Jones, J. & Scholey, J. M. (1993). Sequences of sea urchin kinesin light chain isoforms. J. Mol. Biol. 231, 155–158. Yamazaki, H., Nakata, T., Okada, Y. & Hirokawa, N. (1994). Kif3B forms a heterodimer with Kif3A and works as a new microtubule-based anterograde motor of membrane organelle transport. Mol. Biol. Cell, 5, 32a. Edited by J. Karn (Received 17 May 1995; accepted 28 June 1995)