* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download restriction enzymes
Gene prediction wikipedia , lookup
Zinc finger nuclease wikipedia , lookup
Comparative genomic hybridization wikipedia , lookup
Genetic engineering wikipedia , lookup
Metagenomics wikipedia , lookup
Designer baby wikipedia , lookup
Gel electrophoresis of nucleic acids wikipedia , lookup
Non-coding DNA wikipedia , lookup
DNA supercoil wikipedia , lookup
Genome editing wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
DNA vaccination wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Restriction enzyme wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
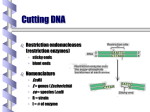
350 Home restriction enzymes R. Ward: Spring 2001 •Restriction Enzymes (endonucleases): molecular scissors that cut DNA •Properties of widely used Type II restriction enzymes: •recognize a single sequence of bases in dsDNA, usually symetrical (palindromic) •cleave both strands, generally within the recognition sequences •leaving “blunt” or “sticky” ends •leave a 3'-hydroxyl on one side of the cut and a 5'-phosphate on the other side of each strand 1 350 Home Examples of restriction enzymes R. Ward: Spring 2001 2 Palindromic sequences Axis of symmetry 5’ – NNNNNNNNNGGATCCNNNNNNNNNN – 3’ 3’ – NNNNNNNNNCCTAGGNNNNNNNNNN –5’ 5’ G 3’ C G C A T 350 Home T A R. Ward: Spring 2001 •v 3 350 Home R. Ward: Spring 2001 •Gene cloning = replication of a target sequence of DNA •insert target sequence into an easily replicated vector •insert the vector into a single bacteria (transformation) •allow the bacteria to amplify •vector has sequences that enable coordinated replication of the recombinant vector DNA •DNA Cloning: Vectors •Properties of useful vectors: •Vector DNA can be introduced into a bacteria •Vector contains a replication origin so it can replicate inside a host cell •vector contains antibiotic resistance gene or other selectable marker gene that allows easy identification of transformed bacterial colonies •Cloning vectors •Plasmid: insert DNA max size= 15 kb; autonomous replication; contains antibiotic resistance genes •Bacteriophage lambda: insert = 25 kb; recombinant DNA packaged into phage particles used to infect E. coli •BACs: bacteria artificial chromosomes insert size 100-500 kb •YACs: yeast articifical chromosomes 250-1000kb (1 mb) •cloning •generate fragments with a restriction enzyme. Sources can be whole DNA sample (genomic), or DNA generated from RNA of particular tissue •mix with linearized (restricted) plasmid (cut with same enzyme •ligate •get two products: plasmid with insert, plasmid only •have site in middle of gene whose function is disrupted upon insertion •bacterial amplification •transform bacteria with plasmid/insert construct •grow (plate) single cell dervied colonies on media carrying selectable agent (antiotic, usually) •grow single colony derived bacterial strains that survive the antibiotic and exhibit evidence of insertion •maintain as colony on plate or as frozen solution. “Library” is a collection of clones 4 350 Home R. Ward: Spring 2001 •Cloning vectors •Plasmid: insert DNA max size= 15 kb; autonomous replication; contains antibiotic resistance genes •Bacteriophage lambda: insert = 25 kb; recombinant DNA packaged into phage particles used to infect E. coli •BACs: bacteria artificial chromosomes insert size 100-500 kb •YACs: yeast articifical chromosomes 250-1000kb (1 mb) •cloning •generate fragments with a restriction enzyme. Sources can be whole DNA sample (genomic), or DNA generated from RNA of particular tissue •mix with linearized (restricted) plasmid (cut with same enzyme •ligate •get two products: plasmid with insert, plasmid only •have site in middle of gene whose function is disrupted upon insertion •transform bacteria with plasmid/insert construct •grow (plate) single cell dervied colonies on media carrying selectable agent (antiotic, usually) •grow single colony derived bacterial strains that survive the antibiotic and exhibit evidence of insertion •maintain as colony on plate or as frozen solution. “Library” is a collection of clones 5 350 Home R. Ward: Spring 2001 6 350 Home R. Ward: Spring 2001 7 350 Home R. Ward: Spring 2001 Main point on BACs and YACs is the large size insert they can carry 8 350 Home R. Ward: Spring 2001 Purifying insert involves separating plasmid from bacterial chromosomal DNA, followed by restriction with original cloning enzyme, then separation of plasmid (or other vector) from insert Methods of separation include electrophoresis You can also use PCR to amplify the insert directly from bacterial colonies or purified vectors by using the DNA sequence info flanking the vector cloning site to design forward and reverse primers. 9 Libraries: Collections of DNA clones 350 Home vGenomic libraries are made from “genomic” DNA directly from the plant’s DNA vcDNA libraries are made from DNA fragments that are created with reverse transcriptase from mRNA isolated from the plant vComplete libraries hypothetically contain one or more copies of every sequence from the source tissue. vLibraries maintained as bacterial or yeast colonies R. Ward: Spring 2001 10 350 Home Subcloning R. Ward: Spring 2001 Lambda (Phage) sub-libraries of individual YAC or BAC inserts enable manipulation of 20kb or so subsequences from that YAC/BAC insert Each sub-library can then be sub-cloned further into plasmids for even higher resolution. 11 350 Home cDNA R. Ward: Spring 2001 Big message here is that mRNA can be converted into DNA, called cDNA cDNA inserts do not have promoter or introns from the original genomic locus that the mRNA came from 12