* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download MD simulering
SNARE (protein) wikipedia , lookup
Multi-state modeling of biomolecules wikipedia , lookup
Mechanosensitive channels wikipedia , lookup
Implicit solvation wikipedia , lookup
Cell membrane wikipedia , lookup
List of types of proteins wikipedia , lookup
Endomembrane system wikipedia , lookup
Theories of general anaesthetic action wikipedia , lookup
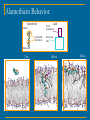
Peptide Aggregation and Pore Formation in a Lipid Bilayer; a Combined CG and AA MD Study Lea Thøgersen, University of Aarhus Pushing the Boundaries of Biomolecular Simulation June 11, 2008 Outline Introduction Results Conclusions Coarse-Grained Molecular Dynamics Alamethicin Modelling Setup NAMD CG vs MARTINI CG Microsecond Action Structure Reverse Coarse-Graining Water Pore Formation Structural Changes Coarse Grained MD – Why? Coarse-graining All-atom MD: time step of 1-2 fs, time frame sampled ~ 100 ns • Reduction in degrees of freedom • Fast frequenzy movements removed • Smoother potential surface • Longer time steps can be taken • Microsecond simulations possible Coarse-grained MD: time step of 20-50 fs, time frame sampled ~ 1 μs Coarse Grained Molecular Dynamics Shelley, Shelley, Reeder, Bandyopadhyay, Klein; A Coarse Grain Model for Phospholipid Simulations J. Phys. Chem. B (2001) 105 4464 Marrink, de Vries, Mark; Coarse Grained Model for Semiquantitative Lipid Simulations J. Phys. Chem. B (2004) 108 750 Shih, Arkhipov, Freddolino, Schulten; Coarse Grained Protein-Lipid Model with Application to Lipoprotein Particles J. Phys. Chem. B (2006) 110 3674 NAMD CG MARTINI CG Marrink, Risselada, Yefimov, Tieleman, de Vries; Monticelli, Kandasamy, Periole, Larson, Tieleman, Marrink; The MARTINI Force Field: Coarse Grained Model for Biomolecular Simulations The MARTINI Coarse-Grained Force Field: Extension to Proteins J. Phys. Chem. B (2007) 111 7812 J. Chem. Theory and Comput. (2008) 4 819 Alamethicin 20 amino acid antimicrobiel peptide. Part of the immune system (for fungi). Forms channels in membranes which allow water and ions to go through. Destroys membrane potential. Gln19 Glu18 Motivation: Potentially a good and simple membrane channel model Insight into this family of proteins could be valuable in the development of antibiotics Gln7 Widely Accepted Channel Model Tieleman, Hess, Sansom; Analysis and Evaluation of Channel Models: Simulations of Alamethicin. Biophys. J. (2002) 83 2392 Spaar, Münster, Salditt; Conformation of Peptides in Lipid Membranes Studied by X-Ray Grazing Incidence Scattering. Biophys. J. (2004) 87 396 Modelling Setup To study alamethicin interaction with membrane and each other 25 peptides 330 DMPC lipids 26452 water ~117000 atoms ~ 11700 beads PBC 120 Å × 124 Å × 90 Å NAMD CG versus MARTINI CG 4 GLY ALA 4 AIB Apolar 1 2 degree of polarity 1-5 1 Polar 2 4 Nda degree of polarity 1-5 C C NAMD MARTINI C (MARTINI) Nda NAMD CG Nda AA N0 2 MARTINI CG Nonpolar donor hydrogenbond acceptor both C5 none C4 (NAMD) Charged 1 donor hydrogenbond acceptor both none Alamethicin Behavior Alamethicin C-term Hydrophilic sidechains Lipid Polar headgroup Non-polar tails N-term 7 ns 120 ns 32 ns Peptide Aggregation I 0-1 μs Peptide Aggregation II MARTINI NAMD 0 μs 1 μs 4 μs Structure of the Clusters 0 0.5 Gln7 1 Gln7 Helix Tilt Alamethicin, DMPC lipid peptid:lipid - 1:15 15N-Aib8 alamethicin θ Aib8 Exp θ = 10˚ 0.04 MD 0.03 0.02 0.01 0 0 30 60 90 Helix tilt / deg. 120 Vosegaard, Bertelsen, Pedersen, Thøgersen, Schiøtt, Tajkhorshid, Skrydstrup, Nielsen; Resolution Enhancement in Solid-State NMR of Oriented Membrane Proteins by Anisotropic Differential Linebroadening JACS (2008) 130 5028 Reverse Coarse-Graining AA t = 0μs CG t = 0μs CG t = 1μs AA - rev CG t = 1μs AA after SA CG water vs AA water δ- Class: Polar δ+ δ+ 5.0Å 900-1000 ns TIP3P model 2.8Å 35-40 ns Structural Changes 18 of 25 peptides remain α-helical Conclusions Alamethicin monomers readily aggregate and form clusters that grow in size over time. Large diversity in form of clusters and structure of peptides. Reverse CG required to obtain detailed water interaction validate results obtained on the long time scale Acknowledgements Emad Tajkhorshid & the NAMD people Theoretical and Computational Biophysics Group University of Illinois at Urbana-Champaign, USA Birgit Schiøtt & the Biomodelling Group Department of Chemistry University of Aarhus, Denmark Niels Christian Nielsen & Thomas Vosegaard Laboratory for Biomolecular NMR spectroscopy Department of Chemistry University of Aarhus, Denmark Funding: &