* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Abstract - UWL faculty websites
Neuronal ceroid lipofuscinosis wikipedia , lookup
Microevolution wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Gene expression profiling wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Designer baby wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Gene nomenclature wikipedia , lookup
Protein moonlighting wikipedia , lookup
Point mutation wikipedia , lookup
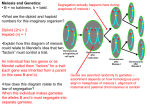
Abstract Saccharomyces cerevisiae or Baker’s yeast is a commonly used model organism for researchers who study cell division and the genes that are involved in its regulation. Several yeast cell division genes have human versions with putative roles in the formation of cancerous tumors. One of these cell division genes, CDC7, is also involved in meiotic division although its precise role in meiosis is unknown. One way to help elucidate CDC7’s role in yeast meiosis is to determine when Cdc7 protein is made using western analysis. In order to perform this analysis, antibodies are required that will specifically recognize the Cdc7 protein. Since none are yet available, yeast strains were constructed that contain the CDC7 gene tagged with a portion of the human myc gene. The tagged CDC7 gene has the same phenotype as the wild-type gene. Antibodies were then purchased that recognize the human myc portion of the Cdc7-myc protein. Experiments were done to determine the appropriate antibody concentrations necessary to detect the tagged Cdc7 protein from mitotic cells on immunoblots. This information will be used to perform a western analysis of protein obtained from meiotic yeast samples to determine when during meiosis the Cdc7 protein is expressed. Introduction One of the ways being investigated to understand the spread of cancer is to determine how proteins affect normal cell growth processes. Saccharomyces cerevisiae or Baker’s yeast is studied by thousands of researchers because yeast contains cell division proteins that are similar to those in human cells. One commonly studied yeast cell growth protein is Cdc7. This protein kinase is required for initiating DNA replication (S phase) during the mitotic cell cycle, although it is not understood how Cdc7 protein controls this initiation in yeast. Saccharomyces cerevisiae is a single-celled eukaryote that can exist as a diploid or as a haploid. Haploid yeast can be one of two forms: “a” or “.” These two forms can mate with each other to form a/ diploid cells. All of these cell types, a, , and a/ diploids, can grow mitotically, but only a/ diploids can undergo the cell division process of meiosis. Meiosis is a process similar but not identical to the mitotic cell cycle. It is worthwhile to examine the role of Cdc7 protein in meiosis to help understand its role in the mitotic cell cycle and possible cancer development. Currently, very little is known about what Cdc7 does in meiosis although it is known to be required for meiosis to occur properly in yeast. One way to examine the meiotic role of Cdc7 is to analyze Cdc7 protein levels throughout meiosis. We constructed a strain containing the CDC7 gene tagged with myc so that Cdc7 protein can be detected by western analysis using commercially available anti-myc antibodies. Results Two haploid temperature sensitive cdc7 mutant yeast strains were separately transformed with a plasmid containing the wild-type CDC7 gene with a portion of the human myc gene fused to its 3’ end (Figure 1). Transformants were selected on media containing uracil. These transformants contained the cdc7ts mutation, the URA3 gene that allows them to grow and be selected on media lacking uracil, and a second copy of CDC7 with the myc tag (Figure 1). The transformants were grown on YPD media which contains uracil, allowing the survival of any cells that removed one of the copies of CDC7 and the URA3 gene by recombination (Figure 1). If the crossover occurred to the left of cdc7ts as drawn in Figure 1, the resulting “popout” would contain the CDC7-myc gene, the desired genotype. If recombination occurred to the right of cdc7ts as depicted (Figure 1) the resulting “popout” would contain the cdc7ts mutation and would be identical to the original strain. This was not the desired genotype. In either case, popouts would lose the wild-type URA3 gene and would be unable to grow on media lacking uracil. 5-FOA media was used to select for the growth of these Ura- yeast popouts because the metabolic poison 5-FOA kills Ura+ cells and thus allows only Ura- cells to grow. To distinguish between the two possible genotypes, the popouts were tested to determine if they were cdc7ts or CDC7-myc by testing for temperature sensitivity. The cells were replica plated to rich YPD media and incubated at 22C or 30C. Popouts that are CDC7-myc, the desired genotype, are not temperature sensitive and should grow at both 22C (the permissive temperature) and 30C (the restrictive temperature). Popouts that are cdc7ts, the undesired genotype, should only grow at the permissive temperature of 22C. Table 1 shows that 8 out of 13 “a” popouts and 4 out of 4 “” popouts were no longer temperature sensitive, indicating that they contained the desired CDC7-myc construct. “a” and “” popouts were crossed together to form diploid strains. The haploid integrants, the haploid popouts, and the diploid strains were grown up in rich media and protein was isolated. The protein was loaded onto a polyacrylamide gel, subjected to electrophoresis, transferred to nitrocellulose membrane, and analysed by immunoblot (Figure 2A). The diploid strains were then induced to undergo meiosis by starvation. Samples were taken after 0, 15, and 16 hours of meiosis. Protein was isolated and immunoblotting was performed as described above. The results are shown in Figure 2B. cdc7ts cdc7ts URA3 CDC7-myc cdc7ts CDC7-myc cdc7ts CDC7-myc Figure 1. Construction of yeast strains containing CDC7 tagged with myc. A) A plasmid containing the CDC7-myc construct was transformed into a cdc7ts strain. This plasmid integrated into the yeast chromosome by recombination, resulting in a strain that had two copies of the CDC7 gene with an intervening URA3 gene. B) Crossing over then occurred spontaneously between the two copies of CDC7 resulting in the loss of the URA3 gene and one of the two CDC7 genes. If the crossover occurred on the left side of cdc7ts, as depicted, then the "popout" contains the desired CDC7-myc construct. Table 1. Results of testing "popouts" for temperature sensitivity.1 “a” popouts YPD 22C 1 2 3 4 5 6 7 8 9 10 11 12 13 + + + + + + + + + + + + + “” popouts YPD 22C 1 2 3 4 1 + + + + YPD 30C + + + + + + o o o o + + o YPD 30C + + + + A"+" indicates growth; a "o" indicates no growth. 1 6 7 2 7 4 5 8 1 6 3 2 3 4 5 8 Figure 2. Results of Cdc7 immunoblot. A) Yeast strains were grown mitotically. Protein was isolated and subjected to SDS-polyacrylamide gel electrophoresis. After transferring the protein to a nitrocellulose membrane, an immunoblot was performed using antibodies specific to the myc tag. Bands on the X-ray film represent the locations of the Cdc7-myc protein in each lane. Lane 1: negative control; Lane 2: positive control; Lane 3: haploid “a” integrant; Lane 4: haploid “a” popout; Lane 5: haploid “” integrant; Lane 6: haploid “” popout; Lane 7: diploid myc-containing strain #1; Lane 8: diploid myc-containing strain #2. The anti-myc antibody was specific to the Cdc7-myc protein; no other bands were observed on the X-ray film. B) Yeast strains were grown mitotically and then were induced to undergo meiosis by starvation. Protein was isolated and an immunoblot was performed using antibodies specific to the myc tag as described above. Lane 1: negative control; Lane 2: positive control; Lane 3: diploid #1 at 0 hrs.; Lane 4: diploid #1 at 15 hrs.; Lane 5: diploid #1 at 16 hrs.; Lane 6: diploid #2 at 0 hrs.; Lane 7: diploid #2 at 15 hrs.; Lane 8: diploid #2 at 16 hrs. The anti-myc antibody was specific to the Cdc7-myc protein. A protease is likely to be responsible for the extra bands observed in the lanes containing meiotic proteins (lanes 4, 5, 7, 8). Discussion Two haploid yeast strains, an “a” and an “”, were engineered to contain the CDC7 gene tagged with a portion of the human myc gene to enable us to detect Cdc7 protein in meiosis using commercially available anti-myc antibodies. A phenotypic assay confirmed the presence of this construct in both haploids. The “a” and “” CDC7-myc haploids were mated together to construct an a/ diploid that could undergo meiosis. This diploid was used to develop a protocol for western analysis so that the expression of the Cdc7 protein could be examined throughout meiosis by immunoblotting. Although meiotic Cdc7p is detectable by immunoblotting, it seems that there is a protease in the meiotic protein samples that is degrading the Cdc7 protein (See Figure 2B). When this experiment is repeated, protease inhibitors will be added to alleviate the problem. Future Work The immunoblot experiment shown in Figure 2B will be repeated using protease inhibitors. Once the degradation problem is resolved a large scale experiment will be set up in which samples will be collected each hour for 12 hours followed by a final time point at 24 hours. Protein will be isolated and an immunoblot will be performed to determine when during meiosis Cdc7 protein is being produced. Acknowledgements We would like to thank Dr. Anne Galbraith for her time and efforts in helping us with this research project. This work was funded by a UW-L Undergraduate Research Grant to Jacqueline Bjornton and a UW-L Faculty Research Grant to Dr. Anne Galbraith.