* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download How Immunity Evolved
Duffy antigen system wikipedia , lookup
Plant disease resistance wikipedia , lookup
Immunocontraception wikipedia , lookup
Lymphopoiesis wikipedia , lookup
Gluten immunochemistry wikipedia , lookup
Complement system wikipedia , lookup
Sociality and disease transmission wikipedia , lookup
DNA vaccination wikipedia , lookup
Herd immunity wikipedia , lookup
Hygiene hypothesis wikipedia , lookup
Social immunity wikipedia , lookup
Adoptive cell transfer wikipedia , lookup
Cancer immunotherapy wikipedia , lookup
Molecular mimicry wikipedia , lookup
Immune system wikipedia , lookup
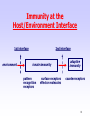
Polyclonal B cell response wikipedia , lookup
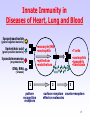
Adaptive immune system wikipedia , lookup
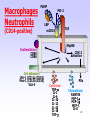
Immunosuppressive drug wikipedia , lookup
Innate Immunity in Diseases of Lung, Heart and Blood: The Idea behind the UA/BWH PGA Donata Vercelli Respiratory Sciences Center Department of Cell Biology and Anatomy College of Medicine, University of Arizona 1 How Immunity Evolved • The immune system evolved under selective pressure imposed by infectious microorganisms. • Multicellular organisms have developed various defense mechanisms that have the capacity to be triggered by infection and to protect the host organism by destroying the invading microbes and neutralizing their virulence factors. 2 How the Innate and Adaptive Immune Systems Recognize Pathogens • Immune responses can be innate or adaptive. • The innate immune system uses germlineencoded receptors for the recognition of microbial pathogens. • The adaptive immune system (found only in vertebrates) uses receptors generated by somatic mechanisms (gene rearrangements) during the ontogeny of each individual organism. 3 The Adaptive Immune System • Somatic mechanisms generate a diverse repertoire of antigen receptors with random specificities, which are clonally distributed on immune cells (T and B lymphocytes) plasticity • The specificity of the receptors expressed on each lymphocyte is not predetermined, and neither is the response that can be induced in lymphocytes upon ligation of their receptors by antigen ambiguity 4 The Cross-talk between Innate and Adaptive Immunity •Induction of an immune response is only appropriate if the antigen recognized is derived from, or belongs to, a pathogen. •Activation of lymphocytes specific for self antigens, or innocuous persistent environmental antigens, is deleterious. Therefore, •The adaptive immune response requires signals that provide information about the origin of the antigen and the type of response to be induced. These signals are provided by the innate immune system. 5 Mechanisms of Innate Immune Recognition 6 How Few Receptors Can Recognize Many Pathogens •Immune recognition is unique in that it occurs between products encoded in different genomes. •The evolution of innate immunity is driven toward recognition of invariant molecular constituents of the infectious agents shared by large groups of pathogens. Therefore A limited number of germline-encoded receptors is able to recognize a great variety of molecular structures associated with pathogens. 7 Receptors and Targets of Innate Recognition • The targets of recognition represent molecular patterns, called PAMPs for pathogenassociated molecular patterns, rather than particular structures. • Host organisms have developed a set of receptors that can specifically recognize PAMPs and are referred to, therefore, as pattern recognition receptors (PRRs). 8 Innate Discrimination between Self and Non-self is Perfect The effect of immune recognition is destruction of the target’ The structures selected for immune recognition must have been absolutely distinct from self antigens to avoid damage to self cells and tissues. Consequences: •The ability of the innate immune system to discriminate between self and nonself is perfect. None of the compounds recognized by the innate immune system are produced by the host organism, and all of them are essential for the physiology and survival of the respective microbes. 9 Pathogen-Associated Molecular Patterns (PAMPs) PAMPs represent invariant structures shared by large groups of microorganisms, and are absolutely essential for the microbe's survival: •LPS: common component of gram-negative bacteria •teichoic acids: common component of grampositive bacteria •double-stranded RNA: a structural signature of several groups of RNA viruses •bacterial DNA (unmethylated CpG sequences) 10 The Pattern Recognition Receptor System of Innate Immunity LPS MD-2 LPS LPS LBP LBP mCD14 TLR2/4 LBP sCD14 TLR2/4 RP105 (CD180) sCD14 MD-1 CD14-positive cells (monocyte/M, PMN) CD14-negative cells (e.g., endothelium, epithelium) CD14-negative cells (B cells) 11 TLRs: Not that non-specific... TLR4 LPS (Gram-) viruses? TLR2 Lipo proteins (Gram+) direct anti-microbial defense TLR6 PGN (w.TLR2) apoptosis of host cells TLR9 TLR5 flagellin DNA septic shock Listeria Pseudomonas adaptive responses 12 Pathogen Recognition and the Control of Adaptive Immunity • Antigen receptors expressed on lymphocytes have randomly generated specificities that cannot determine the origin or biological context of their ligands. • Signaling through an antigen receptor is insufficient on its own to induce the activation of lymphocytes or their differentiation into appropriate effector cells. 13 Costimulation, the Link between Innate and Adaptive Immunity •Activation of T lymphocytes requires a costimulatory second signal in the form of B7 molecules expressed on antigen presenting cells (APC). •The function of costimulatory activity is to signal the presence of pathogens. •The expression of costimulation must be controlled by pathogen recognition. 14 Innate Immunity Provides Cues for the Adaptive Response IL-4 IL-13 pathogen CD3 Signal 1 TCR MHC II PRR CD4 IL-4, IL-13 APC Th2 cell CD28 B7 Signal 2 cytokines 15 Innate and Adaptive Immunity Innate Adaptive Encoding of Receptors germline somatically Distribution of receptors non-clonal clonal Repertoire of receptors limited very large Target invariant variable Recognition perfect imperfect Speed fast slow Long-lasting memory no yes 16 The UA/BWH PGA inflammation MI DVT COPD asthma infection genetics 17 Immunity at the Host/Environment Interface 1st interface environment 2nd interface innate immunity pattern recognition receptors surface receptors effector molecules adaptive immunity counterreceptors 18 Innate Immunity in Diseases of Heart, Lung and Blood lipopolysaccharide (gram-negative bacteria) lipoteichoic acid (gram-positive bacteria) lipoarabinomannan (mycobacteria) DNA, RNA •monocyte/MØ •neutrophils •T cells •epithelium •endothelium •eosinophils •basophils •fibroblasts (viruses) 1 pattern recognition receptors 2 3 surface receptors counterreceptors effector molecules 19 Macrophages Neutrophils PAMP (CD14-positive) MD-2 LBP mCD14 Myd88 Costimulation: CD80 CD86 Cell adhesion: LFA-1 (CD11a/CD18) Mac-1 (CD11b/CD18) VLA-4 TLR COX-2 activation PGs Cytokines: TNF-a IL-1 IL-6 IL-10 IL-12 IL-23 IL-18 TGF-b Chemokines: RANTES MIP-1a MIP-1b MCP-1 IL-8 20 Endothelium (CD14-negative) PAMP MD-2 LBP TLR sCD14 Cell adhesion: ICAM-1 ICAM-2 ICAM-2 V-CAM1 E-selectin OX40L Cytokines: GM-CSF IL-1 IL-6 IL-18 TGF Chemokines: RANTES MCP-1 IL-8 21 Epithelium PAMP LBP (CD14-negative) EGFR MD-2 sCD14 TLR Mucus secretion MUC2 MUC5AC activation Cell adhesion ICAM-1 ICAM-2 ICAM-2 V-CAM1 Cytokines GM-CSF IL-1 Anti-microbial peptides IL-6 b-defensins IL-18 SP-A TGF-b SP-D NOS-2 activation NO Chemokines Eotaxin RANTES IL-8 22 Receptor/Ligand Pairs on Effector Cells EGF PAF EGFR PAFR Cytokines Costimulation Cell adhesion Chemokines IL-1R IL-6R IL-10R IL-12R IL-18R TGF-bR TNFR CCR2 CCR3 CCR5 CCR6 CXCR1 CXCR2 CD80/CD28 CD86/CD28 CD80/CTLA-4 CD86/CTLA-4 LFA-1/ICAM Mac-1/ICAM OX40/OX40L VLA-4/V-CAM 23 Where Should We Start? MD1, 2 LBP TLR2, 4, 5, 6, 9 RP105 CD14 Nod1 Cytokines IL-10 Myd88 Anti-microbial peptides b-defensin 1, 2 SP-A, SP-D … with prototypic innate immunity genes arranged along the PAMP/TLR signaling pathway in different cell types (including VERY recent additions) … Mucus Muc2 Muc5Ac 24