Slide 1
... Neither genes nor environment dominates development; rather there is continual interaction between genes and the environment, with both contributing to the phenotype. However, studies of twins have been used to determine the relative effects of genetic and environmental factors on the development of ...
... Neither genes nor environment dominates development; rather there is continual interaction between genes and the environment, with both contributing to the phenotype. However, studies of twins have been used to determine the relative effects of genetic and environmental factors on the development of ...
Nature Methods article on Programming transcription
... activity of genetically defined cells in the brain and observing how such perturbations affect an animal’s behavior. Several labs have generated transgenic mouse lines that express the light-sensitive proteins in a cell type–specific manner, but obtaining sufficiently high expression levels of the p ...
... activity of genetically defined cells in the brain and observing how such perturbations affect an animal’s behavior. Several labs have generated transgenic mouse lines that express the light-sensitive proteins in a cell type–specific manner, but obtaining sufficiently high expression levels of the p ...
CP_Chromosome 231111_EN
... caused by a phenomenon that, until now, was thought unlikely to occur in mammalian cells: interference between two key genetic processes, DNA transcription1 and replication2. This research, published in the journal Molecular Cell on 23 December 2011, could eventually lead to novel strategies for fig ...
... caused by a phenomenon that, until now, was thought unlikely to occur in mammalian cells: interference between two key genetic processes, DNA transcription1 and replication2. This research, published in the journal Molecular Cell on 23 December 2011, could eventually lead to novel strategies for fig ...
Chapter 16-17 review sheet
... complementary strand, daughter strand, parent strand, RNA primer, and DNA polymerase III, DNA polymerase I. ...
... complementary strand, daughter strand, parent strand, RNA primer, and DNA polymerase III, DNA polymerase I. ...
RNA polymerase II is the key enzyme in the process of
... dimerization partner. Describe then briefly the Myc-network of interacting partners including how Myc-responsive genes either may be activated or repressed depending on the presence of various members in this network. ...
... dimerization partner. Describe then briefly the Myc-network of interacting partners including how Myc-responsive genes either may be activated or repressed depending on the presence of various members in this network. ...
Transcript Maps
... • transcription factor (TF) General term for any protein, other than RNA polymerase, required to initiate or regulate transcription in eukaryotic cells. General factors, required for transcription of all genes, participate in formation of the transcription-initiation complex near the start site. Spe ...
... • transcription factor (TF) General term for any protein, other than RNA polymerase, required to initiate or regulate transcription in eukaryotic cells. General factors, required for transcription of all genes, participate in formation of the transcription-initiation complex near the start site. Spe ...
Transcription/Translation
... additional levels of control. When and how long a protein is active in the cytoplasm represents a post-translational level of control. ...
... additional levels of control. When and how long a protein is active in the cytoplasm represents a post-translational level of control. ...
Basics of Gene Expression Activity
... 13. Make an extension – using what you learned above, how could you dictate how much of a particular protein is made? Describe the “set-up” a cell might have in each case. a. To make a lot of a particular protein - _________________________________________________________ b. To make just a little - ...
... 13. Make an extension – using what you learned above, how could you dictate how much of a particular protein is made? Describe the “set-up” a cell might have in each case. a. To make a lot of a particular protein - _________________________________________________________ b. To make just a little - ...
Eukaryotic Gene Expression
... • Why eukaryotes are different: – Genes are nearly always transcribed individually – 3 RNA Polymerases occur, requiring multiple proteins to initiate transcription ...
... • Why eukaryotes are different: – Genes are nearly always transcribed individually – 3 RNA Polymerases occur, requiring multiple proteins to initiate transcription ...
BSC 219
... different from prokaryotic transcription initiation. Eukaryotic initiation involves a large number of proteins to form an initiation complex that recruits RNA Polymerase to the promoter region. The DNA sequences and some proteins in the complex are variable between promoters. Prokaryotic initiation ...
... different from prokaryotic transcription initiation. Eukaryotic initiation involves a large number of proteins to form an initiation complex that recruits RNA Polymerase to the promoter region. The DNA sequences and some proteins in the complex are variable between promoters. Prokaryotic initiation ...
Bio1100Ch19W
... These prevent cancer by repairing DNA or preventing rapid cell division If mutate a tumor suppressor gene- leads to ________ The Ras gene- (a protooncogene) is mutated in _____ of human ...
... These prevent cancer by repairing DNA or preventing rapid cell division If mutate a tumor suppressor gene- leads to ________ The Ras gene- (a protooncogene) is mutated in _____ of human ...
SBI 4U Genetics 5
... cell’s DNA and causes substitution or frameshift changes. EG. Gasoline fumes, nitrites and compounds found in cigarette smoke Physical mutagens: physically change the DNA ...
... cell’s DNA and causes substitution or frameshift changes. EG. Gasoline fumes, nitrites and compounds found in cigarette smoke Physical mutagens: physically change the DNA ...
Gene Regulation - Two Rivers High School
... Used for: Response to environment O Gene regulation helps a cell respond to its ...
... Used for: Response to environment O Gene regulation helps a cell respond to its ...
A comprehensive catalogue of human RNA-binding
... the team were able to map so-called connectivity quantitative trait loci (cQTLs). These cQTLs are natural genetic variants that influence the regulatory interactions of specific transcription factors and their target genes (for example, a polymorphism that occurs in the coding or promoter sequence o ...
... the team were able to map so-called connectivity quantitative trait loci (cQTLs). These cQTLs are natural genetic variants that influence the regulatory interactions of specific transcription factors and their target genes (for example, a polymorphism that occurs in the coding or promoter sequence o ...
Title
... c. Remain the same Why is the genetic code degenerate? a. Because the DNA is not precisely copied into RNA. b. Because more than one codon in a mRNA can code for a single amino acid. c. Because more than one amino acid can be specified by the same sequence in the mRNA. d. Because the genetic code wa ...
... c. Remain the same Why is the genetic code degenerate? a. Because the DNA is not precisely copied into RNA. b. Because more than one codon in a mRNA can code for a single amino acid. c. Because more than one amino acid can be specified by the same sequence in the mRNA. d. Because the genetic code wa ...
Distinguish between these 3 root types: - mvhs
... will be _________ from the cell. The mRNA for this protein contains a signal recognition sequence that is recognized by a signal recognition particle (SRP). The SRP brings the growing polypeptide to the receptor protein in the ___________________. ...
... will be _________ from the cell. The mRNA for this protein contains a signal recognition sequence that is recognized by a signal recognition particle (SRP). The SRP brings the growing polypeptide to the receptor protein in the ___________________. ...
REGULATION OF GENE EXPRESSION IN EUKARYOTES
... that you have never heard of • Implicated in – Cell proliferation – Cell differentiation – Spermatogenesis – Release of somatostatin (inhibitor growth hormone) – Development of T lymphocytes – Metabolism of the pineal gland – Adaptation to physical stress – Transcription of metabolic enzymes – Criti ...
... that you have never heard of • Implicated in – Cell proliferation – Cell differentiation – Spermatogenesis – Release of somatostatin (inhibitor growth hormone) – Development of T lymphocytes – Metabolism of the pineal gland – Adaptation to physical stress – Transcription of metabolic enzymes – Criti ...
17. Gene regulation
... Ligand binding activates receptor Recognition sites on DNA are similar Receptor--ligand interactions mostly occur in the cytoplasm Other transcription factors mediate environmental responses Chemical signal cAMP Serum growth factor Phorbol esters ...
... Ligand binding activates receptor Recognition sites on DNA are similar Receptor--ligand interactions mostly occur in the cytoplasm Other transcription factors mediate environmental responses Chemical signal cAMP Serum growth factor Phorbol esters ...
Chapter 17 - Denton ISD
... sections called _______, and leaving exons. Some genes can produce multiple polypeptides depending on what is spliced; this is called ___________________. Exon shuffling during cross-over may also be useful in evolution. ...
... sections called _______, and leaving exons. Some genes can produce multiple polypeptides depending on what is spliced; this is called ___________________. Exon shuffling during cross-over may also be useful in evolution. ...
Initiation at Class I Promoters
... 1. Promotes binding of TFIIB, which promotes recruitment of the other factors and RNAP. ...
... 1. Promotes binding of TFIIB, which promotes recruitment of the other factors and RNAP. ...
last year`s final exam
... 23) What is the function of snRNPs? 24) Mature, unfertilized eggs of many species have mRNAs for several genes but the proteins haven’t been made yet. What is preventing their synthesis? 25) What is the first amino acid to be added during synthesis of almost all eukaryotic proteins? 26) Describe one ...
... 23) What is the function of snRNPs? 24) Mature, unfertilized eggs of many species have mRNAs for several genes but the proteins haven’t been made yet. What is preventing their synthesis? 25) What is the first amino acid to be added during synthesis of almost all eukaryotic proteins? 26) Describe one ...
Analytical and Chromatography - Sigma
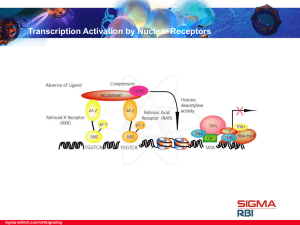
... • Retinoid X receptor (RXR) and Retinoic Acid receptor (RAR) are nuclear receptors that bind either all trans-retinoic (tRA) or 9-cis-retinoic acid (9cis-RA). In the absence of ligand, corepressors, such as Nuclear Receptor Corepressor (NCoR), Silencing Mediator of Retinoid and Thyroid Hormone Recep ...
... • Retinoid X receptor (RXR) and Retinoic Acid receptor (RAR) are nuclear receptors that bind either all trans-retinoic (tRA) or 9-cis-retinoic acid (9cis-RA). In the absence of ligand, corepressors, such as Nuclear Receptor Corepressor (NCoR), Silencing Mediator of Retinoid and Thyroid Hormone Recep ...
Poster Presentation
... When CaCl2 was added we found that only a few of the transcription factors responded by bursting. The transcription factors that responded to calcium are Crz1 and Msn2. When sorbital is added to the solution it causes osmotic shock to the yeast cells. The transcription factors that were found to exh ...
... When CaCl2 was added we found that only a few of the transcription factors responded by bursting. The transcription factors that responded to calcium are Crz1 and Msn2. When sorbital is added to the solution it causes osmotic shock to the yeast cells. The transcription factors that were found to exh ...
Examination in Bi3016 Molecular Cell Biology
... Transcription factors bind specific recognition sites in DNA and regulate gene expression. a. How does a transcription factor interact and recognize a binding site in a DNA strand? b. Explain how transcription factors specificity and affinity to DNA can be increased. How can this also increase the n ...
... Transcription factors bind specific recognition sites in DNA and regulate gene expression. a. How does a transcription factor interact and recognize a binding site in a DNA strand? b. Explain how transcription factors specificity and affinity to DNA can be increased. How can this also increase the n ...
Transcription factor
In molecular biology and genetics, a transcription factor (sometimes called a sequence-specific DNA-binding factor) is a protein that binds to specific DNA sequences, thereby controlling the rate of transcription of genetic information from DNA to messenger RNA. Transcription factors perform this function alone or with other proteins in a complex, by promoting (as an activator), or blocking (as a repressor) the recruitment of RNA polymerase (the enzyme that performs the transcription of genetic information from DNA to RNA) to specific genes.A defining feature of transcription factors is that they contain one or more DNA-binding domains (DBDs), which attach to specific sequences of DNA adjacent to the genes that they regulate. Additional proteins such as coactivators, chromatin remodelers, histone acetylases, deacetylases, kinases, and methylases, while also playing crucial roles in gene regulation, lack DNA-binding domains, and, therefore, are not classified as transcription factors.