Final Presentation
... What is Gene Regulation? • Mechanism by which cells increase or decrease specific gene products. • All organisms react to changes in the external environment using gene regulation. • Due to its relatively small genome and the readily available genome datasets, Saccharomyces cerevisiae is a great mo ...
... What is Gene Regulation? • Mechanism by which cells increase or decrease specific gene products. • All organisms react to changes in the external environment using gene regulation. • Due to its relatively small genome and the readily available genome datasets, Saccharomyces cerevisiae is a great mo ...
Regulation of Gene Expression
... Consider Prokaryotes must be able to adjust to their changing environments o Sometimes the environment can change almost instantly Eukaryotes have to respond as well, although typically not as drastically With multicellular organisms, different types of cells express different sets of genes ...
... Consider Prokaryotes must be able to adjust to their changing environments o Sometimes the environment can change almost instantly Eukaryotes have to respond as well, although typically not as drastically With multicellular organisms, different types of cells express different sets of genes ...
13.4 Gene Expression
... it possible for researchers to switch genes on and off at will, simply by inserting doublestranded RNA into cells. It may provide new ways to treat and perhaps even cure diseases. ...
... it possible for researchers to switch genes on and off at will, simply by inserting doublestranded RNA into cells. It may provide new ways to treat and perhaps even cure diseases. ...
TAF15 Antibody
... evolutionarily conserved proteins known as TBP-associated factors or TAFs. TAFs may participate in basal transcription, serve as coactivators, function in promoter recognition or modify general transcription factors (GTFs) to facilitate complex assembly and transcription initiation. Its gene encodes ...
... evolutionarily conserved proteins known as TBP-associated factors or TAFs. TAFs may participate in basal transcription, serve as coactivators, function in promoter recognition or modify general transcription factors (GTFs) to facilitate complex assembly and transcription initiation. Its gene encodes ...
Worksheet 6 - Iowa State University
... 4. How does sigma recognize the promoter? Can sigma always bind to the promoter? ...
... 4. How does sigma recognize the promoter? Can sigma always bind to the promoter? ...
Identification of Tissue Specific Transcription Factors Using
... probes covering 810 genes encoding transcription factors were taken for the analysis. For every probe we made a plot showing its expression profile for 78 cell/tissue types. We classified all the profiles into three major groups. The first and largest group contains genes that show ubiquitous expres ...
... probes covering 810 genes encoding transcription factors were taken for the analysis. For every probe we made a plot showing its expression profile for 78 cell/tissue types. We classified all the profiles into three major groups. The first and largest group contains genes that show ubiquitous expres ...
outline File - selu moodle
... Genetic code is degenerate Stop codon Start codon Wobble effect at third position Near universal 15.3 Prokaryotic Transcription Begins at a promoter transcribes the transcription unit ends at the terminator Promoter – sequence within DNA Elongation uses RNA polymerase to add ribonucleotides that ...
... Genetic code is degenerate Stop codon Start codon Wobble effect at third position Near universal 15.3 Prokaryotic Transcription Begins at a promoter transcribes the transcription unit ends at the terminator Promoter – sequence within DNA Elongation uses RNA polymerase to add ribonucleotides that ...
Gene Expression
... • An activator and repressor have roles • Conditions tightly controlled – Lactose must be high, but no other sugar present – [Lactose] and [glucose] ...
... • An activator and repressor have roles • Conditions tightly controlled – Lactose must be high, but no other sugar present – [Lactose] and [glucose] ...
problem set
... I & III is inhibited at a 1 g/ml concentration. Therefore, one can determine if a particular gene is transcribed by RNA Pol II by determining if 1 g/ml -amanitin inhibits transcription of the gene. ...
... I & III is inhibited at a 1 g/ml concentration. Therefore, one can determine if a particular gene is transcribed by RNA Pol II by determining if 1 g/ml -amanitin inhibits transcription of the gene. ...
Structure of the DNA-binding motifs of activators
... • Proline-rich domains: CTF • Structure and function – not clearly related ...
... • Proline-rich domains: CTF • Structure and function – not clearly related ...
LKCMedicine Lecture Series by Asst Prof Chng Toh Hean
... transcription of new genes and synthesis of new proteins. The transport of synaptically-localised transcriptional regulators during neuronal activity provides a method of coupling synaptic activation with changes in the transcription. Asst Prof Chng’s lab is interested in studying the molecular basi ...
... transcription of new genes and synthesis of new proteins. The transport of synaptically-localised transcriptional regulators during neuronal activity provides a method of coupling synaptic activation with changes in the transcription. Asst Prof Chng’s lab is interested in studying the molecular basi ...
Lecture 4 – Gene Expression Control and Regulation
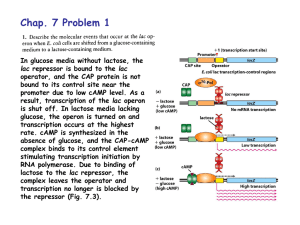
... and a single promoter (the lac operon) • When lactose is not present, repressors bind to the operators and inactivate the promoter; transcription does not proceed • When lactose is present, allolactose binds to the repressors; repressors don’t bind to operators to inactivate the promoter; transcript ...
... and a single promoter (the lac operon) • When lactose is not present, repressors bind to the operators and inactivate the promoter; transcription does not proceed • When lactose is present, allolactose binds to the repressors; repressors don’t bind to operators to inactivate the promoter; transcript ...
401Lecture5sp2013post
... Each probe specific for sequences separated by known distances in linear Fig. 6-35 Lodish et al. 2013 DNA What result would you expect if DNA exists in loops? Would you expect loops to be present at all stages of cell cycle? ...
... Each probe specific for sequences separated by known distances in linear Fig. 6-35 Lodish et al. 2013 DNA What result would you expect if DNA exists in loops? Would you expect loops to be present at all stages of cell cycle? ...
Expression of the transcription factor, TFII-I, in post
... Structurally, the TFII-I protein comprises several domains that define its biological function: an N-terminal leucine zipper domain is followed by six I-repeats (R1-R6), each containing a helix-loop-helix (HLH) domain. Leucine zipper is involved in dimerization, whereas sequence between the first an ...
... Structurally, the TFII-I protein comprises several domains that define its biological function: an N-terminal leucine zipper domain is followed by six I-repeats (R1-R6), each containing a helix-loop-helix (HLH) domain. Leucine zipper is involved in dimerization, whereas sequence between the first an ...
3. Activator, gene-specific transcription facotr
... • cis-acting element ; regulatory DNA sequence /enhancer • gene-specific transcription • extra boost in transcription • regulate chromatin structure & Gene activity ...
... • cis-acting element ; regulatory DNA sequence /enhancer • gene-specific transcription • extra boost in transcription • regulate chromatin structure & Gene activity ...
04/01
... Enhanceosome: protein complex of trans-acting factors bound to appropriate DNA sequences. Proteins interact synergistically to elevate transcription rate. In b-interferon gene transcription, TFs recruit a coactivator (CBP) which is needed for transcription to occur normally. Formation of the enhance ...
... Enhanceosome: protein complex of trans-acting factors bound to appropriate DNA sequences. Proteins interact synergistically to elevate transcription rate. In b-interferon gene transcription, TFs recruit a coactivator (CBP) which is needed for transcription to occur normally. Formation of the enhance ...
8.4 Transcription - School District of La Crosse
... – The large subunit has three binding sites for tRNA. – The small subunit binds to mRNA. ...
... – The large subunit has three binding sites for tRNA. – The small subunit binds to mRNA. ...
Differential Gene Expression
... 1. Most gene transcription requires enhancers. 2. Enhancers are the major determinants of differential transcription in cell types and through developmental stages. 3. There can be multiple signals (e.g. multiple enhancer sites) for a given gene, and each enhancer can be bound by more than one trans ...
... 1. Most gene transcription requires enhancers. 2. Enhancers are the major determinants of differential transcription in cell types and through developmental stages. 3. There can be multiple signals (e.g. multiple enhancer sites) for a given gene, and each enhancer can be bound by more than one trans ...
SEMINAR CANCELED- Rescheduled to January 28, 2016
... Rim101, and genes characteristic of invasive hyphal cells. The late phase includes responses related to phagocytosis by macrophages. Transcription factor gene expression also reflects early and late phases. Transcription factor genes that are required for virulence or proliferation in vivo are enric ...
... Rim101, and genes characteristic of invasive hyphal cells. The late phase includes responses related to phagocytosis by macrophages. Transcription factor gene expression also reflects early and late phases. Transcription factor genes that are required for virulence or proliferation in vivo are enric ...
20141203103493
... Acetylation of histone tails promotes loose chromatin structure that permits transcription ...
... Acetylation of histone tails promotes loose chromatin structure that permits transcription ...
Chapt16_lecture
... Control of Gene Expression • Initiation of Transcription is controlled by controlling gene expression. • Regulatory proteins bind to DNA to either block or stimulate transcription, depending on how they interact with RNA polymerase • Prokaryotic organisms are able to respond to changes in their env ...
... Control of Gene Expression • Initiation of Transcription is controlled by controlling gene expression. • Regulatory proteins bind to DNA to either block or stimulate transcription, depending on how they interact with RNA polymerase • Prokaryotic organisms are able to respond to changes in their env ...
Transcription factor
In molecular biology and genetics, a transcription factor (sometimes called a sequence-specific DNA-binding factor) is a protein that binds to specific DNA sequences, thereby controlling the rate of transcription of genetic information from DNA to messenger RNA. Transcription factors perform this function alone or with other proteins in a complex, by promoting (as an activator), or blocking (as a repressor) the recruitment of RNA polymerase (the enzyme that performs the transcription of genetic information from DNA to RNA) to specific genes.A defining feature of transcription factors is that they contain one or more DNA-binding domains (DBDs), which attach to specific sequences of DNA adjacent to the genes that they regulate. Additional proteins such as coactivators, chromatin remodelers, histone acetylases, deacetylases, kinases, and methylases, while also playing crucial roles in gene regulation, lack DNA-binding domains, and, therefore, are not classified as transcription factors.