* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Non-random Allelic Variation
Site-specific recombinase technology wikipedia , lookup
History of genetic engineering wikipedia , lookup
Adaptive evolution in the human genome wikipedia , lookup
Designer baby wikipedia , lookup
Dual inheritance theory wikipedia , lookup
Koinophilia wikipedia , lookup
Human genetic variation wikipedia , lookup
Quantitative trait locus wikipedia , lookup
Dominance (genetics) wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Gene expression programming wikipedia , lookup
The Selfish Gene wikipedia , lookup
Genetic drift wikipedia , lookup
Hardy–Weinberg principle wikipedia , lookup
Polymorphism (biology) wikipedia , lookup
Natural selection wikipedia , lookup
Population genetics wikipedia , lookup
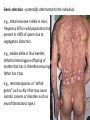
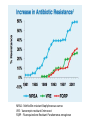
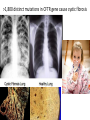
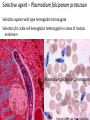
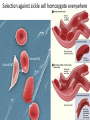
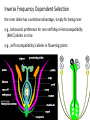
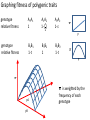
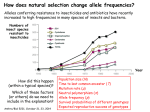
Non-random Allelic Variation AKA Natural Selection Adaptation – a derived character that evolved in response to a specific selective agent (although traits evolve from pre-existing ones and selective agents change in time) or – a feature that is maintained because of natural selection for its function preadaptation – a trait that fortuitously serves a new function adaptation is non-random or directional evolution adaptation results from selection selection can only act on existing variation most adaptations are modifications of pre-existing ones therefore, its results are most often ‘retro-fitted’ and some are suboptimally designed and can only be understood in the context of how they came to be adaptation results from selection Selection defined Wright: “any process in a population that alters gene frequency in a directed fashion without a change in genetic material” Futuyma: “any consistent difference in fitness (i.e., survival and reproduction) among phenotypically different biological entities [be they genes, species, or groups]” “directed” and “consistent” distinguish selection from drift, from randomness Selection is mathematically related to fitness – a measure of reproductive output with no other connotation Most people equate evolution with adaptation In fact, much of evolution is not adaptive Non-adaptive traits include those due to consequences of physics or chemistry, e.g., blood is red drift linkage or ‘genetic hitchhiking’ pleiotropy phylogenetic history Natural Selection Evolution evolution is a two step process 1) origin of genetic variation 2) change in the frequency of alleles and genotypes evolution can occur in the absence of natural selection natural selection can occur without evolution resulting natural selection acts on phenotypes, but evolution describes a change in genotype thus, the degree to which selection results in evolution depends on the heritability (i.e., % genetic variance) of a trait Selection is mathematically related to fitness Selection acts on the phenotype of individuals but, to the extent that selection can be an evolutionary force changing genotype frequency, it is the average reproductive success of heritable phenotypes that counts after all, there may be non-selective deaths of individuals with genotypes of otherwise high average fitness Reductionist view of selection Richard Dawkins The Selfish Gene 1989 “a coach selects a team of oarsmen for a crew race by repeatedly shifting oarsmen among several boats and racing them, after several trials the winning boat will have all the same oarsmen. A crew member finally chosen will have been grouped with both good and inferior ones at different times, but on average his performance has contributed more to the trials than one who was not chosen. Natural selection within populations can be understood as competition among alleles, the winner being the one that confers some characteristic on an organism that provides the allele with the highest rate of survival and reproduction, averaged over all gene combinations in which the allele occurs” Levels of selection Genic selection Individual selection Kin selection Group selection Species selection It is self-evident that selection acts at different levels because these may be in mutual conflict Genic selection – potentially detrimental to the individual e.g., lethal recessive t-allele in mice, frequency 40% in wild populations but present in >90% of sperm due to segregation distortion e.g., medea allele in flour beetles, lethal to heterozygous offspring of mother that has it, therefore ensuring father has it too e.g., retrotransposons or “selfish genes” such as Alu I that may cause somatic cancers or disorders such as neurofribromatosis type 1 In the case of genic selection, we certainly don’t imagine that the tallele or Alu 1 retrotransposon is making a conscious decision or planning its own increase it is simply that through some mechanism of mutation a variant appeared at some time in the past that conferred an ability for the gene to reproduce itself or to reproduce itself more rapidly, and so it did thus, what we describe as selection is simply the consequence of a reproductive advantage stated differently, selection has no forethought if it had, then the t-allele or Alu 1 retrotransposon might realize that by increasing its own reproductive success in the short term it negatively impacts the organism it depends on to survive in the long term Individual selection – potentially detrimental to the species e.g., “cheaters” or individuals who deceive others for personal gain e.g., infanticide in lions, rats, various birds Does a male lion that just claimed a pride decide to kill existing cubs specifically because he consciously wants to optimize his reproductive success relative to other genotypes? probably not Again, without forethought, “selection favors” a particular phenotype (in this case a behavior) because it has the result of improving the reproductive output of individuals that trait relative to those that do not Kin selection i.e., assisting the reproductive success of relatives to improve one’s own inclusive fitness (the notion that an individual’s fitness is increased by the fitness of all who share his or her genes) e.g., delayed dispersal of young and cooperative breeding in a variety of birds, such as Harris’ hawks, acorn woodpeckers, scrub jays, , choughs, white-fronted bee-eaters But appearances may be deceiving Genotyping has revealed what goes on behind closed doors and within nesting cavities nest helpers are sometimes cuckolders moreover, some offspring choose to become nest helpers only after their own attempt at nesting was sabotaged - by their own parents! Altruism and Group Selection vs Kin Selection Altruism – an act for the benefit of another with potential cost and no expectation of personal reward (including inclusive fitness) Altruism is unlikely to evolve due to natural selection because there is no heritable fitness advantage to the provider; but it can result as a ‘unintended’ or ‘misdirected’ consequence of another selected trait Altruism and Group Selection vs Kin Selection Group selection – like altruism, group selection acts for the benefit of a group of individuals that are not necessarily related Group selection is unlikely to evolve due to natural selection or drift because the effective population size of the group is larger than the lineage of the selfish individual (“cheater”) who derives an immediate fitness benefit and whose lineage can evolve more rapidly to exploit altruistic behaviors of the group Species selection e.g., extinction of species with little genetic variation e.g., evolution of sex ratios and population densities Selection is mathematically related to fitness Fitness of a genotype – average of fitnesses of all individuals of the genotype, or “the average per capita lifetime contribution of individuals of that genotype to the population after one or more generations” i i 𝑅 – absolute fitness of genotype i.e., per capita growth rate of genotype i i 𝑅 = (fraction of offspring of genotypei surviving to reproduce) X (average fecundity) (WARNING: there are alternative definitions of absolute fitness – avoid confusion, beware the internet) 𝑅 - average per capita growth rate of population (the equivalent of the intrinsic growth rate “r” of a given genotype but for clonally reproducing species with non-overlapping generations) i i 𝑊 – “relative fitness” i.e., value of 𝑅 relative to the absolute fitness of the maximally fit genotype 𝑅MAX i 𝑊 i 𝑅 = < 1.0 𝑅MAX 𝑅MAX 𝑊MAX = = 1.0 𝑅MAX The rate of genetic change under selection depends on the relative, not absolute, fitnesses of the genotypes 𝑅i number of individuals 𝑅MAX t=1 t=2 generations Different relative fitnesses yield different relative growth rates 𝑤 - “average fitness” i.e., the average fitness of individuals in the population relative to the maximally fit genotype 𝑤 = (𝑊 11)(frequency11) + (𝑊 12)(frequency12) + (𝑊 22)(frequency22) example Given: Genotype frequencies A1A1 = 20% A1A2 = 30% A2A2 = 50% Absolute fitnesses 𝑅11 = 6 𝑅12 = 8 𝑅22 = 10 Relative fitnesses 6 𝑊 11 = 10 = 0.6 8 𝑊 12 = 10 = 0.8 𝑊 22 = 10 = 1.0 = 𝑊 MAX 10 Average fitness 𝑤 = (𝑊 11)(frequency11) + (𝑊 12)(frequency12) + (𝑊 22)(frequency22) 𝑤 = (0.6)(0.2)+(0.8)(0.3)+(1)(0.5) = (0.12)+(0.24)+(0.5) = 0.86 example Given: Genotype frequencies A1A1 = 20% A1A2 = 30% A2A2 = 50% Absolute fitnesses 𝑅11 = 6 𝑅12 = 8 𝑅22 = 10 Relative fitnesses 6 𝑊 11 = 10 = 0.6 8 𝑊 12 = 10 = 0.8 𝑊 22 = 10 = 1.0 = 𝑊 MAX 10 Average fitness 𝑤 = (𝑊 11)(frequency11) + (𝑊 12)(frequency12) + (𝑊 22)(frequency22) 𝑤 = (0.6)(0.2)+(0.8)(0.3)+(1)(0.5) = (0.12)+(0.24)+(0.5) = 0.86 example Given: Genotype frequencies A1A1 = 20% A1A2 = 30% A2A2 = 50% Absolute fitnesses 𝑅11 = 6 𝑅12 = 8 𝑅22 = 10 Relative fitnesses 6 𝑊 11 = 10 = 0.6 8 𝑊 12 = 10 = 0.8 𝑊 22 = 10 = 1.0 = 𝑊 MAX 10 Average fitness 𝑤 = (𝑊 11)(frequency11) + (𝑊 12)(frequency12) + (𝑊 22)(frequency22) 𝑤 = (0.6)(0.2)+(0.8)(0.3)+(1)(0.5) = (0.12)+(0.24)+(0.5) = 0.86 example Given: Genotype frequencies A1A1 = 20% A1A2 = 30% A2A2 = 50% Absolute fitnesses 𝑅11 = 6 𝑅12 = 8 𝑅22 = 10 Relative fitnesses 6 𝑊 11 = 10 = 0.6 8 𝑊 12 = 10 = 0.8 𝑊 22 = 10 = 1.0 = 𝑊 MAX 10 Average fitness 𝑤 = (𝑊 11)(frequency11) + (𝑊 12)(frequency12) + (𝑊 22)(frequency22) 𝑤 = (0.6)(0.2)+(0.8)(0.3)+(1)(0.5) = (0.12)+(0.24)+(0.5) = 0.86 example Given: Genotype frequencies A1A1 = 20% A1A2 = 30% A2A2 = 50% Absolute fitnesses 𝑅11 = 6 𝑅12 = 8 𝑅22 = 10 Relative fitnesses 6 𝑊 11 = 10 = 0.6 8 𝑊 12 = 10 = 0.8 𝑊 22 = 10 = 1.0 = 𝑊 MAX 10 Average fitness 𝑤 = (𝑊 11)(frequency11) + (𝑊 12)(frequency12) + (𝑊 22)(frequency22) 𝑤 = (0.6)(0.2)+(0.8)(0.3)+(1)(0.5) = (0.12)+(0.24)+(0.5) = 0.86 example Given: Genotype frequencies A1A1 = 20% A1A2 = 30% A2A2 = 50% Absolute fitnesses 𝑅11 = 6 𝑅12 = 8 𝑅22 = 10 Relative fitnesses 6 𝑊 11 = 10 = 0.6 8 𝑊 12 = 10 = 0.8 𝑊 22 = 10 = 1.0 = 𝑊 MAX 10 Average fitness 𝑤 = (𝑊 11)(frequency11) + (𝑊 12)(frequency12) + (𝑊 22)(frequency22) 𝑤 = (0.6)(0.2)+(0.8)(0.3)+(1)(0.5) = (0.12)+(0.24)+(0.5) = 0.86 example Given: Genotype frequencies A1A1 = 20% A1A2 = 30% A2A2 = 50% Absolute fitnesses 𝑅11 = 6 𝑅12 = 8 𝑅22 = 10 Relative fitnesses 6 𝑊 11 = 10 = 0.6 8 𝑊 12 = 10 = 0.8 𝑊 22 = 10 = 1.0 = 𝑊 MAX 10 Average fitness 𝑤 = (𝑊 11)(frequency11) + (𝑊 12)(frequency12) + (𝑊 22)(frequency22) 𝑤 = (0.6)(0.2)+(0.8)(0.3)+(1)(0.5) = (0.12)+(0.24)+(0.5) = 0.86 example Given: Genotype frequencies A1A1 = 20% A1A2 = 30% A2A2 = 50% Absolute fitnesses 𝑅11 = 6 𝑅12 = 8 𝑅22 = 10 Relative fitnesses 6 𝑊 11 = 10 = 0.6 8 𝑊 12 = 10 = 0.8 𝑊 22 = 10 = 1.0 = 𝑊 MAX 10 Average fitness 𝑤 = (𝑊 11)(frequency11) + (𝑊 12)(frequency12) + (𝑊 22)(frequency22) 𝑤 = (0.6)(0.2)+(0.8)(0.3)+(1)(0.5) = (0.12)+(0.24)+(0.5) = 0.86 Coefficient of Selection (𝑠i) The intensity of selection against a less fit genotype, or the selective advantage of a more fit genotype i i 𝑠 = 𝑊MAX − 𝑊 previous example, selection against A1A2 (or for A2A2 relative to A1A2) 𝑠22 = 1.0 − 0.8 = 0.2 With selection, allele frequency will change Example using haploid with alleles A and B, with selection against A ′ 𝑝 = 𝑞𝑅𝐴 𝑤 = 𝑞𝑅𝐴 𝑞𝑅𝐴 +𝑞𝑅𝐵 ∆𝑝 = 𝑝′ − 𝑝 −𝑠𝑝𝑞 ∆𝑝 = 1 − 𝑠𝑝 (note this equation applies only to this case) p is negative if s, p, and q are positive The absolute value of p is greatest when s is large and p = q p slows as p 0 For diploids with alleles A1 and A2 𝑝′ = = 1 2 freq𝐴1 𝐴1 𝑤11 + freq𝐴1 𝐴2 𝑤12 𝑤 𝑝2 𝑤11 + 1𝑝𝑞 𝑤12 𝑝2 𝑤11 + 2𝑝𝑞 𝑤12 + 𝑞 2 𝑤22 this equation will always apply to a di-allelic locus in diploids (memorize it) Modes of Selection directional or purifying 𝒔 𝑝 stabilizing or normalizing 𝒕 𝒔 diversifying or disruptive 𝒔 Directional Selection with intermediate heterozygote fitness directional 𝒔 𝑝 genotype genotype frequency relative fitness ∆𝑝 = 1 𝑠𝑝𝑞 2 1−𝑠𝑞 A1A1 p2 1 A1A2 2pq 𝑠 1-( ) 2 A2A2 q2 1-𝑠 (where 1 − 𝑠𝑞 = 𝑤 ; note this equation applies only to this case) Rate of change (∆) is maximal when 𝑝 = q = 0.5 (at any given s) Rate of change (∆) is minimal as q 0 Directional Selection with dominance directional 𝒔 𝑝 genotype relative fitness ∆𝑝 = 𝑠𝑝𝑞 2 1−𝑠𝑞 2 A1A1 1 A1A2 1 A2A2 1-𝑠 (where 1 − 𝑠𝑞 2 = 𝑤 ; note this equation applies only to this case) Even the slightest positive 𝑠 will result in fixation of the A1 allele Rate of change (∆) is maximal when 𝑝 = ⅓ and q = ⅔ ∆ decreases as q 0 because the denominator (𝑤 or average fitness) increases Directional Selection with dominance directional 𝒔 𝑝 Recessive alleles are shielded from selection as q 0 because rare alleles exist almost exclusively in the heterozygous state Rare deleterious recessive alleles exist at low frequencies at many loci Rare advantageous recessive alleles are also shielded from positive selection, therefore initial increase in their frequency is expected to be slow and may be dependent on drift ∆𝑝 equations using 𝑠 differ with every mode of selection (so don’t memorize one and expect it to work for all problems) stabilizing or normalizing 𝒔 𝒕 genotype fitness A1A1 1-𝑠 A1A2 1 A2A2 1-𝑡 diversifying or disruptive 𝒔 genotype fitness A1A1 1 A1A2 1-𝑠 A2A2 1 MRSA - Methicillin-resistant Staphylococcus aureus VRE - Vancomycin-resistant Enterococci FQRP - Fluoroquinolone-Resistant Pseudomonas aeruginosa Nature 497, 24–26 (02 May 2013) doi:10.1038/497024a Persistence of Polymorphism If selection fixes alleles then why is there polymorphism? 1) Recurrent mutation 2) Gene flow 3) Genetic drift 4) Balancing selection Persistence of polymorphism by recurrent mutation mutation A1 A2 𝑞= 𝝁 𝒔 Where 𝑞 = equilibrium frequency of A2 (the mutant allele) 𝝁 = mutation rate 𝒔 = selection coefficient Don’t memorize the equation Remember that the ratio of the numerator to the denominator is what determines the equilibrium frequency of the mutant >1,800 distinct mutations in CFTR gene cause cystic fibrosis Persistence of polymorphism by gene flow A2 is advantageous elsewhere 𝑞∝ 𝑚 𝑞𝑚 𝑠 Where 𝑞 = equilibrium frequency of A2 𝑚 = rate of gene flow 𝑞𝑚 = frequency of allele among immigrants If rate of gene flow 𝑚 >> 𝑠, then populations won’t differentiate Don’t memorize the equation Remember that the ratio of the numerator to the denominator is what determines the equilibrium frequency of the mutant Persistence of polymorphism by gene flow Cline – continuous or monotonic change in allele frequency along a geographic transect; the hallmark of gene flow Clinal variation in aminopeptidase gene in mussels (Mytilus edulis) in Long Island Sound seawater Long Island Sound Hudson River brackish water Long Island Atlantic Ocean Clinal variation in SNP in human HERC2 intron for blue-brown eye color Balancing Selection (not mutually exclusive of normalizing and disruptive selection) Types 1) Heterozygote advantage 2) Antagonistic selection 3) Varying selection 3a) Temporal 3b) Spatial 4) Inverse Frequency dependent selection Heterozygote Advantage (hybrid vigor, heterosis) Selection “for” the heterozygote genotype fitness A1A1 1-𝑠 A1A2 1 A2A2 1-𝑡 Equilibrium frequency of allele = selection against the other allele divided by the sum of selection against both alleles Equilibrium frequency of A1 = 𝑝 Equilibrium frequency of A2 = 𝑞 𝑝= 𝑡 𝑠+𝑡 𝑞= 𝑠 𝑠+𝑡 Equilibrium frequencies depend on a balance of fitnesses of A1A1 and A2A2 Hybrid vigor most cases of heterozygote advantage are equivocal, and may simply represent inbreeding depression hybrid inbred inbred Antagonistic Selection selection “against” the homozygotes truly no different than heterozygote advantage because selection for one genotype is selection against another genotype fitness A1A1 1-𝑠 A1A2 1 A2A2 1-𝑡 Antagonistic Selection – Sickle Cell hemoglobin allele “Sickle cell disease is the most common inherited blood disorder in the United States, affecting 70,000 to 80,000 Americans. The disease is estimated to occur in 1 in 500 African Americans and 1 in 1,000 to 1,400 Hispanic Americans” (US Natl Lib Med) Selection against homozygous wild type hemoglobin genotype – no malaria resistance Selection against homozygous sickle cell hemoglobin genotype – vaso-occlusive crisis, hemolysis, anemia, and complications Selection for heterozygote in geographic areas of malaria endemism – malaria resistance Anopheles mosquito vector current distribution of malaria Selective agent – Plasmodium falciparum protozoan Selection against wild type hemoglobin homozygote Selection for sickle cell hemoglobin heterozygote in areas of malaria endemism Plasmodium falciparum gametocytes Selection against sickle cell homozygote everywhere Normal RBC Sickled RBC Varying Selection typically, individuals do not experience multiple environments Temporally varying environmental conditions different phenotypes are selected for or against at different times requires heterozygote disadvantage Spatially varying selection “multiple niche polymorphism” or “habitat selection” different phenotypes are selected for or against in geographically different environments model generally assumes limited gene flow between demes Varying Selection Apple Maggot Fly Rhagoletes pomonella Spatially segregated on hawthorn and apple fruits Temporally segregated by phenology of hawthorn and apple fruiting Inverse Frequency Dependent Selection the rarer allele has a selective advantage, simply for being rarer e.g., behavioral preference for non-self Major Histocompatibility (MHC) alleles in mice e.g., self-incompatibility S-alleles in flowering plants Alternative Equilibria (not balancing) Types Positive Frequency Dependent Selection Heterozygote Disadvantage (underdominance) Adaptive (fitness) landscapes Two types of frequency dependent selection Fitness of genotype depends on genotype frequencies in the population Inverse Frequency Dependent Selection – balancing, maintains polymorphism Positive Frequency Dependent Selection – not balancing, alternative equilibria Positive Inverse 𝑤22 𝑤𝑖 𝑤11 𝑤12 𝑤12 𝑤22 0 p 1.0 0 𝑤11 p 1.0 Positive Frequency Dependent Selection in Heliconius butterflies a “supergene” region for mimetic color pattern polymorphism without intermediates (Joron et al 2006 PLoS Biol 4: e303) Heterozygote Disadvantage or underdominance initially more frequent allele will become fixed 𝑝 = 𝑞 is unstable this differs from varying (and Disruptive) selection because there is gene flow 𝑤 unstable 𝑝 0 p 1.0 Heterozygote Disadvantage or underdominance initially more frequent allele will become fixed 𝑝 = 𝑞 is unstable this differs from varying (and Disruptive) selection because there is gene flow 𝑤 unstable 𝑝 0 p 1.0 Adaptive Fitness Landscapes until now, we have considered only two-dimensional graphs of single locus diallelic traits Fitness landscape - a multi-dimensional representation of the average fitness of populations with respect to polygenic traits (i.e., genotypes controlled by two or more loci) Graphing fitness of polygenic traits genotype relative fitness A1A1 1 genotype relative fitness B1B1 1-𝑡 A1A2 𝑠 1-( ) 2 B1B2 1 A2A2 1-𝑠 𝑤 𝑝 B2B2 1-𝑡 𝑤 𝑝 𝑤 𝑝A 𝑝B 𝑤 is weighted by the frequency of each genotype in this example, there is only one maximally fit genotype, marked 𝑤 𝑝A 𝑝B 𝑤 𝑝A 𝑝B but there can be multiple equally fit genotypes in a fitness landscape especially if fitness of a phenotype is influenced by many loci or if environmental conditions are varied Since selection is measured as relative fitness, by definition selection can only make populations “climb up” fitness landscapes selection has no forethought; it cannot drive a population down a fitness landscape to cross a “fitness valley” (of lesser average fitness) even if it is to reach a peak of higher fitness populations may be “stuck” in local optima even if they could hypothetically optimize genotypes for even better average fitness however, drift can alter allele frequencies in small populations independently of selection, either up or down fitness gradients because it is random Transilience the interaction of selection and drift that can drive a population down a fitness gradient into and through an adaptive or fitness “valley” across to a new fitness optimum transilience is thought to be particularly important in the evolution of founding populations, because drift has a large effect in small populations sizes, and founding populations may find themselves in novel environments for which their existing genotype frequencies are less than optimal If traits can be non-adaptive, then how are adaptations recognized? Complexity Suitability for function Selection experiments Comparative method Methods of studying selection 1) Correlations among populations H0 – variation is random HA – variation is not random e.g., latitudinal clinal variation in ADH locus in Drosophila on both sides of the equator Methods of studying selection 2) Deviations from expected genotype frequencies 2a) H0 – Hardy-Weinberg Equilibrium Caveat: there are other factors that can cause populations to not be in H-W equilibrium 2b) linkage disequilibrium between traits that produce a suite of functionally related characters despite their ability to be in linkage equilibrium Methods of studying selection 3) temporal patterns of stability or instability of polymorphisms e.g. H0 – drift will cause fixation and monomorphy of alleles but polymorphism is observed e.g., HA – ∆𝑝 too fast to be accounted for by drift or neutrality e.g., industrial melanism in peppered moth (Biston betularia) Methods of studying selection 4) Response to environmental perturbations e.g., change in bill dimensions of Medium ground (Geospiza fortis) finch on Daphne Island, Galapagos after 1982 El Niño Methods of studying selection 5) Genetic demography - relationship of fitness or survival to phenotypic variability e.g., experimental manipulation of tail length (attractive to females) in Long-tailed Widowbird (Euplectes progne) video Methods of studying selection 6) Functional studies e.g., temperature affect on rate of enzyme activity, such as Taq DNA polymerase in thermophilic bacteria (e.g., Thermophilus aquaticus) Methods of studying selection 6) Functional studies e.g., temperature dependence of enzymatic rate activity in Antarctic Icefishes (Notothenioidei) Methods of studying selection 6) Quantitative Trait Locus (QTL) Analysis Polygenic traits with continuous phenotypic variability (vs discrete phenotypes) and non-Mendelian patterns of inheritance Mass selection on large populations can produce rapid phenotypic effects outside the range of previously observed variation by bringing together existing genetic variation and multiple alleles into novel genetic combinations QTL studies e.g., William Castle’s experiments on pelage color in hooded rats 20 generations QTL studies aggressive vs domestic behavior in red fox (silver race) 40 generations Methods of studying selection 6a) QTL mapping linkage of trait in crossing experiments best studied in plants and animals that can be rapidly and selectively bred 6b) Genome Wide Association Studies (GWAS) statistical correlation of SNPS and structural variants to one another and to phenotype best studied in relation to disease in humans due to availability of many completely sequenced genomes, medical records, and interest Methods of studying selection 7) Molecular studies e.g., dN/dS ratio (nonsynonymous : synonymous substitutions) e.g., substitution rates per codon position e.g., codon bias