* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Keshara Senanayake Ms.Reep Chapter 19
Therapeutic gene modulation wikipedia , lookup
DNA vaccination wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Non-coding DNA wikipedia , lookup
Genetic engineering wikipedia , lookup
Minimal genome wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Microevolution wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Genome evolution wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Helitron (biology) wikipedia , lookup
Genomic library wikipedia , lookup
Primary transcript wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
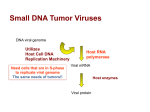
Keshara Senanayake Ms.Reep Chapter 19 - Viruses Chapter 27 - Bacteria and Archea Campbell Biology 9th edition CHAPTER 19 - VIRUSES by Keshara Senanayake oooo baby im virus free ;) A virus is an infectious particle consisting of genes packaged in a protein coat >viruses can’t reproduce/carry out metabolism activities out of a host cell study of viruses had led to the development of techniques that enable scientist to manipulate genes and transfer them from one organism to another (viruses are used as agents of gene transfer in gene therapy) Viruses consist of a nucleic acid surrounded by a protein cost Discovery of Virus Tobacco mosaic disease on tobacco plants (Adolf Mayer (German) found you could transmit the disease by rubbing sap from diseased leaf to healthy leaves deduced caused by “unusually small bacteria” Ivanowsky made a filter for bacteria but disease still transmitted Beijerinck saw pathogen replicated within the host it infected; showed that the mysterious agent could not be cultivated on Petri dish voice concept of smaller/simpler than bacterium (concept of virus) proved by Wendell Stanley by crystallizing the infectious particle electron microscope can see viruses) Structure of virus smallest are 20nm (smaller than ribosome) even largest are barely visible under light microscope Stanley showed that viruses could be crystallized close examination shows it consists of nucleic acid enclosed in a protein coast surrounded by a membranous envelope Viral Genomes their genomes may have double stranded DNA, single-stranded DNA, double-stranded RNA, or single-stranded RNA (depending on type of virus) virus is DNA virus or RNA virus based on kind of nucleic acid genome is usually organized as a single linear or circular molecule of nucleic acid >some genomes consist of multiple molecules of nuclei acid (smallest viruses have 4 genes in their genome, large has several hundred to a thousand, while bacteria have from 200 to a few thousand) Capsid is the protein shell enclosing the viral genome >capsid may be rod-shaped, polyhedral or more complex built from large # of Capsomeres (protein subunits) # of different kinds of proteins in a capsid is small Tobacco mosaic virus has a rigid rod-shaped capsid made from over a thousand molecules of a single type of protein arranged in a helix (rod-shaped viruses are commonly called helical viruses) Adenoviruses (infects respiratory tracts) have 252 identical protein molecules arranged in a polyhedral capsid with 20 triangular facets (an icosahedron) this/similar viruses are icosahedral viruses >have an icosahedra capsid with a glycoprotein spike at each vertex certain viruses have accessory structure to infect their host, like the membranous envelope that surrounds the capsids of influenza viruses viral envelopes contain host cell phospholipids/membrane proteins (and proteins and glycoproteins of viral origins [note: glycoproteins are proteins with carbs covalently attached]) >in an influenza virus the outer envelope is studded with glycoprotein spikes genome consists of 8 different RNA molecules each wrapped in a helical capsid complex capsids are found in viruses that infect bacteria (bacteriophages “simple phages”) first studded include 7 that infected e coli (named Type 1 (T1) Type 2 (T2) …. Type 7 (T7)), the even phages (T2,T4, and T6) are very similar in structure capsid have elongated icosahedral heads enclosing DNA attached to a protein tail w/ fibers by which phages attach to bacterium Viruses lack metabolic enzymes/equipment to make proteins are obligate intracellular parasites (replicate only in host cell) viruses can infect only a limited # of host species (host range) based on that viruses identify host cells by “lock and key” fit between viral surface protein and specific receptor molecules on the outside of the cell viral infections of multicultural eukaryotes is usually limited to particular tissues General Features of Viral Replicative cycles: virus infection begins w/ virus binding to host cell and viral genome going inside mechanism of genome entry depends on type of virus/host cell (T-even use tail apparatus to infect DNA into bacterium others are taken by via endocytosis for envelope viruses by fusion of viral envelope w/ plasma membrane) once the viral genome is inside, the protein it encodes commands the host, reprogramming cell to copy viral nucleic acid/manufacture viral proteins (host make nucleotides for viral nucleic acid, and enzymes/ribosomes/tRNAs/amino acids/ATP/ect) DNA viruses use DNA polymerase of host cell to synthesize new genomes along the template provided by viral RNA RNA viruses to replicated genomes use virally encoded RNA polymerases that use RNA as a template) after viral nucleic acid molecules/Capsomeres are produce they spontaneously self-assemble into new viruses simplest type of viral Replicative cycle ends w/ exit of hundred/thousands of viruses from infected host cell (damages/destroys host cell) (reason for body’s responses/symptoms) viral progeny that exits have potential to infect additional cells many variations on simplified viral Replicative cycle Simple viral Replicative cycle: 1) Virus enters cell and is uncoated releasing viral DNA 2) Host enzymes replicate the viral DNA 3) Meanwhile, host enzymes transcribe the viral genome into viral mRNA which host ribosome use to make more capsid proteins 4) Viral genomes and capsid proteins self-assemble into new viruses particles which exit the cell Replicative cycles of Phages pages are best understood of all viruses research has led to the discovery that some double-stranded DNA viruses can replicate by two alternative mechanisms: lytic and lysogenic cycles Lytic cycle phage Replicative cycle that culminates in death of host cell is lytic cycle refers to last stage on infection in which bacterium lyses (breaks open) and releases the phages that were produced within the cell (each of which can infect a healthy cell and a few successive lytic cycles can destroy an entire bacterial population) Lytic cycle of a phage T4, a virulent phage 1) Attachment. The T4 phage uses its tail fibers to bind to specific receptor sites on the outer surface of an E. coli cell 2) Entry of phage DNA and degradation of host DNA. The sheath of the tail contracts, injecting the phage DNA into the cell and leaving an empty capsid outside. The cell’s DNA is hydrolyzed 3)Synthesis of viral genomes and proteins. The phage DNA directs production of phage proteins and copies of the phage genome by host and viral genome by host and viral enzymes, using components within the cell 4) Assembly. Three separate sets of proteins self-assemble to form phage heads, tails, and tail fibers. The phage genome is packaged inside the capsid as the head forms 5) Release. The phage directors production of an enzyme that damages the bacterial cell wall, allowing fluid to enter. The cell swells and burst, releasing 100-200 virulent phage particles. phages that replicate only by lytic cycle is a virulent phage phage treatments have been used medically in some countries to control bacterial infection bacteria is not defenseless 1) via natural selection bacterial mutants w/ receptors that no longer recognize by a particular type of phage is favored 2) when a phage DNA successful enters a bacterium, the DNA is often identified as foreign and cut by restriction enzymes, which restricts the ability of the phage to infect bacterium >bacterial cell’s own DNA is methylated so it prevents attack by own restriction enzymes note that natural selection also favors phage mutants that can bring to altered receptors/resistant to particular restriction enzyme (so parasite-host relationship continues) and 3) reason is instead of lysing their host cells, many phages coexist with them in a state called lysogeny The Lysogenic cycle: The Lysogenic cycle allows replication of the phage genome without destroying the host Phages that use both modes of replication are temperate phages example is a lambda (λ) phage which is used in biological research resembled a T4 but it’s tail has one short tail fiber > infection of E. Coli by page λ starts when the phage binds to the surface of the cell and injects its linear DNA genome (giggity) in cell DNA molecule forms a circle next step depends on the Replicative mode (lytic cycle or lysogenic cycle) During a lytic cycle the viral genes immediately turn the host cell into a λ producing factory cell lyses and releases viral products During a lysogenic cycle the λ DNA molecule is incorporated into a specific site on E. Coli chromosome by viral proteins that break both circular DNA molecules and join them to each other when integrated into bacterial chromosome the viral DNA is known as a prophage >one prophage gene codes for a protein that prevents the transcription of most of the other prophage genes (so phage genome is “silent”) every time E. Coli cell divides it replicated the phage DNA along w/ its own and passes copies to daughter cells single infect cell quickly gives rise to large population of bacteria carrying virus in prophage form (propagate without killing host cell) lysogenic implies prophages are capable of active phages that lyses their host cells occurs when λ genome is induced to exit the bacterial chromosome and initiate a lytic cycle >an environmental signal (chemical/radiation) triggers the switchover from lysogenic to lytic mode other than the gene for transcription-preventing protein, a few other prophages may be expressed during lysogeny expression of these genes may alter host’s phenotype important medical significance (3 species of bacteria that cause human diseases diphtheria, botulism, and scarlet fever would not be so harmful to humans without prophage genes that cause host bacteria to make toxins) Replicative cycle of Animal Viruses many variations on the basic scheme of viral infection/replication are represented among animal viruses one key variable is the nature of the viral genome (composed of DNA/RNA? Double/single stranded?) nature of genome is basis for common classification of viruses single-stranded RNA viruses are further classified into (3) classes (IV - VI) based on how RNA genome functions in a host cell While few bacteriophages have an envelope/RNA genome, many animal viruses have both (nearly all animal viruses w/ RNA genome have an envelope as do some DNA genomes) Viral Envelopes animal virus equipped w/ an envelope (outer membrane) uses it to the host cell on outer surface of the envelope are viral glycoproteins that bind to specific receptor molecules on the surface of the host cell The Replicative cycle of an enveloped RNA virus (based on 19.7) 1) Glycoproteins on the viral envelope bind to specific receptor molecules (not shown) on the host cell, promoting viral entry into the cell 2) The capsid and viral genome enter the cell. Digestion of the capsid by cellular enzymes release the viral genome 3) The viral genome functions as a template for synthesis of complementary RNA strands by viral RNA polymerase 4) New copies of viral genome RNA are made using complementary RNA strands as templates 5) Complementary RNA strands also function as mRNA which is translated into both capsid proteins (in the cytosol) and glycoproteins for the viral envelope (in the ER and Golgi apparatus) 6) Vesicles transport envelope glycoproteins to the plasma membrane 7) A capsid assembled around each viral genome molecule 8) each new virus buds from the cell, its envelope studded w/ viral glycoproteins embedded in membrane derived from the host cell >Ribosome bound to the ER of the host cell make the protein part of envelope glycoproteins cellular enzymes in ER/Golgi then add sugars resulting viral glycoproteins embedded in host cell-derived membrane are transported to the cell surface. In an exocytose like process, new viral capsids are wrapped in membrane as they bud from cell SO, the viral envelope is derived from the host cell’s plasma membrane (although some molecules of membrane are specified by viral genes) enveloped virus are free to infect other cells this Replicative cycle does not kill host some viruses have envelopes not derived from plasma membrane (like herpes viruses temporarily cloaked in membrane derived from nuclear envelope of host; then shed this membrane in the cytoplasm and acquire a new envelope made from membrane of the Golgi apparatus) these viruses have doublestranded DNA genome and replicate within the host cells nucleus using viral + cellular enzymes to replicate/transcribe DNA (herpes viruses copied of DNA can remain behind as mini-chromosomes in nuclei of certain nerve cells latent until physical/emotional distress triggers a new round of active virus production) RNA as Viral Genetic Material although some phages/plant viruses are RNA viruses, but the broadest range of RNA genomes is found among the viruses that infect animals >among the (3) types of single-stranded RNA genomes found in animal viruses, the genome of class IV viruses can directly serve as mRNA can be translated into viral protein immediately after infection >19.7 from above is a class V, where the RNA genome serves as a template for mRNA synthesis RNA genome is transcribed into complementary RNA strands which acts as mRNA and as a template for the synthesis of additional copies of genomic RNA all viruses that require RNA RNA synthesis to make mRNA use a viral enzyme capable of carrying out this process (no enzymes like this in cells viral enzyme is packaged w/ genome inside the viral capsid) RNA virus w/ most complicated Replicative cycles are retroviruses (class VI) equipped w/ an enzyme called reverse transcriptase which transcribes an RNA template into DNA (providing RNA DNA flow [the opposite of the usual]) applies to HIV, the retrovirus that causes AIDS HIV/other retroviruses are enveloped viruses w/ two identical molecules of single stranded RNA and two molecules of reverse transcriptase The Replicative cycle of HIV: 1) The envelope glycoprotein enable the virus to bind to specific receptors on certain white blood cells 2) The virus fuses w/ the cell’s plasma membrane. The capsid proteins are removed, releasing the viral proteins and RNA 3) Reverse transcriptase catalyzes the synthesis of a DNA strand complementary to the viral RNA 4) Reverse transcriptase catalyzes the synthesis of a 2nd DNA strand molecule complementary to the first 5) The double stranded DNA is incorporated as a provirus into the cell’s DNA 6) Proviral genes are transcribed into RNA molecules, which serves as genomes for the next viral generation and as mRNAs for translation into viral proteins 7) The viral proteins include capsid proteins and reverse transcriptase (made in cytoplasm) and envelope glycoproteins (made in ER) 8) Vesicles transport the glycoproteins to the cell’s plasma membrane 9) Capsids are assembled around viral genomes and reverse transcriptase molecules 10) New viruses bud off from the host cell after HIV enter cell, its reverse transcriptase molecules are released into cytoplasm where they catalyze synthesis of viral DNA which then enters the cell’s nucleus and integrates into the DNA of a chromosomes. The integrated viral DNA is a provirus, never leaves host’s genome (remaining a permanent resident prophage in contrast leaves the host genome at the start of a lytic cycle) host’s RNA polymerase transcribes the proviral DNA into RNA molecules (which can function both as mRNA for the synthesis of viral proteins and as genomes for the new viruses that will be assembled and released from the cell) Evolution of Viruses note an isolated virus is unable to replicate its genes/regenerate its own supply of ATP (but it has genetic program written in the universal language of life) viruses are found in every form of life since they depend on cells for their own propagation viruses are not descendents of precellular forms of life but evolved after the first cells appeared >biologist favor the hypothesis that viruses originated form naked bits of cellular nucleic acids that moved from one cell to another (via injured cell surfaces) evolution of genes coding for capsid proteins helped infection of uninjured cells candidates for original sources of viral genomes include plasmids (small, circular DNA. Exist apart from cell’s genome, can replicate independently of genome, and are occasionally transferred between cells) and transposons (DNA segments that can move from one location to another within a cell’s genome) Plasmids/transposons/viruses are mobile genetic elements viral genome can have more in common w/ the genome of its host than w/ genomes of viruses that infect other host recent sequencing of viral genomes has shown that the genetic sequencing of many viral genomes are similar to distantly related viruses (some animal viruses share similar sequence w/ plant viruses can show natural selection for certain viral genes favored) Mimi virus is the largest virus discovered Viruses, viroids, and prions pathogens in animals/plants vaccine is a harmless variant or a derivation of a pathogen that stimulates the immune system to mount defenses against the harmful pathogen antibiotics are powerless against viruses antibiotics kill bacteria by inhibiting enzymes specific to bacteria but have not effect on eukaryotic/virally encoded enzymes most antiviral drugs resemble nucleosides and as a result interfere w/ viral nucleic acid synthesis viruses on the rise are emerging viruses general outbreak is an epidemic global epidemic is an pandemic >three process contribute to the emergence of viral diseases 1) mutation of existing viruses RNA viruses have an unusual high rate of mutation because errors in replicating their RNA genomes are not corrected by proofreading some mutations change existing viruses into new genetic varieties (strains) that cause disease in individuals who are immune to ancestral form of that virus 2) dissemination of a viral disease from a small isolated human population 3) spread of existing viruses from other animals (3) types of influenza (B and C [only humans] and A [infects a wide range of animals]) Viral Diseases in Plants over 2,000 types of viral diseases in plants most have same basic structure and mode of replication as animal viruses most have an RNA genome, many a helical capsid (like TMV) or icosahedra capsid Viral disease of plants spread via (2) major routes (1) Horizontal transmissions a plant infected from an external source of the virus invading virus must get past the plant’s outer protective layer of cells (epidermis) a plant becomes more susceptible to viral infections it has been damages by wind/injury/herbivores (especially insects, post a double threat because they can act as carriers of viruses) or farmers/gardeners who sure tools can spread it (2) Vertical transmission plant inherits a viral infection from a parent (occur in asexual propagation [through cutting] or in sexual reproduction via infected seeds) once virus enters a plant cell/begins replicating, viral genomes/associated proteins can spread throughout the plant via plasmodesmata Passage of viral macromolecules from cell to cell is facilitated by virally encoded proteins that cause enlargement of plasdeomesmata Viroids and Prions: The Simplest Infectious Agents viroids are circular RNA molecules that infect plants >do not encode proteins but can replicate in host plant cells (using host cell enzymes) seem to cause errors in the regulatory systems that control plant growth signs of viroids disease are abnormal development/stunted growth in viroids a single molecule can be infectious agent that spreads a disease but viroids are nucleus acids infectious proteins, prions, appear to cause a # of brain degenerative disease in animals (mad cow disease) prions are most likely transmitted in food two characteristics of prions that are alarming 1) prions act very slowly (incubation period of at least ten years before symptoms develop) lengthy incubation period prevents source of infection from being identified until long after the first cases appeared (allowing many more infections to occur) 2) prions are virtually indestructible not destroyed/deactivated by heating to normal cooking temperature >no known cure for prions disease prions is a misfolded form of a protein normally present in brain cells when prion gets into a cell contain the normal protein molecule the prion converts the normal protein molecule into a misfolded type several prions then aggregate into a complex that can convert other normal proteins into prions which join the chain. Prions aggregate interferes w/ normal cellular functions and cause disease symptoms Campbell 9th edition Chapter 27 - BACTERIA AND ARCHEA prokaryotic species are well adapted to more “normal” habitats than other species found ability to adapt to a broad range of habitats explain why prokaryotes are the most abundant organisms on earth Structural and functional adaptations contribute to prokaryotic success most prokaryotes are unicellular (some species remain attached to each other after cell divisions) typical diameters 0.5 - 5 micrometers (eukaryotic cells are 10-100 micrometers) prokaryotes have many shapes are well organized achieving all of an organism’s life functions within a single cell Most common shapes of prokaryotes (a) Spherical (like Cocci) (b) Rod-shaped ( like bacilli) (c) Spiral ( like Spirilla) Cell Surface Structures key feature of nearly all prokaryotic cells is cell wall maintains cell shape/protects cell/prevents it from bursting in an hypotonic environment (most lose water and shrunk away from wall [plasmolyze] water los inhibits cell reproduction) most bacterial cell walls contain peptidoglycan (polymer made up of modified sugars cross-linked by short polypeptides) encloses the entire bacterium/anchors other molecules that extend from surface (archeal cell walls have a variety of polysaccharides/proteins but LACK peptidoglycan) Gram stain has helped scientist identify many bacterial species samples are first stained w/ crystal violet dye/iodine then rinsed in alcohol and the stained w/ red dye (safranin) structure of bacterium’s cell wall determines the staining response >gram positive bacteria w/ simpler walls with a relatively large amount of peptidoglycan >gram negative less peptidoglycan and are more complex w/ an outer membrane that contains lipopolysaccharides (carb + lipid) (also more dangerous) lipid portion of lipopolysaccharides in walls of gram-negative bacteria are toxic outer membrane of a gram negative bacterium protects it from body’s defenses tends to be more resistant to antibiotics (because outer membrane impedes entry of drugs) >antibiotics inhibits peptidoglycan cross-linking (effective) theirs a sticky layer of polysaccharides/protein that surrounds the cell wall of prokaryotes >capsule if well defined >slime layer if less organized both enable prokaryotes to adhere to their substrate or to other individuals in a colony also protect against dehydration and shield pathogenic prokaryotes from attacks by immune system some prokaryotes stick to substrate/one another via hair like appendages called fimbriae usually shorter and more numerous than pilus, appendages that pull two cells together prior DNA transfer from one cell to another (often referred to as sex pilus) Motility ½ of all prokaryotes are capable of taxis, directed movement toward or away from a stimulus (chemo taxis towards (+) or away (-) from chemicals) Flagella help prokaryote move, may be scattered over the entire surface of cell/concentrated at one or both ends. prokaryotic flagella differ from eukaryotic flagella: 1/10 th in width and are not covered by an extension of the plasma membrane (also different in composition/mechanism of propulsion) >bacterial/archeal flagella are similar in size/rotation mechanism (composed of different proteins) show that bacteria/Achaea/eukaryotes arose independently >similar functions but not related by common descent, so they are analogous not homologous structures Evolutionary Origins of Bacterial Flagella bacterial flagella has (3) main parts (motor, hook, and filament) originated as simpler structures that were modified over time only ½ of flagellum’s protein components are necessary for its function (other is inessential or not encoded in the genomes of some species. Of the 21 proteins required by all species, 19 are modified versions of proteins that perform other tasks in bacteria) >some proteins found in motor are homologous to similar proteins in secretory system found in bacteria other proteins in motor are homologous to proteins that function in ion transport >proteins that comprise the rod, hook, and filament are all related to each other and are descended from an ancestral protein that formed a pilus-like tube >suggest bacterial flagellum evolved as other proteins were added to an ancestral secretory system example of exaptation (process in which existing structures take on new functions through descent w/ modification) Internal Organization and DNA cells of prokaryotes are simpler than those of eukaryotes in both their internal structure/physical arrangement of their DNA >prokaryotes lack complex compartmentalization found in eukaryotic cells (some prokaryotic cells do have specialized membranes that perform metabolic functions usually infoldings in plasma membrane) genome of prokaryote is structurally different from an eukaryotic genome (has less DNA) and in prokaryotes the genome consists of a circular chromosome w/ many fewer proteins than found in the linear chromosome of eukaryotes prokaryotes lack a membrane-bound nucleus (chromosome located in the nucleoid) in addition to its single chromosome a typical prokaryote cell may have smaller rings of independently replicating DNA molecules called plasmids most carrying a few genes DNA replication, transcription, and translation are similar processes in prokaryotes and eukaryotes >some differences (prokaryotic ribosome are smaller than eukaryotic ribosome/differ in their protein and RNA content allow certain antibiotics to bind to ribosome and block protein synthesis in prokaryotes but not eukaryotes) Reproduction and Adaptation prokaryotes can reproduce quickly in a favorable environment by binary fission, a single prokaryotic cell divides into 2 cells, which divide into 4, 8 ect >under optimal conditions most divide in 1 - 3 hours (some in 20 minutes) exponential but prokaryotic reproduction is limited eventually exhaust their nutrient supply/poisoned by metabolic wastes, face competition, or consumed >prokaryotes are small, reproduce by binary fission, and have short generation times prokaryotes can withstand harsh conditions due to particular biochemical adaptation; others because of particular structural adaptations Certain bacteria develop resistant cells called endospores when they lack essential nutrients >original cell produces a copy of its chromosomes and surrounds it w/ a tough structure forming an endosperm water is removed from endosperm and its metabolism halts original cell lyses, releasing endosperm durable (needs 121 Celsius under high pressure, in less hostile environments it says dormant but viable for centuries can rehydrate and resume metabolism when environment improves) because of short generation times prokaryotic populations can evolve substantially in short periods of time Rapid Reproduction, mutation, and genetic recombination promote genetic diversity in prokaryotes prokaryotes have considerable genetic variation (3) factors give rise to high levels of genetic diversity > (1) rapid reproduction (2) mutation (3) genetic recombination Rapid Reproduction and Mutation most genetic variation in sexual populations results from the way existing alleles are arranged in new combinations during meiosis and fertilization variation can result from prokaryotes rapid reproduction/mutation after repeated rounds of binary fission, most offspring are genetically identical but if errors occur during DNA replication (insertion, deletions, or base-pair substitutions) some offspring may be different new mutations, though rare, can increase genetic diversity quickly in species w/ short generation times and large populations can lead to rapid evolution: individuals that are genetically better equipped for their environment tend to survive/reproduce than unfit individuals Genetic Recombination additional diversity arises from genetic recombination combining of DNA from two sources in eukaryotes the sexual processes of meiosis + fertilization combined DNA from two individuals in a zygote (does not happen in prokaryotes) Instead (3) other mechanisms can bring together prokaryotic DNA from different individuals (cells) transformation, transduction, and conjugation when individuals are members of different species this movement from one organism to another is called horizontal gene transfer 1) In transformation genotype and possibly phenotype of a prokaryotic cell are altered by the uptake of foreign DNA from its surroundings (a nonpathogenic cell can take up a piece of DNA carrying the allele for pathogenticity and replaces its own allele with the foreign allele, an exchange of homologous DNA segments. The cell is now a recombinant: its chromosomes contain DNA derived from two different cells 2) In transduction, phages (from “bacteria phages” the viruses that infect bacteria) carry prokaryotic genes from one host cell to another transduction results from accidents that occur during the phage Replicative cycle >a virus that carries prokaryotic DNA may not be able to replicate because it lacks some/all of its genetic material but the virus can attach to another prokaryotic cell (recipient) and inject prokaryotic DNA acquired from the first cell (donor) if some of this DNA is then incorporated into the recipient cell’s chromosome by DNA recombination, a recombinant is formed Conjugation and Plasmids in conjugation two prokaryotic cells (usually same species) are temporarily joined in bacteria, the DNA transfer is always one-way, one cell donates DNA and the other receives it in E. Coli pilus of the donor cell attaches to the recipient pilus then retracts pulling the two cells together temporary “mating bridge” forms between cells through which the donor may transfer DN to the recipient the ability to form pilus and donate DNA during conjugation results from the presence of a particular piece of DNA called the F factor can exist either as plasmid or as a segment of DNA within the bacterial chromosome The F Factor as a plasmid F factor in its plasmid form the F plasmid Cells containing F plasmid (F+ Cell) function are DNA donors during conjugation. Cells without F factor (F-) function as DNA recipients during conjugation. F+ condition is transferred meaning that F+ cell converts F- cell to F+ if a copy of entire F plasmid is transferred The F Factor in the chromosome chromosomal genes can be transferred during conjugation when donor’s cell’s F factor is integrated into the chromosome >Hfr (high frequency recombination) cell is a cell w/ F factor built into its chromosome Hfr cell functions as a donor during conjugation w/ F- cell when its chromosomal DNA enters F- cell homologous regions of Hfr and F- chromosome align allowing segments of their DNA to be exchanged results in the production of a recombinant bacterium that has genes derived from (2) different cells (has genetic variation) R Plasmids and Antibiotic Resistance sometimes mutation in a chromosomal gene of a pathogen can confer resistance I.e: a mutation in one gene may make it less likely that the pathogen will transport a particular antibiotic into its cell another gene mutation may change the intracellular target protein for an antibiotic molecule Some bacteria have “resistance genes” code for enzymes that specifically destroy or hinder antibiotics carried by plasmids known as R (resistance) plasmids antibiotics will kill antibiotic-sensitive bacteria but not those w/ R plasmids that counter that antibiotic natural selection would cause the fraction of the bacterial population carrying genes for antibiotic resistance to increase many R plasmids like F plasmids have genes that encode pilus and enable DNA transfer from one bacterial cell to another by conjugation Diverse nutritional and metabolic adaptations have evolved in prokaryotes because of extensive genetic variation >prokaryotes an be categorized by how they get energy/carbon used to making organic molecules that make up cells have broader range of metabolic adaptation organisms that get energy via light are phototrophs, via chemicals are chemotroph >need only CO2 as a carbon source are Autotrophs heterotroph require at least one organic nutrient (4 classes: photoautotroph, chemoautotroph, photoheterotroph, and chemoheterotroph) Role of Oxygen in Metabolism obligate aerobes must use O2 for cellular respiration obligate anaerobes are poisoned by O2 (some live via fermentation) some extract chemicals by anaerobic respiration (use substances other than O2 to accept electrons) Facultative anaerobes use O2 if present but can also carry out fermentation or anaerobic respiration in an anaerobic environment Nitrogen metabolism nitrogen is essential for amino acids/nucleic acids prokaryotes can metabolize nitrogen in many forms Cynobacteria and some methanogens (Achaea) convert N2 to ammonia (NH3) “Nitrogen Fixation” >cells can incorporate “fixed’ nitrogen into amino acids/other organic molecules vitals for plants which can’t use atmospheric nitrogen but can use nitrogen compounds prokaryotes produce from ammonia Metabolic Cooperation cooperation between prokaryotic cells allow them to do things they can’t as individual cells cooperation takes place between specialized cells of a filament most cells in a filament carry out only photosynthesis, while a few specialized cells called heterocyst carry out only nitrogen fixation which is surrounded by a thickened cell wall that restricts entry of O2 produced by neighboring photosynthetic cells intracellular connections let heterocyst to transport fixed nitrogen to neighboring cells/receive carbohydrates metabolic cooperation w/ different prokaryotic species occurs in surface-coating colonies known as biofilms cells in biofilms secrete signaling molecules that recruit nearby cells causing colonies to grow (also make polysaccharides and proteins to stick cell to substrate/each other) channels in biofilms let nutrients reach cell in interior and waste to be expelled contribute to tooth decay (brush teeth to break biofilms) Lessons from Molecular Systematic small-subunit rRNA is used as a marker for evolutionary relationships seems that many prokaryotes classified as bacteria are more related to eukaryotes and need their own domain rise to Achaea molecular systematic shows the significance of horizontal gene transfer in evolution of prokaryotes over many years prokaryotes have acquired genes from even distantly related species and continue to do so so big portions of the genomes of many prokaryotes are mosaic of genes imported from other species Achaea share traits w/ bacteria and eukaryotes unique >extremephiles are Achaea that live in extreme conditions extreme halophiles live in high saline environments extreme thermophiles thrive in very hot environments don’t denature in high temp because their DNA and proteins have adaptations that make them stable at high temperatures some Achaea live in moderate conditions methagons are archeae that release methane as a byproduct of their unique ways of obtaining energy (use CO2 to oxidize H2 produced energy + methane waste (are poisoned by O2)) many extreme halophiles/all machineguns are Achaea in clade euryarchaeota, has some thermophiles (most of which are in 2nd clade crenarchaeota) many of both are not extremophiles Bacteria prokaryotes people are more aware of every major mode of nutrition/metabolism is represented among bacteria (pg 568 + 569 for major groups of bacteria … probably not needed for the test) Prokaryotes play crucial roles in the biosphere Chemical recycling >all atoms in organic molecules were part of inorganic substances like soil/air/water will eventually return >ecosystem depends on continual recycling of chemical elements between nonliving/living components and prokaryotes play a major role chemo heterotrophic prokaryotes are decomposers (break down dead organism unlocking carbon supplies) without them all life would cease prokaryotes convert molecules to forms that can be taken up by other organisms under some conditions, prokaryotes can increase the availability of nutrients that plants require for growth (nitrogen, prosperous, and potassium) prokaryotes can also decrease the availability of key plant nutrients (when they “immobilize” nutrients by using them to synthesize molecules that remain in their cells) Prokaryotes play a role in ecological Interactions prokaryotes often form symbiotic associations w/ larger organisms (host) and the smaller is known as the symbiotic many cases mutualism happens (both benefit) some commenalism happens (one benefits; other neutral some engage in parasitism (parasite eat cell contents of host harm but not usually kill…immediately) parasites that cause disease are pathogens (many are prokaryotes) very existence of an ecosystem can depend on prokaryotes (like ecological communities near hydrothermal vents no light so they depend on bacteria that harvest compounds like H2S released from vents) Prokaryotes have both beneficial/harmful impacts only a fraction are harmful (other play essential roles) Mutualitics bacteria in humans (like in the gut) are helpful help digest food intestine can’t break down by itself some signals human genes to build the network of intestinal blood vessels necessary to absorb nutrient molecules All pathogenic prokaryotes are bacteria some bacterial diseases are transmitted by other species (fleas/ticks) Pathogen prokaryotes cause illness by poisons, exotoxins and endotoxins >Exotoxins are proteins secreted by certain bacteria and other organisms >Endotoxins are lipopolysaccharide components of the outer membrane of gram negative bacteria in contrast to exotoxins, endotoxins are released only when the bacteria die/their cell walls break down Antibiotics save lives resistance is currently evolving (rapid production/natural selection/horizontal gene transfer) Prokaryotes in Research and Technology prokaryotes has led to new applications in biotechnology (E.coli in gene cloning and Agrobacterium tumefaciens in producing transgenic plants) bacteria can also make natural plastics (PHA) Prokaryotes are useless in bioremediation, the use of organism to remove pollutants from soil, air, or water through genetic engineering humans can modify bacteria to produce vitaments/antibitocis/hormones and so many more applications usefulness of prokaryotes derives from their diverse forms of nutrition/metabolism all metabolic versatility evolved prior to the appearance of the structural novelties in the evolution of eukaryotic organisms