* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download TamarEldad
Survey
Document related concepts
Site-specific recombinase technology wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Designer baby wikipedia , lookup
Gene expression profiling wikipedia , lookup
Microevolution wikipedia , lookup
Point mutation wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Gene nomenclature wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Transcript
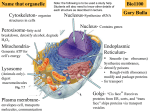
Predicting Protein Function Annotation using ProteinProtein Interaction Networks By Tamar Eldad Advisor: Dr. Yanay Ofran 89-385 Computational Biology - Projects Workshop Bar-Ilan University, the Mina and Everard Goodman Faculty of Life Sciences 1 Protein Function Prediction Exponential increase in the number of proteins being identified by sequence genomics projects Impossible to perform functional assay for every uncharacterized gene Turn to sophisticated computational methods for assistance in annotating the huge volume of sequence and structure data being produced homology-based annotation transfer sequence patterns structure similarity structure patterns genomic context microarray data 2 What is Function? Biological function has more than one aspect Sub-cellular to whole-organism context Physiological aspect Phenotype The need of a well-defined vocabulary 3 Protein Sequence: Protein Structure: 4 The Gene Ontology The Gene Ontology project is a major bioinformatics initiative with the aim of standardizing the representation of gene and gene product attributes across species and databases. The project provides a controlled vocabulary of terms for describing gene product characteristics and gene product annotation data. 6 The Gene Ontology Cellular component Molecular function Biological process DAG (1….N parent nodes) General Specific Term is assigned to Gene Product 7 The Gene Ontology 8 A New Approach Classical Biology – collect a set of features for each protein Systems Biology – study protein function in the context of a network Assemblies represent more than the sum of their parts 9 Protein Interactions Data on thousands of interactions in humans and most model species have become available mass spectrometry genome-wide chromatin immunoprecipitation yeast two-hybrid assays combinatorial reverse genetic screens rapid literature mining techniques 10 PPI Networks Data are represented as networks, with nodes representing proteins and edges representing the detected PPIs. 11 Existing Methods Alignment – aligning sequence-matching proteins between species and checking if they also share network alignment can teach us about conserved pathways between species Integration - data from different types of networks (i.e. protein, genetic, and transcriptional interaction networks) are integrated in order to get a better picture of the whole biological system Querying - find sub-networks similar to functional units (by comparing interactions and the proteins themselves) - likely to be functioning units too 12 New Method conserved network motifs between two species convey evidence for function similarity of the individual proteins that make up these motifs 1e-09 5e-15 8e-13 2e-10 HUMAN YEAST 13 New Method What do we need? 1. list of proteins in human cell 2. list of proteins in yeast cell 3. interactions in each cell 4. sequence similarity grades 5. known GO annotations 6. function distance calculation 14 Protein Lists - UniProt DB 15 Interaction Databases HPRD - The Human Protein Reference Database. Dip - Database of Interacting Proteins. Mips -Munich information center of proteins sequences IntAct – interaction molecular database. Reliable interaction performs one of these conditions: 1. was at least observed in 2 different experiments. OR 2. was reported in 3 different articles. 16 Sequence Similarity Grades BLAST - bl2seq YEAST HUMAN 1 2 3 4 1 - 0.008 3e-18 X 2 10 - 0.02 3.6 17 GO annotations –UniProt DB 18 Evidence Codes 19 Function Distance Calculation 20 Implementation 1. Prepare similarity matrix for cutoff e-value 2. Find all components of size N – 1 (DFS search) 3. Compare sub-graphs found using similarity matrix 4. Add N-th non-similar component to each pair of matching graphs 5. Get GO function annotation of N-th components 6. Calculate average distance of N-th component’s function 21 Quality Assurance 1. Compare to random-pair annotation No-sequence similarity 2. Compare to sequence-similar annotation BLAST Only proteins under cut-off value Human genes only 22 Detailed Results graph1 new comp go func graph2 new comp go func term type Eval average ,4814,4256,591,1584, Q12495 GO:0005515 ,4253,1335,2447,2353, Q9UHD2 GO:0005515 MolecularFunction 4 0.079 ,4814,4256,591,1584, Q12495 GO:0030528 ,4253,1335,2447,2353, Q9UHD2 GO:0030528 MolecularFunction 3 0.079 ,4814,4256,591,1584, Q12495 GO:0006334 ,4253,1335,2447,2353, Q9UHD2 GO:0006334 BiologicalProcess 0 0.079 ,4814,4256,591,1584, Q12495 GO:0005515 ,4253,1335,2447,2353, O15111 GO:0005515 MolecularFunction 1 0.079 ,4814,4256,591,1584, Q12495 GO:0005515 ,4253,1335,2447,2353, O15111 GO:0005515 MolecularFunction 12 0.079 ,4819,2,236,234, P16649 GO:0016584 ,4354,2303,2890,3693, P55060 GO:0016584 BiologicalProcess 1 0.062 ,4819,2,236,234, P16649 GO:0016565 ,4354,2303,2890,3693, Q96KB5 GO:0016565 MolecularFunction 1 0.062 ,4819,2,236,234, P16649 GO:0016584 ,4354,2303,2890,3693, Q15699 GO:0016584 BiologicalProcess 8 0.062 ,4819,2,236,234, P16649 GO:0016584 ,4354,2303,2890,3693, Q15699 GO:0016584 BiologicalProcess 5 0.062 ,4867,2966,168,1224, P13393 GO:0000120 ,4387,1383,1452,2289, P63279 GO:0000120 CellularComponent 4 0.041 ,4867,2966,168,1224, P13393 GO:0000120 ,4387,1383,1452,2289, P63279 GO:0000120 CellularComponent 3 0.041 ,4867,2966,168,1224, P13393 GO:0000126 ,4387,1383,1452,2289, P63279 GO:0000126 CellularComponent 7 0.041 dist 23 Results E-value 5e-05 24 Play with Parameters • Change graph size • Lower e-value • Start with larger amount of connected components • Use only graphs with higher connectivity • Non-similar proteins can be any protein in the graph • Different network topology • Limit number of paired proteins 25 Results 26 Conclusions Most results are random Significant improvement only for Biological Process prediction Still far behind Homology Based Transfer 27 Summary Functional annotation is one of the greatest challenges in the post-genomic era PPI data for functional annotation as a new approach for promoting this field Method tried out is unsuccessful Other Ideas: Find a more specific search pattern Start from best results – what specializes them? 28 References Friedberg,I. (2006) Automated function prediction: the genomic challenge. Brief. Bioinform. Accepted for publication Sharan R, Ulitsky I, Shamir R: Network-based prediction of protein function. Mol Syst Biol 2007, 3:88. Sharan R, Ideker T: Modeling cellular machinery through biological network comparison. Nature Biotechnology 24, 4: 427 - 433. http://www.geneontology.org/ http://www.chem.qmul.ac.uk/iubmb/enzyme/ 29 Thanks Advisor – Dr. Yanay Ofran Guys at the lab – Rotem, Vered, Sivan Roi Adadi & Omer Erel 30 Alignment Querying Integration Similarity Matrix E-value = 0.0005 YEAST HUMAN 1 2 3 4 1 - 0.008 TRUE 3e-18 TRUE X FALSE 2 10 FALSE - FALSE 0.02 FALSE 3.6 Neighboring matrix HUMAN CELL INTERACTIONS 1 2 3 4 1 - TRUE FALSE TRUE 2 TRUE - FALSE FALSE