* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Eukaryotic Genomes - Building Directory
Survey
Document related concepts
Cre-Lox recombination wikipedia , lookup
Gene expression wikipedia , lookup
Genomic imprinting wikipedia , lookup
Ridge (biology) wikipedia , lookup
Secreted frizzled-related protein 1 wikipedia , lookup
Non-coding DNA wikipedia , lookup
Community fingerprinting wikipedia , lookup
Transcriptional regulation wikipedia , lookup
List of types of proteins wikipedia , lookup
Genome evolution wikipedia , lookup
Promoter (genetics) wikipedia , lookup
Gene expression profiling wikipedia , lookup
Molecular evolution wikipedia , lookup
Gene regulatory network wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Transcript
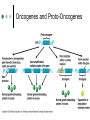
Eukaryotic Genomes Chromatin Structure If stretched out, each chromosome would be approximately 4cm long Chromatin is made up of DNA and the proteins that condense it If DNA did not associate with proteins in this way, what would be the problem? Chromatin Structure Nucleosomes The DNA double helix wraps around proteins called histones This gives the appearance of “beads on a string” The “beads” (nucleosomes) are joined by “string” (linker DNA) Chromatin Structure The nucleosomes interact with one another to coil and fold multiple times in a specific conformation 30-nm fiber Looped domains Mitotic chromosomes (most tightly condensed) The Regulation of Gene Expression All cells in an organism contain an identical genome (set of genes) However, the genes expressed in the cells of each type are unique Most of the DNA in eukaryotic genomes are noncoding – unsure of its purpose 25,000 genes in humans Only about 1.5% codes for protein The expression of specific genes is most commonly regulated at transcription, often in response to external signals Genes are turned on and off Transcription is the most common regulatory point in the pathway of gene expression, but it can happen at a number of places as well Cancer Mutations that alter genes that regulate cell growth and division can lead to cancer These mutations can be random and spontaneous More commonly, however, these mutations are caused by environmental influences • Examples? Oncogenes and Proto-Oncogenes Oncogenes: cancer-causing genes Proto-oncogenes: genes that stimulate normal cell growth and division Proto-oncogenes (normal) can become oncogenes (abnormal; cancer-causing) in a number of ways Translocation of gene Gene amplification – more copies of gene Point mutations All of these possibilities cause more growthstimulating protein to be produced Oncogenes and Proto-Oncogenes Tumor-Suppressor Genes The products of tumor-suppressor genes inhibit cell division and help prevent uncontrolled cell growth If a mutation occurs that affects tumor-suppressor genes, cancer may result p53 is a tumor-suppressor gene that activates other genes to repair DNA damage and inhibit further cell division The Multistep Model of Cancer Development Usually, more than one mutation is required to produce a cancerous cell Since cancer results from an accumulation of mutations, the longer we live the more likely we are to develop cancer In most cases of cancer, at least one active oncogene appears and at least one tumorsuppressor gene is mutated or lost Inherited Predisposition to Cancer Certain types of cancer can run in families Since cancer is caused by the accumulation of multiple genetic changes, an individual who inherits an oncogene or mutant tumor-suppressor gene is “that much closer” to accumulating the required number of mutations to develop cancer Transposons Transposons (“jumping genes”) are sequences of DNA that can move around to different positions within a single cell Can cause mutations and change the amount of DNA in a cell Discovered by Barbara McClintock in 1933 in maize (corn) Transposons Transposons are mutagens and damage the DNA of the host cell in a number of ways: A transposon that inserts itself in the middle of a functional gene will likely damage or disable that gene If a transposon leaves a gene, the gap will probably not be repaired correctly