* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download blumberg-lab.bio.uci.edu
Survey
Document related concepts
Protein design wikipedia , lookup
Protein folding wikipedia , lookup
Protein domain wikipedia , lookup
Homology modeling wikipedia , lookup
Protein moonlighting wikipedia , lookup
Protein structure prediction wikipedia , lookup
Intrinsically disordered proteins wikipedia , lookup
Protein purification wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
Western blot wikipedia , lookup
Protein mass spectrometry wikipedia , lookup
List of types of proteins wikipedia , lookup
Transcript
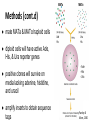
Presented by Evelia Ayala and Muaeen Obadi Interactome •Interactome: the set of protein-protein interactions that occur in a cell •Molecular interactions mediate biological functions that maintain normal cellular activities Li et al., 2004 Why focus on the interactome? ● large amounts of data produced by genomic projects ● fully sequenced genomes are loaded with novel genes ● identification of interactions would contribute to functional interpretation of genome Why use two hybrid screening? ● limitations of other methods ● functional prediction - functions of characterized proteins give insight into novel proteins ● emphasis on complementary approaches ● yeast-two hybrid system was the only well-established genomic approach for a genomewide scale analysis Two Hybrid screening ● method to detect protein-protein interactions ● transcription factor split into two fragments: bait protein prey protein binding domain (BD) & activation domain (AD) ● each fragment is fused to a different protein BD: bait protein & AD: prey protein ● if the two proteins interact, the BD & AD bind to allow for transcription of reporter gene Pandey & Mann, 2000 Methods ● Open reading frames (ORFs) individually cloned into bait expression vector & prey expression vector ● each bait plasmid introduced into MATa cell ● each prey plasmid introduced into MATα cell bait vector prey vector transform MATa MATα Methods (cont.d) ● mate MATa & MATα haploid cells ● diploid cells will have active Ade, His, & Ura reporter genes ● positive clones will survive on media lacking adenine, histidine, and uracil ● amplify inserts to obtain sequence tags Ade His Ura His - Ade - His - Ura Pandey & Mann, 2000 Outline of extensive two-hybrid analysis ● transformants divided into bait & prey pools ● pools systematically mated ● positive clones selected ● sequence tagging to obtain interaction sequence tags (ISTs) Results (condensed) - ISTs: pair of tag sequences for bait & prey Comparing results - comparing with an independent yeast two-hybrid project that used different strategies The Overlap in results ● They only share a small overlap of interactions. ● The known interactions from the YPD were shared significantly higher between data sets. Genome Wide Two Hybrid Interaction Map ● Removal of redundancy led to the core data. ● To confirm no background noise they analyzed dataset from YPD. ● Confirmed large cluster of interactions. Interaction Networks The positives of the approach ● For genome wide interaction mapping this is the most feasible approach. ● Hypothetical networks arise which become of interest for studies. ● Better understanding of cellular processes can be developed by testable hypothesis of these networks. The cons of the approach ● Dataset from different projects lack the ability to provide large overlap. ● In the nature of large scale screenings, interactions tend to escape detection. ● The two hybrid method itself has the concern of reliability. Topics of Discussion ● The project fails to restate ~90% of the two hybrid interaction identified in conventional studies. ● These protein to protein interaction have been added the the database creating large networks. The need for bioinformatics to read these networks has become necessary. ● To avoid complexes sharing protein components in silico there needs be data collected on architecture and their occurrence. Further Readings - Ito, T., Tashiro, K., Muta, S., Ozawa, R., Chiba, T., Nishizawa, M., Yamamoto, K., Kuhara., Sakaki, Y. (2000). Toward a protein-protein interaction map of the budding yeast: A comprehensive system to examine two-hybrid interactions in all possible combinations between the yeast proteins. PNAS, 97: 1143-1147. - Uetz et al. (2000). A comprehensive analysis of protein-protein interactions in Saccharomyces cerevisiae. Nature, 403: 623-627.