* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download KOX1, KAP1
Cell nucleus wikipedia , lookup
Biochemical switches in the cell cycle wikipedia , lookup
Magnesium transporter wikipedia , lookup
Protein moonlighting wikipedia , lookup
Signal transduction wikipedia , lookup
Cellular differentiation wikipedia , lookup
List of types of proteins wikipedia , lookup
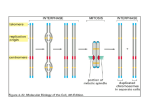
Extra Slides DATA TO SUPPORT STATEMENTS IN PRESENTATION Outline of talk General background Introduction to the project Experimental design Experiments and results Conclusions Future experiments Prior demonstrations of KAP1 complexes Proteins involved in complex Method 1 HP1a, KAP1 Yeast 2 Hybrid 2 KOX1, KAP1 Yeast 2 Hybrid 3 NuRD/HDAC, KAP1 4 KIP21, KAP1 5 KIP41, KAP1 6 KOX1, KAP1, HP1a Summary Reference HP1a was used as a bait and TIF1b (KAP1) was isolated Le Douarin et al 1996 KRAB domain of KOX1 was used as a bait and KAP1 was isolated Moosmann et al. 1996 Yeast 2 Hybrid Mi2a (= NuRD/HDAC) was shown interact with KAP-1PHD/BROMO but not Mi2b Schultz et al. 2001 Yeast 2 Hybrid KIP21 and KIP41 (=SETDB1) were shown interact with KAP-1PHD/BROMO Schultz et al. 2002 Gel Electrophoretic Mobility Shift assay (in vitro) DNA+GAL4-KRAB+KAP1 ternary complex was shown to complex with HP1a Ryan et al. 1999 Structure of CSD complex Epigenetic Gene Silencing Epigenetic effects are those changes in gene function which are heritable through mitosis and/or meiosis and are not due to changes in DNA sequence. Main types of epigenetic information: – Cytosine DNA Methylation – Genomic impritning – Histone modifications CSD interacting proteins Chromatin based epigenetics Controls chromosome domains and also helps in cell differentiation X –inactivation (XIST locus. Genomic imprinting, parent-of origin-specific allele silencing) Developmental reprogramming of cell lineages Plasticity of stem cells Implications human biology and disease, including cancer and aging Expression Vectors Gal4 DBD HP1a pBridge HP1 KAP1 KAP1 TRP1 BAIT 9525 bp MET25 HA/NLS HP1a (BD-HP1a)-(HA-KAP1) Gal4 DBD KAP1 GAL4 AD HA PREY pACT2 Hela cDNA Lib 10114 bp LEU2 Gene Library (AD-?) ? Gal4 AD HeLa cell cDNA library Source of human genes (cDNA). Human cervical carcinoma cell line Estimated number of Independent Clones: 3.5 x 106 – Average insert (cDNA) size: 2.0 kb – cDNA size range: 0.5 – 4.0 kb HeLa cancer cells Hybrid proteins used in this study BAITS – – – – – BD-HP1a (BD-HP1A)-(HA-KAP1) BD-KAP1 (BD-KAP1)-(HA-HP1) (BD-HP1A)-(HACAF1p150) PREYS – AD-? (HeLa cell cDNA library; Clontech) – – – – – AD-KIP21 (SETDB1) AD-KIP41 (SETDB1) AD-Mi2b AD-KOX1 AD-Y2H6.2 (POGZ) Prior demonstrations of KAP1 & HP1 complexes Proteins involved in complex Method 1 HP1a, KAP1 Yeast 2 Hybrid 2 KOX1, KAP1 Yeast 2 Hybrid 3 NuRD/HDAC, KAP1 4 KIP21, KAP1 5 KIP41, KAP1 6 7 KOX1, KAP1, HP1a HP1a, POGZ (Y2H6.2) Summary Reference HP1a was used as a bait and TIF1b (KAP1) was isolated Le Douarin et al 1996 KRAB domain of KOX1 was used as a bait and KAP1 was isolated Moosmann et al. 1996 Yeast 2 Hybrid Mi2a (= NuRD/HDAC) was shown interact with KAP-1PHD/BROMO but not Mi2b Schultz et al. 2001 Yeast 2 Hybrid KIP21 and KIP41 (=SETDB1) were shown interact with KAP-1PHD/BROMO Schultz et al. 2002 Gel Electrophoretic Mobility Shift assay (in vitro) DNA+GAL4-KRAB+KAP1 ternary complex was shown to complex with HP1a Co-immunoprecipitation and Yeast Two Hybrid genetic screen Novel prey (POGZ / Y2H6.2) shown to interact with HP1a Ryan et al. 1999 Lechner et al, Unpublished Histone Code HP1 gene silencing Strahl et al. Nature 2000 Long term Goals Build complete HP1 network with all its partner proteins and to build a chromatin network database ! Map both the temporal and Spatial formation of these complexes Get in depth and complete understanding of gene regulation in cells Statement of Problem Studies show that HP1a binds with several proteins and presumably performs different activities (mainly in regulating chromatin structure and function). But, little is known about what complexes exist in vivo and what controls their formation Assumptions The main assumption in this experiment is that there are regulatory factors that help in the formation of HP1 complexes. Yeast system does not interfere with any of the interactions. (relatively ‘safe’; most of the initial HP1 interactions were done in yeast screens.) Feasibility of this screen can be demonstrated by showing a ternary complex formation with positive controls. Limitations Yeast might inactivate or destroy the foreign (human) proteins. Some of these human proteins might be toxic to yeast. Functional homology might be present between yeast proteins and the human proteins. Presence of false positives or false negatives. Bait proteins that auto-activate cannot be used. Cannot directly extrapolate the results to higher eukaryotes without further Histone H3 tail bound to Chromo Domain of Drosophila HP1a Yelow: Me2k9 Red/Orange: Me3k9 Y : Tyrosine W: Tryptophan E : Glutamine T : Threonine K : Lysine V : Valine N : Asparagine S : Serine A : Alanine Jacobs and Khorasanizadeh (2002), Science express Mock transformation Number of Transformants = ~ 2 x 106 per ml; For a 5ml final volume of transformed cells number of library colonies screened is around ~107 ! Large enough to cover all clones in the library. (Number of independent clones in the library: 3.5 x 106; clontech ) HIS3 expression seems to be Leaky on –L/T/H plates S. No PARENT PLATE PREY 1 SD - LT pACT-Mi2b 2 SD - LT pACT-KIP21 3 SD - LT pACT-KIP41 4 SD - LT pACT TO COLONIES SD - LT 10 -20 SD -LTH 0 SD - LT 100-150 SD -LTH 20 -30 SD - LT 100-150 SD -LTH 1 SD - LT Lawn SD -LTH 20 -30 All colonies have BAIT in them (pBRIDGE-Gal4HP1a-HA-KAP1). BAIT and empty pACT seem to grow on –L/T/H but not – L/T/H/A (observation from mock transformation). HIS3 titration with competitive inhibitor 3-AT This assay is to determine the amount of 3-AT needed to reduce noise (growth when there is no interaction) on selection plates. S.no -- L/T/H with 3-AT @ different concentrations plasmid 0mM 1 pACT ~60 2 KIP21 TNC 3 KIP41 SV40T + p53 positive control 1mM ~24 3mM 5mM 10mM 15mM 3 No colonies ~100 10-20 No colonies TNC ~100 0 No colonies Lawn not plated Lawn Lawn Lawn Law n -L/T/H/A Lawn NOTE : 3-AT is 3-AMINO-1,2,4-TRIAZOLE, a competitive inhibiotor of HIS3 gene product. TNC means too numerous to count All colonies have BAIT in them (pBRIDGE-Gal4HP1a-HA-KAP1). Cell Lysate prepared for Western blot used for Co-IPs Anti-HA used for IMMUNOPRECIPITATION Harsh conditions used to obtain cell lysate Blot demonstrates that the Anti-HA antibody is not as sensitive as expected. Who's calling the shots? Some transcription factors have, or recruit proteins that have, histone modification and remodeling activities (Fig. 1). Presumably, gene activation requires at least one such factor that can bind its recognition sequence within 'inactive' chromatin and recruit other factors that collaborate in altering local chromatin structure. These altered regions of chromatin would then expose binding sites for other factors, including the basal transcription machinery3. Histone modifications may also be necessary to allow RNA polymerase to transit across nucleosomal DNA sequences7. Also, whereas acetylation can be reversed by various histone deacetylases, there are no known histone demethylases. Therefore, once a genomic region is methylated, modified nucleosomes must be replaced rather than altered to remove this epigenetic mark. Further work will be required to understand the mechanisms responsible for the spread of local histone modifications and the impact of these modifications on chromatin structure and transcriptional regulation. Nature Genetics 36, 438 - 440 (2004) Figure 1. Transcription factors recruit chromatin modification enzymes, which in turn regulate chromatin structure in the vicinity of the promoter. Dashed lines indicate spread of chromatin changes outward from the promoter region. In this model, changes are reversible until nucleosomes are modified by histone methylation. Ac, acetylated; Me, methylated. Fig. 3. Model to explain the role of positive and negative factors in heterochromatin and euchromatin. Methylated amino acids in the histone H3 tail are indicated by red lettering, and acetylated residues are shown in blue. The underlying sequence of the satellite repeats promotes the formation of a regular array of stable nucleosomes, which are favoured substrates for methylation at H3 lysine 9 (K9) by SUVAR39H1. Binding of HP1 to long arrays of nucleosomes containing H3 methylated at K9 promotes the formation of the higher-order heterochromatin structure. Transcriptionally active euchromatin is generated by transcription factors binding to clustered recognition sequences resulting in the formation of DNase I hypersensitive sites (HS). The HS generate and maintain the open structure of the euchromatin by promoting H3 K9 acetylation and K4 methylation of neighbouring nucleosomes. They can also act as barriers preventing the spread of heterochromatin into neighbouring euchromatin. Trends Genet. 2002 May;18(5):252-8. How to identify different classes of interaction? My Interests How do cells in different tissues have different functions when they have the same genome? Is entire human genome expressed? – NO There is approximately one gene every ~75,000 base pairs. And only a fraction (~2%)of this part codes for polypeptides. Global gene expression patterns?