* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Enzyme Inhibition and Drug Action
Western blot wikipedia , lookup
Fatty acid synthesis wikipedia , lookup
Butyric acid wikipedia , lookup
Polyadenylation wikipedia , lookup
Restriction enzyme wikipedia , lookup
Citric acid cycle wikipedia , lookup
Nicotinamide adenine dinucleotide wikipedia , lookup
Ultrasensitivity wikipedia , lookup
Metalloprotein wikipedia , lookup
Oxidative phosphorylation wikipedia , lookup
Evolution of metal ions in biological systems wikipedia , lookup
Biochemistry wikipedia , lookup
NADH:ubiquinone oxidoreductase (H+-translocating) wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Catalytic triad wikipedia , lookup
Biosynthesis wikipedia , lookup
Discovery and development of neuraminidase inhibitors wikipedia , lookup
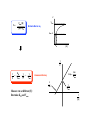
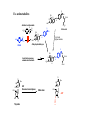
Enzyme Inhibition and Drug Action •Malfunction of enzyme •Introduction of enzyme by microorganism Disease Inhibition of enzyme - Interesting but difficult drug strategy O NH OH N N Inhib. of enzymes from microorganisms H2N N CO2H N H N CO2H Antibact. sulfonamide O R S NH O H2N H2N H2N OH - Enzyme only in microorg. - Different structure of enzyme, human and microorg OH OH N N O Folic acid O N N H From food Higher animals N H Dehydropteridinsyre PABA O O OH N H2N N H N NH N H N H Tetrahydrofolic acid Trimetoprim OCH3 N H2N N N N H N H Dehydrofolic acid NH2 Essintial processes, animals and bacteria N N H2N CO2H NH OH CO2H Folatreduktase OCH3 OCH3 CO2H CO2H Enzyme inhibition k3 k1 [ E-S ] E + S [ E-P ] E + P E: Enzyme S: Substrate P: Product k2 Two last steps ≈irreversible, E-S to E-P rate limiting Reaction velocity, V=k3 [E-S] Rate of form. ES: k1[E][S] Rate of decomp. ES: (k1 + k3)[E][S] Assume steady state ([E-S] doesn’t change) k1[E][S] = (k1 + k3)[E][S] [E-S] = [E] [S] (k2 + k3) / k1 [E-S] = Michaelis const.: KM = (k2 + k3) / k1 [E] = [E tot] - [E-S] [E] [S] KM [E-S] = ([Etot] - [E-S] [S] KM [E-S] = [Etot] [S] [S] + KM k3 k1 [ E-S ] E + S [ E-P ] E + P E: Enzyme S: Substrate P: Product k2 [E-S] = [Etot] [S] V=k3 [E-S] [S] + KM V = k3[Etot] [S] [S] + KM V = Vmax [S] [S] + KM Vmax: All enzyme sites occupied by S [S] [S]>>KM, Michaelis Menten eq. Vmax=k3 [Etot] ≈1 [S] + KM V Vmax Vmax : 2 KM [S] V V = Vmax Vmax [S] Michaelis Menten eq. [S] + KM Vmax : 2 KM [S] 1 V 1 V = KM Vmax 1 [S] + 1 Vmax Slope = Lineweaver-Burk eq. -1 KM Measure rate at different [S]: Determine KM and Vmax KM Vmax 1 Vmax 1 [S] Reversible and irreversible enzyme inhibitors E + 1 [ E-I ] E + 1 Reversible inhibition •Competitive •Non-competitive E +S [ E-S ] [ E-I ] If covalently bond to enzyme, bond relatively easily be broken i.e. hydrol. of ester P E + I P [ E-I ] 1 With inhibitor No inhib. V Binding to the same site Inhib. can be reversed by increasing [S] Vmax unchanged KM increase Structural resemblance S and I Designed I drugs - Antimetabolites Slope = -1 KM KM Vmax 1 Vmax 1 [S] O Ex. antimetabolites N N H2N Antibact. sulfonamide H2N O R S NH O OH PABA N CO2H N H N CO2H O H2N N N Folic acid OH OH N O H2N NH OH N H From food Higher animals N H Dehydropteridinsyre O NH OH N N Essintial processes, animals and bacteria H2N N CO2H N H N H CO2H O O Me HN O HO N O HO Me HN HIV Reverse transcriptase O RNA chain HO N O AZT OH Thymidin N N N OH Ex. transition state analogs NH2 N N N N HO Adenosine deaminase N HN OH N N HO O HO N HN N N HO O HO O H2N OH OH O HO OH Inosine Adenine sp3 OH N HN N HO O HO OH Coformycin Also metab. of anticancer / antiviral adenine deriv. Reversible inhibition •Competitive •Non-competitive E +S P [ E-S ] +I [E-S-I] E + I [ E-I ] +S 1 Binding to different sites Inhib. can be reversed by increasing [S] Vmax decrease KM unchanged Inhib. and substrate very different structures Difficult to design inhib. With inhibitor No inhib. V Slope = -1 KM KM Vmax 1 Vmax 1 [S] Irreversible enzyme inhibitors E + 1 [ E-I ] Often covalent bonds between E and I Enzyme is permanently modified and inactivated •Affinity labels and active site directed irreversible inhibitors •Mechanism based irreversible enzyme inactivators Structural recemblance with substrate Electrophilic - alkylate nucleophilic subst in enzyme active site Low selectivity - generally highly toxic •Affinity labels and active site directed irreversible inhibitors •Mechanism based irreversible enzyme inactivators Suicide substrate - kcat inhibitors - Trojan horse inhib. - latent alkylating agent ≈ Pro-drug, must be activated by the enzyme Penicillins are cleaved by β-lactamase O OH O N O R N H H O β-Lactamase O S R OH HN N H H S Clavulanic acid irreversibly inhibits β-lactamase OH O O OH N H Nu O O O O H H H O O O HN H OH OH OH O OH HN H O OH Nu Nu O O O OH HN HH O Nu OH