* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Abstract - WSU Horticulture
Protein mass spectrometry wikipedia , lookup
Cooperative binding wikipedia , lookup
Bimolecular fluorescence complementation wikipedia , lookup
Western blot wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
Protein purification wikipedia , lookup
Protein structure prediction wikipedia , lookup
Protein–protein interaction wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Protein domain wikipedia , lookup
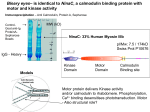
Structure-Function Relationships of Calcium/Calmodulin-dependent Protein Kinase (CCaMK): Agricultural and Ecological Implications Biological nitrogen fixation in legumes involves a complex microbiological process in which a certain type of bacteria (Rhizobium) fixes atmospheric nitrogen and converts it into a form the plant can use. This symbiotic system would reduce the cost of heavy use of nitrogen fertilizers in agriculture. Nitrogen fixation in legumes (bacterial symbiosis) has received considerable attention and it is becoming clear that calcium/calmodulin-mediated signaling plays a key role in sensing and transducing the bacterial signal (Nod factor). A calcium/calmodulin-dependent protein kinase (CCaMK), cloned and characterized in our laboratory, is known to play a critical role in decoding the calcium signal, and it is required for bacterial symbiosis. CCaMK is an important effector protein of Ca2+/calmodulin-mediated signaling and has been established as a critical regulator of both bacterial and fungal symbioses in plants. It contains a Ser/Thr kinase domain in its N-terminal, and two other regulatory domains: a calmodulin-binding/autoinhibitory domain (CaMBD/AID) and a visinin-like domain. In addition, recent studies have revealed the presence of two phosphorylation sites, S343 and S344. Recently, we discovered two different regulatory mechanisms of CCaMK. The first was the discovery of an intramolecular interaction during the phosphorylation of this kinase. Our published data revealed that S343D and S344D mutants were drastically compromised in their affinity to calmodulin (CaM), but our unpublished data indicated that the synthetic-peptides corresponding to the mutated CaM-binding domains were able to interact with CaM. These results led us to hypothesize that there is an intramolecular interaction involving S343 and S344, and other region(s), most likely the kinase domain of CCaMK, which is critical for its activity. In order to test this hypothesis, we created a series of progressive deletions in the CCaMK protein to determine the amino acid in the kinase domain that interacts with these phosphorylation sites. The deletion mutants were subjected to calmodulin-binding assay to observe their recovery of affinity to calmodulin. We were able to identify a critical amino acid (T227) that interacts independently with these two phosphorylation sites. However, our results also suggest that there must be other interactor(s) that may be involved during phosphorylation of S344 and S343. The second approach involved altering CCaMK’s calmodulin-binding capacity. It has been reported that CCaMK is regulated by calcium and calcium/calmodulin separately. This dual regulation is critical for the activation of this kinase and its interaction with its substrate protein, IPD3. Furthermore, CCaMK contains a regulatory area such as a calmodulin-binding domain in which the calmodulin binds and promotes substrate phosphorylation. Our data revealed that a site direct mutation in the calmodulin-binding domain of CCaMK increased its binding to CaM. This result suggests that the affinity of CCaMK to calmodulin could be altered by site-directed mutagenesis, providing an effective approach to study how calmodulin regulates CCaMK in terms of phosphorylation activity; and therefore the regulation of symbioses in Medicago truncatula. Site direct mutation W342F within the calmodulinbinding domain showed a considerable increase in calmodulin binding affinity which resulted in altered nodule development.