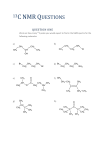
* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download slides - NMRbox
Ancestral sequence reconstruction wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Magnesium transporter wikipedia , lookup
Peptide synthesis wikipedia , lookup
Ribosomally synthesized and post-translationally modified peptides wikipedia , lookup
Protein moonlighting wikipedia , lookup
Cell-penetrating peptide wikipedia , lookup
Protein (nutrient) wikipedia , lookup
Protein folding wikipedia , lookup
Genetic code wikipedia , lookup
Metalloprotein wikipedia , lookup
List of types of proteins wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Expanded genetic code wikipedia , lookup
Western blot wikipedia , lookup
Protein adsorption wikipedia , lookup
Pharmacometabolomics wikipedia , lookup
Homology modeling wikipedia , lookup
Isotopic labeling wikipedia , lookup
Intrinsically disordered proteins wikipedia , lookup
Biochemistry wikipedia , lookup
Bottromycin wikipedia , lookup
Protein structure prediction wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
Frank Delaglio June 21 2016 Version 3 Identify many H-H short range NOE distances Supplement with torsions from J-Coupling values Assume standard peptide geometry Use simulated annealing to find a structure which matches distances Identify many H-H short range NOE distances Supplement with torsions from J-Coupling values Assume standard peptide geometry Use simulated annealing to find a structure which matches distances The network of distances is complicated, likewise the NOE spectra are complicated NOE distances are only qualitative A given peak might be the only evidence of an interaction A mis-assigned peak can be similarly problematic F F Y Beta Sheet Y Alpha Helix F [email protected] CA vs CB Chemical Shift Colored by Amino Acid Type Subtract Residue-Specific Random Coil Shift to form Secondary Shift Chemical Shift and Backbone Structure Motif Match database triplet with target, based on sum-of squares difference in chemical shifts, plus residue type homology term. Use central residue as predictor of phi and psi. Yang Shen, and Ad Bax, J. Biomol. NMR, 56, 227-241(2013). The TALOS-N database contains 580 proteins. On average, TALOS-N makes consistent predictions for about 90% of the residues. About 3.5% of the unambiguous predictions made by TALOS-N differ from the crystal structure. On average, the RMSD as reported by TALOS-N for the consensus predictions was 8.7 degrees for φ, and 8.5 degrees for ψ. The actual RMSD of the "correct" predictions relative to the crystal structures was 12.3 degrees for φ, and 12.1 degrees for ψ (which includes the uncertainty in the X-ray derived angles). TALOS-N can identify the chemical shift signature of a given χ1 rotamer for about 50% of the residues, all corresponding to cases where no extensive rotamer averaging is taking place. http://spin.niddk.nih.gov/bax/nmrserver/talosn Y Shen and A Bax: Protein backbone and sidechain torsion angles predicted from NMR chemical shifts using artificial neural networks. J. Biomol. NMR, 56, 227-241 (2013). SPARTA+ http://spin.niddk.nih.gov/bax/nmrserver/sparta Y Shen and A Bax: SPARTA+: a modest improvement in empirical NMR chemical shift prediction by means of an artificial neural network. J. Biomol. NMR, 48, 13-22 (2010). Consistent blind protein structure generation from NMR chemical shift data Proc Natl Acad Sci USA, (2008) 105, 4685-4690 Yang Shen Oliver Lange Frank Delaglio Paolo Rossi James M. Aramini Gaohua Liu Alexander Eletsky Yibing Wu Kiran K. Singarapu Alexander Lemak Alexandr Ignatchenko Cheryl H. Arrowsmith Thomas Szyperski Gaetano T. Montelione David Baker Ad Bax Using SPARTA Chemical Shift Prediction to Improve ROSETTA Scoring Function 0.57 0.69 0.64 0.66 0.60 2.07 0.70 1.10 2.03 Structures of Two Designed Proteins with High Sequence Identity NMR structures of Ga88 and Gb88 NMR structures vs csRosetta models Patrick A. Alexander, Yanan He, Yihong Chen, John Orban, and Philip N. Bryan PNAS, 2007, 104:11963-11968 PNAS, 2008, 105:14412-14417 Mean-to-mean backbone RMSD 1.31A 1.07A NMR Structure CS-Rosetta Structure https://csrosetta.bmrb.wisc.edu/csrosetta?key=6106ea029d9c Luke Arbogast Robert Brinson Yves Aubin John Marino 2D HN/N correlated NMR to reveal High Order Structure Practical 2D H/C correlated Methyl NMR of mAbs at natural abundance Multivariate analysis for easy evaluation of NMR fingerprint data NMR spectral fingerprinting without spectra Goal: Use NMR to provide direct answers about properties such as protein fold, excipient effects, glycosylation, stability, and aggregation. Changes in these properties can reduce the efficacy of a biotherapeutic, or cause harmful immune responses. Strategy: Use 2H, 13C, and 15N isotopic labeling to guide the development of natural abundance methods. [email protected] Fourier transforms are used to convert time-domain data to frequencydomain, and the information content is similar in all domains. Time-Domain Interferogram = Frequency Domain = Conventional FID NUS FID Interferogram NUS Interferogram Fourier Spectrum NUS Fourier Spectrum Conventional FID NUS FID Interferogram NUS Interferogram Fourier Spectrum NUS IST Reconstruction Scaled Intensity FT vs NUS FT vs FT Scaled Intensity Scaled Intensity Top 10 Drugs by US Sales Accounts for $60 Billion of $315 Billion Four of the Top 10 are Biologics Product Sales 4/2014 - 3/2015 Used For Type Humira $8,290,106,091 Anti-inflammatory mAb Abilify $7,995,192,015 Antipsychotic Small molecule Sovaldi $6,957,331,432 Hepatitis C Small molecule Crestor $5,958,997,432 Reduce cholesterol Small molecule Enbrel $5,953,627,734 Autoimmune diseases Protein/IgG1 Harvoni $5,398,133,616 Hepatitis C Small molecule x 2 Nexium $5,394,307,899 Reduce stomach acid Small molecule Advair Diskus $4,789,231,826 Asthma/COPD Small molecule x 2 Lantus Solostar $4,770,782,304 Diabetes Protein Remicade $4,614,448,608 Autoimmune diseases mAb Biologics are life-changing and life-saving therapeutics. They are also expensive, an issue for everyone in the healthcare system. As originator biologics go off patent, less expensive biosimilars can be produced. Development and manufacture requires monitoring high-order structure, aggregation, stability, and modifications such as glycosylation. Statistics from IMS Health Inc. as quoted on medscape.com, May 6, 2015 Most Drugs work by Targeting a Specific Protein or Class of Proteins Most familiar drugs are small organic molecules, often discovered by synthetic chemists making many variations of a molecular scaffold. Often, more than one kind of drug can bind to a target, which also means, often one given drug will also bind to undesired targets, causing side effects. This is often discovered late in the development process, and is why many new drugs fail (failure rate is ~90% and hasn’t changed much). Salicylic acid Indomethacin Aspirin Acetaminophen COX-2: Cyclooxygenase-2 (prostaglandin synthase-2, blue) complexed with indomethacin (red) Ibuprofen Proteins Themselves Can be Used as Drugs: Biologic Therapeutics Insulin Epogen (Erythropoietin) Filgrastim (G-CSF) Regulates Glucose Levels Stimulates Red Blood Cell Production Stimulates White Blood Cell Production 51 amino acids 166 amino acids 177 amino acids Hierarchy of Protein High Order Structure If all of these aren’t right, and don’t stay right, the protein therapeutic is wrong. Changes in these properties can reduce the efficacy of a protein-based therapeutic, or cause dangerous immune responses. MET GLN ILE PHE VAL LYS THR LEU THR GLY LYS THR ILE THR LEU GLU VAL GLU PRO SER Tertiary Structure (Protein Fold) Primary Structure (Amino Acid Sequence) Quaternary Structure (Complex or Aggregate of Two or More Proteins) Secondary Structure (Helix, Sheet, Turn, Coil) Harder to Measure Unfolded G-CSF, 15N Labeled Native G-CSF, 15N Labeled Y Aubin, DJ Hodgson, WB Thach, G Gingras, and S Sauvé: Monitoring Effects of Excipients, Formulation Parameters and Mutations on the High Order Structure of Filgrastim by NMR. Pharm Res., 32, 3365-3375 (2015). 15N ppm Minor Oxidized Species 1H 1H ppm ppm Knowledge of NMR assignments and structure allows careful peak-by-peak analysis which can correlate spectral changes with specific and subtle structural details such as a sidechain reorientation. NMR assignment is complicated, and generally requires 13C / 15N labeled protein. Y Aubin, DJ Hodgson, WB Thach, G Gingras, and S Sauvé: Monitoring Effects of Excipients, Formulation Parameters and Mutations on the High Order Structure of Filgrastim by NMR. Pharm Res., 32, 3365-3375 (2015). Antibody Proteins as Drugs: A Natural Source of Diverse Binding Partners Instead of synthesizing and testing large numbers of small organic molecules, genetic engineering can be used to select and duplicate antibodies that bind with high affinity and specificity to most any target … humans generate about 10 billion antibody variations ... IgG Antibody, ~150 kDa Two identical heavy chains, two identical light chains, symmetric. Glycans at two amino acids Variable Regions in blue and purple Pharma Loves mAbs and Igs – Find Hit Quickly, Re-use Biomanufacturing Platform Fv fusion mAb conjugate mAb mAb mAb mAb mAb mAb mAb Fc fusion mAb mAb mAb mAb protein mAb Fc fusion virus peptide mAb mAb mAb mAb mAb mAb mAb mAb mAb mAb mAb Example from Amgen Drug Development Pipeline - Adapted from www.amgenpipeline.com mAb NIST Principal Investigators: John Schiel and Trina Formolo Standard Reference Material: issued under NIST trademark with specified property values and associated uncertainties. Humanized mAb (IgG1κ) expressed in murine culture. Frozen bulk “Drug-like substance” donated by MedImmune. Extensive interlaboratory characterization by 65+ Biopharma, Instrument, Academic, FDA participants. Data Publically available at igg.nist.gov Amino Acid Sequencing Amino Acid Analysis N- and C-terminal Sequencing Peptide Mapping by MS S-S Bridge Analysis Glycosylation Analysis Molecular Weight Information Isoelectric Focusing SDS-PAGE Extinction Coefficient Post-Translational Modifications Spectroscopic Profiles: CD, NMR LC: SEC, RP, IEX NIST RM 8670 NISTmAb 150 kDa, 4 Chains, Symmetric Model based on Protein Data Bank Structures 1HZH 2GJ7 3IXT Size of a mAb ~1,300 Amino Acids Amino Acid Count Malate Synthase G 723 Amino Acids Protein Data Bank (PDB) NMR Structure Depositions by Year Since mAbs are much larger than the proteins usually studied by NMR, expectation is that fingerprinting mAbs by NMR would not be practical, especially without isotopic enrichment. Methyl groups are excellent reporters of protein fold, and 13C has higher natural abundance than 15N (1.07% vs 0.33%). Rapid rotation of methyl groups mitigates the effects of slow molecular tumbling in large proteins, for greatly improved spectra. Non-Uniform Sampling (NUS) can further increase spectral quality obtainable in a given amount of measurement time, making NMR fingerprinting of mAbs practical. Uniform Sampling 50% NUS NISTmAb Methyl Groups Ala Ile Leu Met Thr Val LW Arbogast, RG Brinson, and JP Marino: Mapping Monoclonal Antibody Structure by 2D 13C NMR at Natural Abundance. Anal. Chem., 87,3556–3561 (2015). G2F Glycan: ~55% G1F Glycan: ~30% G0F Glycan: ~15% Two identical heavy chains of the Native NISTmAb each have a glycan bonded to the sidechain N of Asn 297 b1-4 galactosidase G0F Glycan: ~100% PNGase F Quenched after Partial Reaction Deglyosylated NISTmAb: ~40% Asparagine 297 Aspartic Acid 30 spectra shown in overlay, normalized to uniform maximum intensity, colored by sample type. The four 16-scan spectra in the series are not shown for high noise. Each spectrum is represented exactly as a single object in a multdimensional space. The coordinates of the object are all of the spectral intensities. In this representation, similar spectra cluster together. Spectra with some features in common lie along lines and curves. PCA projects this space to a small number of dimensions along directions of maximum variance, so that it can be readily viewed and characterized. Component 3 Score Component 2 Score Component 3 Score G0F Native Partial Deglycosylation 40% NUS Native Component 2 Score As shown, the Native, G0F, and Deglycosylated samples are well-clustered. Note also that the NUS reconstructions are systematically different from the conventional data. In practice, this kind of PCA analysis is very sensitive to processing details such as baseline correction and phasing. Y Aubin, DJ Hodgson, WB Thach, G Gingras, and S Sauvé: Monitoring Effects of Excipients, Formulation Parameters and Mutations on the High Order Structure of Filgrastim by NMR. Pharm Res., 32, 3365-3375 (2015). pH 6.2 pH 5.5 pH 5.0 pH 4.5 pH 2.1 pH 4.0 pH 3.5 pH 2.6 pH 3.0 pH 3.4 The PCA analysis is accomplished in seconds, without the need for peak detection or assignment. PCA and other methods of multivariate analysis can reveal systematic behavior and outliers that might be hard to identify directly from inspection of spectra, even for an expert. Multivariate approaches become more useful with larger numbers of spectra, without becoming harder to do. Fourier transforms are used to convert time-domain data to frequencydomain, and the information content is similar in all domains. Time-Domain Interferogram = Frequency Domain = PCA on Spectra Of G-CSF PCA on Equivalent Interferograms of G-CSF PCA on Spectra Of G-CSF PCA on Equivalent Interferograms of G-CSF Apodization Removed in the Indirect Dimension PCA on Spectra Of G-CSF PCA on Equivalent Interferograms of G-CSF Masked with 50% NUS Schedule Apodization Removed in the Indirect Dimension PCA Results on Interferograms of G-CSF 50% NUS Component 2 vs 3 PCA Results on Interferograms of G-CSF 50% NUS Component 2 vs 3 2D H/C methyl spectral fingerprinting is practical at natural abundance for molecules as large as mAbs. Multivariate methods such as PCA can be used on spectra to reveal details of High Order Structure, post-translational modification, and excipient effects. Since NMR fingerprinting can potentially be performed without the need to identify peaks, it might be possible to develop even more efficient measurement strategies which do not produce spectra that can be analyzed visually, but nevertheless encode all the structural information of interest. Labeled samples in preparation now will allow us to explore backbone and sidechain dynamics, dipolar couplings, NMR HD exchange, etc., and relate these to high order structure, glycosylation, aggregation, and stability. If sufficient numbers of spectra can be measured, NMR spectral fingerprinting is a good potential target for machine learning approaches. NIST Disclaimer: Certain commercial equipment, instruments, and materials are identified in this presentation in order to specify the experimental procedure. Such identification does not imply recommendation or endorsement by the National Institute of Standards and Technology, nor does it imply that the material or equipment identified is necessarily the best available for the purpose.